diff --git a/README.md b/README.md
index 40b26c77..7ed74c16 100644
--- a/README.md
+++ b/README.md
@@ -1,36 +1,34 @@
-# Azure Machine Learning service sample notebooks
-
----
+# Azure Machine Learning service example notebooks
This repository contains example notebooks demonstrating the [Azure Machine Learning](https://azure.microsoft.com/en-us/services/machine-learning-service/) Python SDK
which allows you to build, train, deploy and manage machine learning solutions using Azure. The AML SDK
allows you the choice of using local or cloud compute resources, while managing
and maintaining the complete data science workflow from the cloud.
-* Read [instructions on setting up notebooks](./NBSETUP.md) to run these notebooks.
+
-* Find quickstarts, end-to-end tutorials, and how-tos on the [official documentation site for Azure Machine Learning service](https://docs.microsoft.com/en-us/azure/machine-learning/service/).
+## How to use and navigate the example notebooks?
-## Getting Started
+You can set up you own Python environment or use Azure Notebooks with Azure ML SDK pre-installed. Read [these instructions](./NBSETUP.md) to set up your environment and clone the example notebooks.
-These examples will provide you with an effective way to get started using AML. Once you're familiar with
-some of the capabilities, explore the repository for specific topics.
+You should always run the [Configuration](./configuration.ipynb) notebook first when setting up a notebook library on a new machine or in a new environment. It configures your notebook library to connect to an Azure Machine Learning workspace, and sets up your workspace and compute to be used by many of the other examples.
-- [Configuration](./configuration.ipynb) configures your notebook library to easily connect to an
- Azure Machine Learning workspace, and sets up your workspace to be used by many of the other examples. You should
- always run this first when setting up a notebook library on a new machine or in a new environment
-- [Train in notebook](./how-to-use-azureml/training/train-within-notebook) shows how to create a model directly in a notebook while recording
- metrics and deploy that model to a test service
-- [Train on remote](./how-to-use-azureml/training/train-on-remote-vm) takes the previous example and shows how to create the model on a cloud compute target
-- [Production deploy to AKS](./how-to-use-azureml/deployment/production-deploy-to-aks) shows how to create a production grade inferencing webservice
+If you want to...
+
+ * ...try out and explore Azure ML, start with image classification tutorials [part 1 training](./tutorials/img-classification-part1-training.ipynb) and [part 2 deployment](./tutorials/img-classification-part2-deploy.ipynb).
+ * ...learn about experimentation and tracking run history, first [train within Notebook](./how-to-use-azureml/training/train-within-notebook/train-within-notebook.ipynb), then try [training on remote VM](./how-to-use-azureml/training/train-on-remote-vm/train-on-remote-vm.ipynb) and [using logging APIs](./how-to-use-azureml/training/logging-api/logging-api.ipynb).
+ * ...train deep learning models at scale, first learn about [Machine Learning Compute](./how-to-use-azureml/training/train-on-amlcompute/train-on-amlcompute.ipynb), and then try [distributed hyperparameter tuning](./how-to-use-azureml/training-with-deep-learning/train-hyperparameter-tune-deploy-with-pytorch/train-hyperparameter-tune-deploy-with-pytorch.ipynb) and [distributed training](./how-to-use-azureml/training-with-deep-learning/distributed-pytorch-with-horovod/distributed-pytorch-with-horovod.ipynb).
+ * ...deploy model as realtime scoring service, first learn the basics by [training within Notebook and deploying to Azure Container Instance](./how-to-use-azureml/training/train-within-notebook/train-within-notebook.ipynb), then learn how to [register and manage models, and create Docker images](./how-to-use-azureml/deployment/register-model-create-image-deploy-service/register-model-create-image-deploy-service.ipynb), and [production deploy models on Azure Kubernetes Cluster](./how-to-use-azureml/deployment/production-deploy-to-aks/production-deploy-to-aks.ipynb).
+ * ...deploy models as batch scoring service, first [train a model within Notebook](./how-to-use-azureml/training/train-within-notebook/train-within-notebook.ipynb), learn how to [register and manage models](./how-to-use-azureml/deployment/register-model-create-image-deploy-service/register-model-create-image-deploy-service.ipynb), then [create Machine Learning Compute for scoring compute](./how-to-use-azureml/training/train-on-amlcompute/train-on-amlcompute.ipynb), and [use Machine Learning Pipelines to deploy your model](./how-to-use-azureml/machine-learning-pipelines/pipeline-mpi-batch-prediction.ipynb).
+ * ...monitor your deployed models, learn about using [App Insights](./how-to-use-azureml/deployment/enable-app-insights-in-production-service/enable-app-insights-in-production-service.ipynb) and [model data collection](./how-to-use-azureml/deployment/enable-data-collection-for-models-in-aks/enable-data-collection-for-models-in-aks.ipynb).
## Tutorials
The [Tutorials](./tutorials) folder contains notebooks for the tutorials described in the [Azure Machine Learning documentation](https://aka.ms/aml-docs)
-## How to use AML
+## How to use Azure ML
-The [How to use AML](./how-to-use-azureml) folder contains specific examples demonstrating the features of the Azure Machine Learning SDK
+The [How to use Azure ML](./how-to-use-azureml) folder contains specific examples demonstrating the features of the Azure Machine Learning SDK
- [Training](./how-to-use-azureml/training) - Examples of how to build models using Azure ML's logging and execution capabilities on local and remote compute targets.
- [Training with Deep Learning](./how-to-use-azureml/training-with-deep-learning) - Examples demonstrating how to build deep learning models using estimators and parameter sweeps
@@ -38,3 +36,21 @@ The [How to use AML](./how-to-use-azureml) folder contains specific examples dem
- [Machine Learning Pipelines](./how-to-use-azureml/machine-learning-pipelines) - Examples showing how to create and use reusable pipelines for training and batch scoring
- [Deployment](./how-to-use-azureml/deployment) - Examples showing how to deploy and manage machine learning models and solutions
- [Azure Databricks](./how-to-use-azureml/azure-databricks) - Examples showing how to use Azure ML with Azure Databricks
+
+---
+## Documentation
+
+ * Quickstarts, end-to-end tutorials, and how-tos on the [official documentation site for Azure Machine Learning service](https://docs.microsoft.com/en-us/azure/machine-learning/service/).
+
+ * [Python SDK reference]( https://docs.microsoft.com/en-us/python/api/overview/azure/ml/intro?view=azure-ml-py)
+
+
+---
+
+## Projects using Azure Machine Learning
+
+Visit following repos to see projects contributed by Azure ML users:
+
+ - [Fine tune natural language processing models using Azure Machine Learning service](https://github.com/Microsoft/AzureML-BERT)
+ - [Fashion MNIST with Azure ML SDK](https://github.com/amynic/azureml-sdk-fashion)
+
\ No newline at end of file
diff --git a/how-to-use-azureml/automated-machine-learning/automl_setup.cmd b/how-to-use-azureml/automated-machine-learning/automl_setup.cmd
index cd4a92c1..0f2478ea 100644
--- a/how-to-use-azureml/automated-machine-learning/automl_setup.cmd
+++ b/how-to-use-azureml/automated-machine-learning/automl_setup.cmd
@@ -23,6 +23,10 @@ if errorlevel 1 goto ErrorExit
call python -m ipykernel install --user --name %conda_env_name% --display-name "Python (%conda_env_name%)"
+REM azureml.widgets is now installed as part of the pip install under the conda env.
+REM Removing the old user install so that the notebooks will use the latest widget.
+call jupyter nbextension uninstall --user --py azureml.widgets
+
echo.
echo.
echo ***************************************
diff --git a/how-to-use-azureml/automated-machine-learning/automl_setup_linux.sh b/how-to-use-azureml/automated-machine-learning/automl_setup_linux.sh
index 442dfc0a..2f2e96cc 100644
--- a/how-to-use-azureml/automated-machine-learning/automl_setup_linux.sh
+++ b/how-to-use-azureml/automated-machine-learning/automl_setup_linux.sh
@@ -22,11 +22,13 @@ fi
if source activate $CONDA_ENV_NAME 2> /dev/null
then
echo "Upgrading azureml-sdk[automl,notebooks,explain] in existing conda environment" $CONDA_ENV_NAME
- pip install --upgrade azureml-sdk[automl,notebooks,explain]
+ pip install --upgrade azureml-sdk[automl,notebooks,explain] &&
+ jupyter nbextension uninstall --user --py azureml.widgets
else
conda env create -f $AUTOML_ENV_FILE -n $CONDA_ENV_NAME &&
source activate $CONDA_ENV_NAME &&
python -m ipykernel install --user --name $CONDA_ENV_NAME --display-name "Python ($CONDA_ENV_NAME)" &&
+ jupyter nbextension uninstall --user --py azureml.widgets &&
echo "" &&
echo "" &&
echo "***************************************" &&
diff --git a/how-to-use-azureml/automated-machine-learning/automl_setup_mac.sh b/how-to-use-azureml/automated-machine-learning/automl_setup_mac.sh
index 054223f8..298fc67d 100644
--- a/how-to-use-azureml/automated-machine-learning/automl_setup_mac.sh
+++ b/how-to-use-azureml/automated-machine-learning/automl_setup_mac.sh
@@ -22,13 +22,15 @@ fi
if source activate $CONDA_ENV_NAME 2> /dev/null
then
echo "Upgrading azureml-sdk[automl,notebooks,explain] in existing conda environment" $CONDA_ENV_NAME
- pip install --upgrade azureml-sdk[automl,notebooks,explain]
+ pip install --upgrade azureml-sdk[automl,notebooks,explain] &&
+ jupyter nbextension uninstall --user --py azureml.widgets
else
conda env create -f $AUTOML_ENV_FILE -n $CONDA_ENV_NAME &&
source activate $CONDA_ENV_NAME &&
conda install lightgbm -c conda-forge -y &&
python -m ipykernel install --user --name $CONDA_ENV_NAME --display-name "Python ($CONDA_ENV_NAME)" &&
- pip install numpy==1.15.3
+ jupyter nbextension uninstall --user --py azureml.widgets &&
+ pip install numpy==1.15.3 &&
echo "" &&
echo "" &&
echo "***************************************" &&
diff --git a/how-to-use-azureml/automated-machine-learning/classification-with-deployment/auto-ml-classification-with-deployment.ipynb b/how-to-use-azureml/automated-machine-learning/classification-with-deployment/auto-ml-classification-with-deployment.ipynb
index da6a3d99..68effc07 100644
--- a/how-to-use-azureml/automated-machine-learning/classification-with-deployment/auto-ml-classification-with-deployment.ipynb
+++ b/how-to-use-azureml/automated-machine-learning/classification-with-deployment/auto-ml-classification-with-deployment.ipynb
@@ -1,523 +1,522 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Automated Machine Learning\n",
- "_**Classification with Deployment**_\n",
- "\n",
- "## Contents\n",
- "1. [Introduction](#Introduction)\n",
- "1. [Setup](#Setup)\n",
- "1. [Train](#Train)\n",
- "1. [Deploy](#Deploy)\n",
- "1. [Test](#Test)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Introduction\n",
- "\n",
- "In this example we use the scikit learn's [digit dataset](http://scikit-learn.org/stable/modules/generated/sklearn.datasets.load_digits.html) to showcase how you can use AutoML for a simple classification problem and deploy it to an Azure Container Instance (ACI).\n",
- "\n",
- "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
- "\n",
- "In this notebook you will learn how to:\n",
- "1. Create an experiment using an existing workspace.\n",
- "2. Configure AutoML using `AutoMLConfig`.\n",
- "3. Train the model using local compute.\n",
- "4. Explore the results.\n",
- "5. Register the model.\n",
- "6. Create a container image.\n",
- "7. Create an Azure Container Instance (ACI) service.\n",
- "8. Test the ACI service."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Setup\n",
- "\n",
- "As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import json\n",
- "import logging\n",
- "import os\n",
- "import random\n",
- "\n",
- "from matplotlib import pyplot as plt\n",
- "from matplotlib.pyplot import imshow\n",
- "import numpy as np\n",
- "import pandas as pd\n",
- "from sklearn import datasets\n",
- "\n",
- "import azureml.core\n",
- "from azureml.core.experiment import Experiment\n",
- "from azureml.core.workspace import Workspace\n",
- "from azureml.train.automl import AutoMLConfig\n",
- "from azureml.train.automl.run import AutoMLRun"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "ws = Workspace.from_config()\n",
- "\n",
- "# choose a name for experiment\n",
- "experiment_name = 'automl-local-classification'\n",
- "# project folder\n",
- "project_folder = './sample_projects/automl-local-classification'\n",
- "\n",
- "experiment=Experiment(ws, experiment_name)\n",
- "\n",
- "output = {}\n",
- "output['SDK version'] = azureml.core.VERSION\n",
- "output['Subscription ID'] = ws.subscription_id\n",
- "output['Workspace'] = ws.name\n",
- "output['Resource Group'] = ws.resource_group\n",
- "output['Location'] = ws.location\n",
- "output['Project Directory'] = project_folder\n",
- "output['Experiment Name'] = experiment.name\n",
- "pd.set_option('display.max_colwidth', -1)\n",
- "pd.DataFrame(data=output, index=['']).T"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Opt-in diagnostics for better experience, quality, and security of future releases."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.telemetry import set_diagnostics_collection\n",
- "set_diagnostics_collection(send_diagnostics = True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Train\n",
- "\n",
- "Instantiate a AutoMLConfig object. This defines the settings and data used to run the experiment.\n",
- "\n",
- "|Property|Description|\n",
- "|-|-|\n",
- "|**task**|classification or regression|\n",
- "|**primary_metric**|This is the metric that you want to optimize. Classification supports the following primary metrics:
accuracy
AUC_weighted
average_precision_score_weighted
norm_macro_recall
precision_score_weighted|\n",
- "|**iteration_timeout_minutes**|Time limit in minutes for each iteration.|\n",
- "|**iterations**|Number of iterations. In each iteration AutoML trains a specific pipeline with the data.|\n",
- "|**n_cross_validations**|Number of cross validation splits.|\n",
- "|**X**|(sparse) array-like, shape = [n_samples, n_features]|\n",
- "|**y**|(sparse) array-like, shape = [n_samples, ], [n_samples, n_classes]
Multi-class targets. An indicator matrix turns on multilabel classification. This should be an array of integers.|\n",
- "|**path**|Relative path to the project folder. AutoML stores configuration files for the experiment under this folder. You can specify a new empty folder.|"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "digits = datasets.load_digits()\n",
- "X_train = digits.data[10:,:]\n",
- "y_train = digits.target[10:]\n",
- "\n",
- "automl_config = AutoMLConfig(task = 'classification',\n",
- " name = experiment_name,\n",
- " debug_log = 'automl_errors.log',\n",
- " primary_metric = 'AUC_weighted',\n",
- " iteration_timeout_minutes = 20,\n",
- " iterations = 10,\n",
- " n_cross_validations = 2,\n",
- " verbosity = logging.INFO,\n",
- " X = X_train, \n",
- " y = y_train,\n",
- " path = project_folder)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Call the `submit` method on the experiment object and pass the run configuration. Execution of local runs is synchronous. Depending on the data and the number of iterations this can run for a while.\n",
- "In this example, we specify `show_output = True` to print currently running iterations to the console."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "local_run = experiment.submit(automl_config, show_output = True)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "local_run"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Deploy\n",
- "\n",
- "### Retrieve the Best Model\n",
- "\n",
- "Below we select the best pipeline from our iterations. The `get_output` method on `automl_classifier` returns the best run and the fitted model for the last invocation. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "best_run, fitted_model = local_run.get_output()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Register the Fitted Model for Deployment\n",
- "If neither `metric` nor `iteration` are specified in the `register_model` call, the iteration with the best primary metric is registered."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "description = 'AutoML Model'\n",
- "tags = None\n",
- "model = local_run.register_model(description = description, tags = tags)\n",
- "local_run.model_id # This will be written to the script file later in the notebook."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create Scoring Script"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%writefile score.py\n",
- "import pickle\n",
- "import json\n",
- "import numpy\n",
- "import azureml.train.automl\n",
- "from sklearn.externals import joblib\n",
- "from azureml.core.model import Model\n",
- "\n",
- "\n",
- "def init():\n",
- " global model\n",
- " model_path = Model.get_model_path(model_name = '<>') # this name is model.id of model that we want to deploy\n",
- " # deserialize the model file back into a sklearn model\n",
- " model = joblib.load(model_path)\n",
- "\n",
- "def run(rawdata):\n",
- " try:\n",
- " data = json.loads(rawdata)['data']\n",
- " data = numpy.array(data)\n",
- " result = model.predict(data)\n",
- " except Exception as e:\n",
- " result = str(e)\n",
- " return json.dumps({\"error\": result})\n",
- " return json.dumps({\"result\":result.tolist()})"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create a YAML File for the Environment"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "To ensure the fit results are consistent with the training results, the SDK dependency versions need to be the same as the environment that trains the model. Details about retrieving the versions can be found in notebook [12.auto-ml-retrieve-the-training-sdk-versions](12.auto-ml-retrieve-the-training-sdk-versions.ipynb)."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "experiment_name = 'automl-local-classification'\n",
- "\n",
- "experiment = Experiment(ws, experiment_name)\n",
- "ml_run = AutoMLRun(experiment = experiment, run_id = local_run.id)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "dependencies = ml_run.get_run_sdk_dependencies(iteration = 7)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "for p in ['azureml-train-automl', 'azureml-sdk', 'azureml-core']:\n",
- " print('{}\\t{}'.format(p, dependencies[p]))"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.conda_dependencies import CondaDependencies\n",
- "\n",
- "myenv = CondaDependencies.create(conda_packages=['numpy','scikit-learn'], pip_packages=['azureml-sdk[automl]'])\n",
- "\n",
- "conda_env_file_name = 'myenv.yml'\n",
- "myenv.save_to_file('.', conda_env_file_name)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Substitute the actual version number in the environment file.\n",
- "# This is not strictly needed in this notebook because the model should have been generated using the current SDK version.\n",
- "# However, we include this in case this code is used on an experiment from a previous SDK version.\n",
- "\n",
- "with open(conda_env_file_name, 'r') as cefr:\n",
- " content = cefr.read()\n",
- "\n",
- "with open(conda_env_file_name, 'w') as cefw:\n",
- " cefw.write(content.replace(azureml.core.VERSION, dependencies['azureml-sdk']))\n",
- "\n",
- "# Substitute the actual model id in the script file.\n",
- "\n",
- "script_file_name = 'score.py'\n",
- "\n",
- "with open(script_file_name, 'r') as cefr:\n",
- " content = cefr.read()\n",
- "\n",
- "with open(script_file_name, 'w') as cefw:\n",
- " cefw.write(content.replace('<>', local_run.model_id))"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create a Container Image"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.image import Image, ContainerImage\n",
- "\n",
- "image_config = ContainerImage.image_configuration(runtime= \"python\",\n",
- " execution_script = script_file_name,\n",
- " conda_file = conda_env_file_name,\n",
- " tags = {'area': \"digits\", 'type': \"automl_classification\"},\n",
- " description = \"Image for automl classification sample\")\n",
- "\n",
- "image = Image.create(name = \"automlsampleimage\",\n",
- " # this is the model object \n",
- " models = [model],\n",
- " image_config = image_config, \n",
- " workspace = ws)\n",
- "\n",
- "image.wait_for_creation(show_output = True)\n",
- "\n",
- "if image.creation_state == 'Failed':\n",
- " print(\"Image build log at: \" + image.image_build_log_uri)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Deploy the Image as a Web Service on Azure Container Instance"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.webservice import AciWebservice\n",
- "\n",
- "aciconfig = AciWebservice.deploy_configuration(cpu_cores = 1, \n",
- " memory_gb = 1, \n",
- " tags = {'area': \"digits\", 'type': \"automl_classification\"}, \n",
- " description = 'sample service for Automl Classification')"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.webservice import Webservice\n",
- "\n",
- "aci_service_name = 'automl-sample-01'\n",
- "print(aci_service_name)\n",
- "aci_service = Webservice.deploy_from_image(deployment_config = aciconfig,\n",
- " image = image,\n",
- " name = aci_service_name,\n",
- " workspace = ws)\n",
- "aci_service.wait_for_deployment(True)\n",
- "print(aci_service.state)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Delete a Web Service"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "#aci_service.delete()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Get Logs from a Deployed Web Service"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "#aci_service.get_logs()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Test"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "#Randomly select digits and test\n",
- "digits = datasets.load_digits()\n",
- "X_test = digits.data[:10, :]\n",
- "y_test = digits.target[:10]\n",
- "images = digits.images[:10]\n",
- "\n",
- "for index in np.random.choice(len(y_test), 3, replace = False):\n",
- " print(index)\n",
- " test_sample = json.dumps({'data':X_test[index:index + 1].tolist()})\n",
- " predicted = aci_service.run(input_data = test_sample)\n",
- " label = y_test[index]\n",
- " predictedDict = json.loads(predicted)\n",
- " title = \"Label value = %d Predicted value = %s \" % ( label,predictedDict['result'][0])\n",
- " fig = plt.figure(1, figsize = (3,3))\n",
- " ax1 = fig.add_axes((0,0,.8,.8))\n",
- " ax1.set_title(title)\n",
- " plt.imshow(images[index], cmap = plt.cm.gray_r, interpolation = 'nearest')\n",
- " plt.show()"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "savitam"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Automated Machine Learning\n",
+ "_**Classification with Deployment**_\n",
+ "\n",
+ "## Contents\n",
+ "1. [Introduction](#Introduction)\n",
+ "1. [Setup](#Setup)\n",
+ "1. [Train](#Train)\n",
+ "1. [Deploy](#Deploy)\n",
+ "1. [Test](#Test)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Introduction\n",
+ "\n",
+ "In this example we use the scikit learn's [digit dataset](http://scikit-learn.org/stable/modules/generated/sklearn.datasets.load_digits.html) to showcase how you can use AutoML for a simple classification problem and deploy it to an Azure Container Instance (ACI).\n",
+ "\n",
+ "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
+ "\n",
+ "In this notebook you will learn how to:\n",
+ "1. Create an experiment using an existing workspace.\n",
+ "2. Configure AutoML using `AutoMLConfig`.\n",
+ "3. Train the model using local compute.\n",
+ "4. Explore the results.\n",
+ "5. Register the model.\n",
+ "6. Create a container image.\n",
+ "7. Create an Azure Container Instance (ACI) service.\n",
+ "8. Test the ACI service."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Setup\n",
+ "\n",
+ "As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import json\n",
+ "import logging\n",
+ "\n",
+ "from matplotlib import pyplot as plt\n",
+ "import numpy as np\n",
+ "import pandas as pd\n",
+ "from sklearn import datasets\n",
+ "\n",
+ "import azureml.core\n",
+ "from azureml.core.experiment import Experiment\n",
+ "from azureml.core.workspace import Workspace\n",
+ "from azureml.train.automl import AutoMLConfig\n",
+ "from azureml.train.automl.run import AutoMLRun"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "ws = Workspace.from_config()\n",
+ "\n",
+ "# choose a name for experiment\n",
+ "experiment_name = 'automl-local-classification'\n",
+ "# project folder\n",
+ "project_folder = './sample_projects/automl-local-classification'\n",
+ "\n",
+ "experiment=Experiment(ws, experiment_name)\n",
+ "\n",
+ "output = {}\n",
+ "output['SDK version'] = azureml.core.VERSION\n",
+ "output['Subscription ID'] = ws.subscription_id\n",
+ "output['Workspace'] = ws.name\n",
+ "output['Resource Group'] = ws.resource_group\n",
+ "output['Location'] = ws.location\n",
+ "output['Project Directory'] = project_folder\n",
+ "output['Experiment Name'] = experiment.name\n",
+ "pd.set_option('display.max_colwidth', -1)\n",
+ "outputDf = pd.DataFrame(data = output, index = [''])\n",
+ "outputDf.T"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Opt-in diagnostics for better experience, quality, and security of future releases."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.telemetry import set_diagnostics_collection\n",
+ "set_diagnostics_collection(send_diagnostics = True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Train\n",
+ "\n",
+ "Instantiate a AutoMLConfig object. This defines the settings and data used to run the experiment.\n",
+ "\n",
+ "|Property|Description|\n",
+ "|-|-|\n",
+ "|**task**|classification or regression|\n",
+ "|**primary_metric**|This is the metric that you want to optimize. Classification supports the following primary metrics:
accuracy
AUC_weighted
average_precision_score_weighted
norm_macro_recall
precision_score_weighted|\n",
+ "|**iteration_timeout_minutes**|Time limit in minutes for each iteration.|\n",
+ "|**iterations**|Number of iterations. In each iteration AutoML trains a specific pipeline with the data.|\n",
+ "|**n_cross_validations**|Number of cross validation splits.|\n",
+ "|**X**|(sparse) array-like, shape = [n_samples, n_features]|\n",
+ "|**y**|(sparse) array-like, shape = [n_samples, ], [n_samples, n_classes]
Multi-class targets. An indicator matrix turns on multilabel classification. This should be an array of integers.|\n",
+ "|**path**|Relative path to the project folder. AutoML stores configuration files for the experiment under this folder. You can specify a new empty folder.|"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "digits = datasets.load_digits()\n",
+ "X_train = digits.data[10:,:]\n",
+ "y_train = digits.target[10:]\n",
+ "\n",
+ "automl_config = AutoMLConfig(task = 'classification',\n",
+ " name = experiment_name,\n",
+ " debug_log = 'automl_errors.log',\n",
+ " primary_metric = 'AUC_weighted',\n",
+ " iteration_timeout_minutes = 20,\n",
+ " iterations = 10,\n",
+ " n_cross_validations = 2,\n",
+ " verbosity = logging.INFO,\n",
+ " X = X_train, \n",
+ " y = y_train,\n",
+ " path = project_folder)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Call the `submit` method on the experiment object and pass the run configuration. Execution of local runs is synchronous. Depending on the data and the number of iterations this can run for a while.\n",
+ "In this example, we specify `show_output = True` to print currently running iterations to the console."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "local_run = experiment.submit(automl_config, show_output = True)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "local_run"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Deploy\n",
+ "\n",
+ "### Retrieve the Best Model\n",
+ "\n",
+ "Below we select the best pipeline from our iterations. The `get_output` method on `automl_classifier` returns the best run and the fitted model for the last invocation. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "best_run, fitted_model = local_run.get_output()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Register the Fitted Model for Deployment\n",
+ "If neither `metric` nor `iteration` are specified in the `register_model` call, the iteration with the best primary metric is registered."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "description = 'AutoML Model'\n",
+ "tags = None\n",
+ "model = local_run.register_model(description = description, tags = tags)\n",
+ "\n",
+ "print(local_run.model_id) # This will be written to the script file later in the notebook."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create Scoring Script"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%writefile score.py\n",
+ "import pickle\n",
+ "import json\n",
+ "import numpy\n",
+ "import azureml.train.automl\n",
+ "from sklearn.externals import joblib\n",
+ "from azureml.core.model import Model\n",
+ "\n",
+ "\n",
+ "def init():\n",
+ " global model\n",
+ " model_path = Model.get_model_path(model_name = '<>') # this name is model.id of model that we want to deploy\n",
+ " # deserialize the model file back into a sklearn model\n",
+ " model = joblib.load(model_path)\n",
+ "\n",
+ "def run(rawdata):\n",
+ " try:\n",
+ " data = json.loads(rawdata)['data']\n",
+ " data = numpy.array(data)\n",
+ " result = model.predict(data)\n",
+ " except Exception as e:\n",
+ " result = str(e)\n",
+ " return json.dumps({\"error\": result})\n",
+ " return json.dumps({\"result\":result.tolist()})"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create a YAML File for the Environment"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "To ensure the fit results are consistent with the training results, the SDK dependency versions need to be the same as the environment that trains the model. Details about retrieving the versions can be found in notebook [12.auto-ml-retrieve-the-training-sdk-versions](12.auto-ml-retrieve-the-training-sdk-versions.ipynb)."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "experiment_name = 'automl-local-classification'\n",
+ "\n",
+ "experiment = Experiment(ws, experiment_name)\n",
+ "ml_run = AutoMLRun(experiment = experiment, run_id = local_run.id)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "dependencies = ml_run.get_run_sdk_dependencies(iteration = 7)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "for p in ['azureml-train-automl', 'azureml-sdk', 'azureml-core']:\n",
+ " print('{}\\t{}'.format(p, dependencies[p]))"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.conda_dependencies import CondaDependencies\n",
+ "\n",
+ "myenv = CondaDependencies.create(conda_packages=['numpy','scikit-learn'], pip_packages=['azureml-sdk[automl]'])\n",
+ "\n",
+ "conda_env_file_name = 'myenv.yml'\n",
+ "myenv.save_to_file('.', conda_env_file_name)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Substitute the actual version number in the environment file.\n",
+ "# This is not strictly needed in this notebook because the model should have been generated using the current SDK version.\n",
+ "# However, we include this in case this code is used on an experiment from a previous SDK version.\n",
+ "\n",
+ "with open(conda_env_file_name, 'r') as cefr:\n",
+ " content = cefr.read()\n",
+ "\n",
+ "with open(conda_env_file_name, 'w') as cefw:\n",
+ " cefw.write(content.replace(azureml.core.VERSION, dependencies['azureml-sdk']))\n",
+ "\n",
+ "# Substitute the actual model id in the script file.\n",
+ "\n",
+ "script_file_name = 'score.py'\n",
+ "\n",
+ "with open(script_file_name, 'r') as cefr:\n",
+ " content = cefr.read()\n",
+ "\n",
+ "with open(script_file_name, 'w') as cefw:\n",
+ " cefw.write(content.replace('<>', local_run.model_id))"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create a Container Image"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.image import Image, ContainerImage\n",
+ "\n",
+ "image_config = ContainerImage.image_configuration(runtime= \"python\",\n",
+ " execution_script = script_file_name,\n",
+ " conda_file = conda_env_file_name,\n",
+ " tags = {'area': \"digits\", 'type': \"automl_classification\"},\n",
+ " description = \"Image for automl classification sample\")\n",
+ "\n",
+ "image = Image.create(name = \"automlsampleimage\",\n",
+ " # this is the model object \n",
+ " models = [model],\n",
+ " image_config = image_config, \n",
+ " workspace = ws)\n",
+ "\n",
+ "image.wait_for_creation(show_output = True)\n",
+ "\n",
+ "if image.creation_state == 'Failed':\n",
+ " print(\"Image build log at: \" + image.image_build_log_uri)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Deploy the Image as a Web Service on Azure Container Instance"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.webservice import AciWebservice\n",
+ "\n",
+ "aciconfig = AciWebservice.deploy_configuration(cpu_cores = 1, \n",
+ " memory_gb = 1, \n",
+ " tags = {'area': \"digits\", 'type': \"automl_classification\"}, \n",
+ " description = 'sample service for Automl Classification')"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.webservice import Webservice\n",
+ "\n",
+ "aci_service_name = 'automl-sample-01'\n",
+ "print(aci_service_name)\n",
+ "aci_service = Webservice.deploy_from_image(deployment_config = aciconfig,\n",
+ " image = image,\n",
+ " name = aci_service_name,\n",
+ " workspace = ws)\n",
+ "aci_service.wait_for_deployment(True)\n",
+ "print(aci_service.state)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Delete a Web Service"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "#aci_service.delete()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Get Logs from a Deployed Web Service"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "#aci_service.get_logs()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Test"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "#Randomly select digits and test\n",
+ "digits = datasets.load_digits()\n",
+ "X_test = digits.data[:10, :]\n",
+ "y_test = digits.target[:10]\n",
+ "images = digits.images[:10]\n",
+ "\n",
+ "for index in np.random.choice(len(y_test), 3, replace = False):\n",
+ " print(index)\n",
+ " test_sample = json.dumps({'data':X_test[index:index + 1].tolist()})\n",
+ " predicted = aci_service.run(input_data = test_sample)\n",
+ " label = y_test[index]\n",
+ " predictedDict = json.loads(predicted)\n",
+ " title = \"Label value = %d Predicted value = %s \" % ( label,predictedDict['result'][0])\n",
+ " fig = plt.figure(1, figsize = (3,3))\n",
+ " ax1 = fig.add_axes((0,0,.8,.8))\n",
+ " ax1.set_title(title)\n",
+ " plt.imshow(images[index], cmap = plt.cm.gray_r, interpolation = 'nearest')\n",
+ " plt.show()"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "savitam"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.6"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/automated-machine-learning/classification-with-whitelisting/auto-ml-classification-with-whitelisting.ipynb b/how-to-use-azureml/automated-machine-learning/classification-with-whitelisting/auto-ml-classification-with-whitelisting.ipynb
index d32a7965..840d965b 100644
--- a/how-to-use-azureml/automated-machine-learning/classification-with-whitelisting/auto-ml-classification-with-whitelisting.ipynb
+++ b/how-to-use-azureml/automated-machine-learning/classification-with-whitelisting/auto-ml-classification-with-whitelisting.ipynb
@@ -1,403 +1,398 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Automated Machine Learning\n",
- "_**Classification using whitelist models**_\n",
- "\n",
- "## Contents\n",
- "1. [Introduction](#Introduction)\n",
- "1. [Setup](#Setup)\n",
- "1. [Data](#Data)\n",
- "1. [Train](#Train)\n",
- "1. [Results](#Results)\n",
- "1. [Test](#Test)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Introduction\n",
- "\n",
- "In this example we use the scikit-learn's [digit dataset](http://scikit-learn.org/stable/datasets/index.html#optical-recognition-of-handwritten-digits-dataset) to showcase how you can use AutoML for a simple classification problem.\n",
- "\n",
- "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
- "This notebooks shows how can automl can be trained on a a selected list of models,see the readme.md for the models.\n",
- "This trains the model exclusively on tensorflow based models.\n",
- "\n",
- "In this notebook you will learn how to:\n",
- "1. Create an `Experiment` in an existing `Workspace`.\n",
- "2. Configure AutoML using `AutoMLConfig`.\n",
- "3. Train the model on a whilelisted models using local compute. \n",
- "4. Explore the results.\n",
- "5. Test the best fitted model."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Setup\n",
- "\n",
- "As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import logging\n",
- "import os\n",
- "import random\n",
- "\n",
- "from matplotlib import pyplot as plt\n",
- "from matplotlib.pyplot import imshow\n",
- "import numpy as np\n",
- "import pandas as pd\n",
- "from sklearn import datasets\n",
- "\n",
- "import azureml.core\n",
- "from azureml.core.experiment import Experiment\n",
- "from azureml.core.workspace import Workspace\n",
- "from azureml.train.automl import AutoMLConfig\n",
- "from azureml.train.automl.run import AutoMLRun"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "ws = Workspace.from_config()\n",
- "\n",
- "# Choose a name for the experiment and specify the project folder.\n",
- "experiment_name = 'automl-local-whitelist'\n",
- "project_folder = './sample_projects/automl-local-whitelist'\n",
- "\n",
- "experiment = Experiment(ws, experiment_name)\n",
- "\n",
- "output = {}\n",
- "output['SDK version'] = azureml.core.VERSION\n",
- "output['Subscription ID'] = ws.subscription_id\n",
- "output['Workspace Name'] = ws.name\n",
- "output['Resource Group'] = ws.resource_group\n",
- "output['Location'] = ws.location\n",
- "output['Project Directory'] = project_folder\n",
- "output['Experiment Name'] = experiment.name\n",
- "pd.set_option('display.max_colwidth', -1)\n",
- "pd.DataFrame(data = output, index = ['']).T"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Opt-in diagnostics for better experience, quality, and security of future releases."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.telemetry import set_diagnostics_collection\n",
- "set_diagnostics_collection(send_diagnostics = True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Data\n",
- "\n",
- "This uses scikit-learn's [load_digits](http://scikit-learn.org/stable/modules/generated/sklearn.datasets.load_digits.html) method."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from sklearn import datasets\n",
- "\n",
- "digits = datasets.load_digits()\n",
- "\n",
- "# Exclude the first 100 rows from training so that they can be used for test.\n",
- "X_train = digits.data[100:,:]\n",
- "y_train = digits.target[100:]"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Train\n",
- "\n",
- "Instantiate an `AutoMLConfig` object to specify the settings and data used to run the experiment.\n",
- "\n",
- "|Property|Description|\n",
- "|-|-|\n",
- "|**task**|classification or regression|\n",
- "|**primary_metric**|This is the metric that you want to optimize. Classification supports the following primary metrics:
accuracy
AUC_weighted
balanced_accuracy
average_precision_score_weighted
precision_score_weighted|\n",
- "|**iteration_timeout_minutes**|Time limit in minutes for each iteration.|\n",
- "|**iterations**|Number of iterations. In each iteration AutoML trains a specific pipeline with the data.|\n",
- "|**n_cross_validations**|Number of cross validation splits.|\n",
- "|**X**|(sparse) array-like, shape = [n_samples, n_features]|\n",
- "|**y**|(sparse) array-like, shape = [n_samples, ], [n_samples, n_classes]
Multi-class targets. An indicator matrix turns on multilabel classification. This should be an array of integers.|\n",
- "|**path**|Relative path to the project folder. AutoML stores configuration files for the experiment under this folder. You can specify a new empty folder.|\n",
- "|**whitelist_models**|List of models that AutoML should use. The possible values are listed [here](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-configure-auto-train#configure-your-experiment-settings).|"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "automl_config = AutoMLConfig(task = 'classification',\n",
- " debug_log = 'automl_errors.log',\n",
- " primary_metric = 'AUC_weighted',\n",
- " iteration_timeout_minutes = 60,\n",
- " iterations = 10,\n",
- " n_cross_validations = 3,\n",
- " verbosity = logging.INFO,\n",
- " X = X_train, \n",
- " y = y_train,\n",
- " enable_tf=True,\n",
- " whitelist_models=[\"TensorFlowLinearClassifier\", \"TensorFlowDNN\"],\n",
- " path = project_folder)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Call the `submit` method on the experiment object and pass the run configuration. Execution of local runs is synchronous. Depending on the data and the number of iterations this can run for a while.\n",
- "In this example, we specify `show_output = True` to print currently running iterations to the console."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "local_run = experiment.submit(automl_config, show_output = True)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "local_run"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Results"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Widget for Monitoring Runs\n",
- "\n",
- "The widget will first report a \"loading\" status while running the first iteration. After completing the first iteration, an auto-updating graph and table will be shown. The widget will refresh once per minute, so you should see the graph update as child runs complete.\n",
- "\n",
- "**Note:** The widget displays a link at the bottom. Use this link to open a web interface to explore the individual run details."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.widgets import RunDetails\n",
- "RunDetails(local_run).show() "
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "\n",
- "#### Retrieve All Child Runs\n",
- "You can also use SDK methods to fetch all the child runs and see individual metrics that we log."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "children = list(local_run.get_children())\n",
- "metricslist = {}\n",
- "for run in children:\n",
- " properties = run.get_properties()\n",
- " metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)}\n",
- " metricslist[int(properties['iteration'])] = metrics\n",
- "\n",
- "rundata = pd.DataFrame(metricslist).sort_index(1)\n",
- "rundata"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Retrieve the Best Model\n",
- "\n",
- "Below we select the best pipeline from our iterations. The `get_output` method returns the best run and the fitted model. The Model includes the pipeline and any pre-processing. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "best_run, fitted_model = local_run.get_output()\n",
- "print(best_run)\n",
- "print(fitted_model)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Best Model Based on Any Other Metric\n",
- "Show the run and the model that has the smallest `log_loss` value:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "lookup_metric = \"log_loss\"\n",
- "best_run, fitted_model = local_run.get_output(metric = lookup_metric)\n",
- "print(best_run)\n",
- "print(fitted_model)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Model from a Specific Iteration\n",
- "Show the run and the model from the third iteration:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "iteration = 3\n",
- "third_run, third_model = local_run.get_output(iteration = iteration)\n",
- "print(third_run)\n",
- "print(third_model)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Test\n",
- "\n",
- "#### Load Test Data"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "digits = datasets.load_digits()\n",
- "X_test = digits.data[:10, :]\n",
- "y_test = digits.target[:10]\n",
- "images = digits.images[:10]"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Testing Our Best Fitted Model\n",
- "We will try to predict 2 digits and see how our model works."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Randomly select digits and test.\n",
- "for index in np.random.choice(len(y_test), 2, replace = False):\n",
- " print(index)\n",
- " predicted = fitted_model.predict(X_test[index:index + 1])[0]\n",
- " label = y_test[index]\n",
- " title = \"Label value = %d Predicted value = %d \" % (label, predicted)\n",
- " fig = plt.figure(1, figsize = (3,3))\n",
- " ax1 = fig.add_axes((0,0,.8,.8))\n",
- " ax1.set_title(title)\n",
- " plt.imshow(images[index], cmap = plt.cm.gray_r, interpolation = 'nearest')\n",
- " plt.show()"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "savitam"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Automated Machine Learning\n",
+ "_**Classification using whitelist models**_\n",
+ "\n",
+ "## Contents\n",
+ "1. [Introduction](#Introduction)\n",
+ "1. [Setup](#Setup)\n",
+ "1. [Data](#Data)\n",
+ "1. [Train](#Train)\n",
+ "1. [Results](#Results)\n",
+ "1. [Test](#Test)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Introduction\n",
+ "\n",
+ "In this example we use the scikit-learn's [digit dataset](http://scikit-learn.org/stable/datasets/index.html#optical-recognition-of-handwritten-digits-dataset) to showcase how you can use AutoML for a simple classification problem.\n",
+ "\n",
+ "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
+ "This notebooks shows how can automl can be trained on a a selected list of models,see the readme.md for the models.\n",
+ "This trains the model exclusively on tensorflow based models.\n",
+ "\n",
+ "In this notebook you will learn how to:\n",
+ "1. Create an `Experiment` in an existing `Workspace`.\n",
+ "2. Configure AutoML using `AutoMLConfig`.\n",
+ "3. Train the model on a whilelisted models using local compute. \n",
+ "4. Explore the results.\n",
+ "5. Test the best fitted model."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Setup\n",
+ "\n",
+ "As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import logging\n",
+ "\n",
+ "from matplotlib import pyplot as plt\n",
+ "import numpy as np\n",
+ "import pandas as pd\n",
+ "from sklearn import datasets\n",
+ "\n",
+ "import azureml.core\n",
+ "from azureml.core.experiment import Experiment\n",
+ "from azureml.core.workspace import Workspace\n",
+ "from azureml.train.automl import AutoMLConfig"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "ws = Workspace.from_config()\n",
+ "\n",
+ "# Choose a name for the experiment and specify the project folder.\n",
+ "experiment_name = 'automl-local-whitelist'\n",
+ "project_folder = './sample_projects/automl-local-whitelist'\n",
+ "\n",
+ "experiment = Experiment(ws, experiment_name)\n",
+ "\n",
+ "output = {}\n",
+ "output['SDK version'] = azureml.core.VERSION\n",
+ "output['Subscription ID'] = ws.subscription_id\n",
+ "output['Workspace Name'] = ws.name\n",
+ "output['Resource Group'] = ws.resource_group\n",
+ "output['Location'] = ws.location\n",
+ "output['Project Directory'] = project_folder\n",
+ "output['Experiment Name'] = experiment.name\n",
+ "pd.set_option('display.max_colwidth', -1)\n",
+ "outputDf = pd.DataFrame(data = output, index = [''])\n",
+ "outputDf.T"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Opt-in diagnostics for better experience, quality, and security of future releases."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.telemetry import set_diagnostics_collection\n",
+ "set_diagnostics_collection(send_diagnostics = True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Data\n",
+ "\n",
+ "This uses scikit-learn's [load_digits](http://scikit-learn.org/stable/modules/generated/sklearn.datasets.load_digits.html) method."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "digits = datasets.load_digits()\n",
+ "\n",
+ "# Exclude the first 100 rows from training so that they can be used for test.\n",
+ "X_train = digits.data[100:,:]\n",
+ "y_train = digits.target[100:]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Train\n",
+ "\n",
+ "Instantiate an `AutoMLConfig` object to specify the settings and data used to run the experiment.\n",
+ "\n",
+ "|Property|Description|\n",
+ "|-|-|\n",
+ "|**task**|classification or regression|\n",
+ "|**primary_metric**|This is the metric that you want to optimize. Classification supports the following primary metrics:
accuracy
AUC_weighted
balanced_accuracy
average_precision_score_weighted
precision_score_weighted|\n",
+ "|**iteration_timeout_minutes**|Time limit in minutes for each iteration.|\n",
+ "|**iterations**|Number of iterations. In each iteration AutoML trains a specific pipeline with the data.|\n",
+ "|**n_cross_validations**|Number of cross validation splits.|\n",
+ "|**X**|(sparse) array-like, shape = [n_samples, n_features]|\n",
+ "|**y**|(sparse) array-like, shape = [n_samples, ], [n_samples, n_classes]
Multi-class targets. An indicator matrix turns on multilabel classification. This should be an array of integers.|\n",
+ "|**path**|Relative path to the project folder. AutoML stores configuration files for the experiment under this folder. You can specify a new empty folder.|\n",
+ "|**whitelist_models**|List of models that AutoML should use. The possible values are listed [here](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-configure-auto-train#configure-your-experiment-settings).|"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "automl_config = AutoMLConfig(task = 'classification',\n",
+ " debug_log = 'automl_errors.log',\n",
+ " primary_metric = 'AUC_weighted',\n",
+ " iteration_timeout_minutes = 60,\n",
+ " iterations = 10,\n",
+ " n_cross_validations = 3,\n",
+ " verbosity = logging.INFO,\n",
+ " X = X_train, \n",
+ " y = y_train,\n",
+ " enable_tf=True,\n",
+ " whitelist_models=[\"TensorFlowLinearClassifier\", \"TensorFlowDNN\"],\n",
+ " path = project_folder)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Call the `submit` method on the experiment object and pass the run configuration. Execution of local runs is synchronous. Depending on the data and the number of iterations this can run for a while.\n",
+ "In this example, we specify `show_output = True` to print currently running iterations to the console."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "local_run = experiment.submit(automl_config, show_output = True)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "local_run"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Results"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Widget for Monitoring Runs\n",
+ "\n",
+ "The widget will first report a \"loading\" status while running the first iteration. After completing the first iteration, an auto-updating graph and table will be shown. The widget will refresh once per minute, so you should see the graph update as child runs complete.\n",
+ "\n",
+ "**Note:** The widget displays a link at the bottom. Use this link to open a web interface to explore the individual run details."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.widgets import RunDetails\n",
+ "RunDetails(local_run).show() "
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "\n",
+ "#### Retrieve All Child Runs\n",
+ "You can also use SDK methods to fetch all the child runs and see individual metrics that we log."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "children = list(local_run.get_children())\n",
+ "metricslist = {}\n",
+ "for run in children:\n",
+ " properties = run.get_properties()\n",
+ " metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)}\n",
+ " metricslist[int(properties['iteration'])] = metrics\n",
+ "\n",
+ "rundata = pd.DataFrame(metricslist).sort_index(1)\n",
+ "rundata"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Retrieve the Best Model\n",
+ "\n",
+ "Below we select the best pipeline from our iterations. The `get_output` method returns the best run and the fitted model. The Model includes the pipeline and any pre-processing. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "best_run, fitted_model = local_run.get_output()\n",
+ "print(best_run)\n",
+ "print(fitted_model)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Best Model Based on Any Other Metric\n",
+ "Show the run and the model that has the smallest `log_loss` value:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "lookup_metric = \"log_loss\"\n",
+ "best_run, fitted_model = local_run.get_output(metric = lookup_metric)\n",
+ "print(best_run)\n",
+ "print(fitted_model)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Model from a Specific Iteration\n",
+ "Show the run and the model from the third iteration:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "iteration = 3\n",
+ "third_run, third_model = local_run.get_output(iteration = iteration)\n",
+ "print(third_run)\n",
+ "print(third_model)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Test\n",
+ "\n",
+ "#### Load Test Data"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "digits = datasets.load_digits()\n",
+ "X_test = digits.data[:10, :]\n",
+ "y_test = digits.target[:10]\n",
+ "images = digits.images[:10]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Testing Our Best Fitted Model\n",
+ "We will try to predict 2 digits and see how our model works."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Randomly select digits and test.\n",
+ "for index in np.random.choice(len(y_test), 2, replace = False):\n",
+ " print(index)\n",
+ " predicted = fitted_model.predict(X_test[index:index + 1])[0]\n",
+ " label = y_test[index]\n",
+ " title = \"Label value = %d Predicted value = %d \" % (label, predicted)\n",
+ " fig = plt.figure(1, figsize = (3,3))\n",
+ " ax1 = fig.add_axes((0,0,.8,.8))\n",
+ " ax1.set_title(title)\n",
+ " plt.imshow(images[index], cmap = plt.cm.gray_r, interpolation = 'nearest')\n",
+ " plt.show()"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "savitam"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.6"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/automated-machine-learning/classification/auto-ml-classification.ipynb b/how-to-use-azureml/automated-machine-learning/classification/auto-ml-classification.ipynb
index 03ead4b0..f70e1faa 100644
--- a/how-to-use-azureml/automated-machine-learning/classification/auto-ml-classification.ipynb
+++ b/how-to-use-azureml/automated-machine-learning/classification/auto-ml-classification.ipynb
@@ -1,418 +1,413 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Automated Machine Learning\n",
- "_**Classification with Local Compute**_\n",
- "\n",
- "## Contents\n",
- "1. [Introduction](#Introduction)\n",
- "1. [Setup](#Setup)\n",
- "1. [Data](#Data)\n",
- "1. [Train](#Train)\n",
- "1. [Results](#Results)\n",
- "1. [Test](#Test)\n",
- "\n"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Introduction\n",
- "\n",
- "In this example we use the scikit-learn's [digit dataset](http://scikit-learn.org/stable/datasets/index.html#optical-recognition-of-handwritten-digits-dataset) to showcase how you can use AutoML for a simple classification problem.\n",
- "\n",
- "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
- "\n",
- "In this notebook you will learn how to:\n",
- "1. Create an `Experiment` in an existing `Workspace`.\n",
- "2. Configure AutoML using `AutoMLConfig`.\n",
- "3. Train the model using local compute.\n",
- "4. Explore the results.\n",
- "5. Test the best fitted model."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Setup\n",
- "\n",
- "As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import logging\n",
- "import os\n",
- "import random\n",
- "\n",
- "from matplotlib import pyplot as plt\n",
- "from matplotlib.pyplot import imshow\n",
- "import numpy as np\n",
- "import pandas as pd\n",
- "from sklearn import datasets\n",
- "\n",
- "import azureml.core\n",
- "from azureml.core.experiment import Experiment\n",
- "from azureml.core.workspace import Workspace\n",
- "from azureml.train.automl import AutoMLConfig\n",
- "from azureml.train.automl.run import AutoMLRun"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "ws = Workspace.from_config()\n",
- "\n",
- "# Choose a name for the experiment and specify the project folder.\n",
- "experiment_name = 'automl-local-classification'\n",
- "project_folder = './sample_projects/automl-local-classification'\n",
- "\n",
- "experiment = Experiment(ws, experiment_name)\n",
- "\n",
- "output = {}\n",
- "output['SDK version'] = azureml.core.VERSION\n",
- "output['Subscription ID'] = ws.subscription_id\n",
- "output['Workspace Name'] = ws.name\n",
- "output['Resource Group'] = ws.resource_group\n",
- "output['Location'] = ws.location\n",
- "output['Project Directory'] = project_folder\n",
- "output['Experiment Name'] = experiment.name\n",
- "pd.set_option('display.max_colwidth', -1)\n",
- "pd.DataFrame(data = output, index = ['']).T"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Opt-in diagnostics for better experience, quality, and security of future releases."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.telemetry import set_diagnostics_collection\n",
- "set_diagnostics_collection(send_diagnostics = True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Data\n",
- "\n",
- "This uses scikit-learn's [load_digits](http://scikit-learn.org/stable/modules/generated/sklearn.datasets.load_digits.html) method."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from sklearn import datasets\n",
- "\n",
- "digits = datasets.load_digits()\n",
- "\n",
- "# Exclude the first 100 rows from training so that they can be used for test.\n",
- "X_train = digits.data[100:,:]\n",
- "y_train = digits.target[100:]"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Train\n",
- "\n",
- "Instantiate an `AutoMLConfig` object to specify the settings and data used to run the experiment.\n",
- "\n",
- "|Property|Description|\n",
- "|-|-|\n",
- "|**task**|classification or regression|\n",
- "|**primary_metric**|This is the metric that you want to optimize. Classification supports the following primary metrics:
accuracy
AUC_weighted
average_precision_score_weighted
norm_macro_recall
precision_score_weighted|\n",
- "|**iteration_timeout_minutes**|Time limit in minutes for each iteration.|\n",
- "|**iterations**|Number of iterations. In each iteration AutoML trains a specific pipeline with the data.|\n",
- "|**n_cross_validations**|Number of cross validation splits.|\n",
- "|**X**|(sparse) array-like, shape = [n_samples, n_features]|\n",
- "|**y**|(sparse) array-like, shape = [n_samples, ], [n_samples, n_classes]
Multi-class targets. An indicator matrix turns on multilabel classification. This should be an array of integers.|\n",
- "|**path**|Relative path to the project folder. AutoML stores configuration files for the experiment under this folder. You can specify a new empty folder.|"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "automl_config = AutoMLConfig(task = 'classification',\n",
- " debug_log = 'automl_errors.log',\n",
- " primary_metric = 'AUC_weighted',\n",
- " iteration_timeout_minutes = 60,\n",
- " iterations = 25,\n",
- " n_cross_validations = 3,\n",
- " verbosity = logging.INFO,\n",
- " X = X_train, \n",
- " y = y_train,\n",
- " path = project_folder)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Call the `submit` method on the experiment object and pass the run configuration. Execution of local runs is synchronous. Depending on the data and the number of iterations this can run for a while.\n",
- "In this example, we specify `show_output = True` to print currently running iterations to the console."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "local_run = experiment.submit(automl_config, show_output = True)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "local_run"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Optionally, you can continue an interrupted local run by calling `continue_experiment` without the `iterations` parameter, or run more iterations for a completed run by specifying the `iterations` parameter:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "local_run = local_run.continue_experiment(X = X_train, \n",
- " y = y_train, \n",
- " show_output = True,\n",
- " iterations = 5)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Results"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Widget for Monitoring Runs\n",
- "\n",
- "The widget will first report a \"loading\" status while running the first iteration. After completing the first iteration, an auto-updating graph and table will be shown. The widget will refresh once per minute, so you should see the graph update as child runs complete.\n",
- "\n",
- "**Note:** The widget displays a link at the bottom. Use this link to open a web interface to explore the individual run details."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.widgets import RunDetails\n",
- "RunDetails(local_run).show() "
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "\n",
- "#### Retrieve All Child Runs\n",
- "You can also use SDK methods to fetch all the child runs and see individual metrics that we log."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "children = list(local_run.get_children())\n",
- "metricslist = {}\n",
- "for run in children:\n",
- " properties = run.get_properties()\n",
- " metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)}\n",
- " metricslist[int(properties['iteration'])] = metrics\n",
- "\n",
- "rundata = pd.DataFrame(metricslist).sort_index(1)\n",
- "rundata"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Retrieve the Best Model\n",
- "\n",
- "Below we select the best pipeline from our iterations. The `get_output` method returns the best run and the fitted model. The Model includes the pipeline and any pre-processing. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "best_run, fitted_model = local_run.get_output()\n",
- "print(best_run)\n",
- "print(fitted_model)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Best Model Based on Any Other Metric\n",
- "Show the run and the model that has the smallest `log_loss` value:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "lookup_metric = \"log_loss\"\n",
- "best_run, fitted_model = local_run.get_output(metric = lookup_metric)\n",
- "print(best_run)\n",
- "print(fitted_model)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Model from a Specific Iteration\n",
- "Show the run and the model from the third iteration:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "iteration = 3\n",
- "third_run, third_model = local_run.get_output(iteration = iteration)\n",
- "print(third_run)\n",
- "print(third_model)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Test \n",
- "\n",
- "#### Load Test Data"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "digits = datasets.load_digits()\n",
- "X_test = digits.data[:10, :]\n",
- "y_test = digits.target[:10]\n",
- "images = digits.images[:10]"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Testing Our Best Fitted Model\n",
- "We will try to predict 2 digits and see how our model works."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Randomly select digits and test.\n",
- "for index in np.random.choice(len(y_test), 2, replace = False):\n",
- " print(index)\n",
- " predicted = fitted_model.predict(X_test[index:index + 1])[0]\n",
- " label = y_test[index]\n",
- " title = \"Label value = %d Predicted value = %d \" % (label, predicted)\n",
- " fig = plt.figure(1, figsize = (3,3))\n",
- " ax1 = fig.add_axes((0,0,.8,.8))\n",
- " ax1.set_title(title)\n",
- " plt.imshow(images[index], cmap = plt.cm.gray_r, interpolation = 'nearest')\n",
- " plt.show()"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "savitam"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Automated Machine Learning\n",
+ "_**Classification with Local Compute**_\n",
+ "\n",
+ "## Contents\n",
+ "1. [Introduction](#Introduction)\n",
+ "1. [Setup](#Setup)\n",
+ "1. [Data](#Data)\n",
+ "1. [Train](#Train)\n",
+ "1. [Results](#Results)\n",
+ "1. [Test](#Test)\n",
+ "\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Introduction\n",
+ "\n",
+ "In this example we use the scikit-learn's [digit dataset](http://scikit-learn.org/stable/datasets/index.html#optical-recognition-of-handwritten-digits-dataset) to showcase how you can use AutoML for a simple classification problem.\n",
+ "\n",
+ "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
+ "\n",
+ "In this notebook you will learn how to:\n",
+ "1. Create an `Experiment` in an existing `Workspace`.\n",
+ "2. Configure AutoML using `AutoMLConfig`.\n",
+ "3. Train the model using local compute.\n",
+ "4. Explore the results.\n",
+ "5. Test the best fitted model."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Setup\n",
+ "\n",
+ "As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import logging\n",
+ "\n",
+ "from matplotlib import pyplot as plt\n",
+ "import numpy as np\n",
+ "import pandas as pd\n",
+ "from sklearn import datasets\n",
+ "\n",
+ "import azureml.core\n",
+ "from azureml.core.experiment import Experiment\n",
+ "from azureml.core.workspace import Workspace\n",
+ "from azureml.train.automl import AutoMLConfig"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "ws = Workspace.from_config()\n",
+ "\n",
+ "# Choose a name for the experiment and specify the project folder.\n",
+ "experiment_name = 'automl-local-classification'\n",
+ "project_folder = './sample_projects/automl-local-classification'\n",
+ "\n",
+ "experiment = Experiment(ws, experiment_name)\n",
+ "\n",
+ "output = {}\n",
+ "output['SDK version'] = azureml.core.VERSION\n",
+ "output['Subscription ID'] = ws.subscription_id\n",
+ "output['Workspace Name'] = ws.name\n",
+ "output['Resource Group'] = ws.resource_group\n",
+ "output['Location'] = ws.location\n",
+ "output['Project Directory'] = project_folder\n",
+ "output['Experiment Name'] = experiment.name\n",
+ "pd.set_option('display.max_colwidth', -1)\n",
+ "outputDf = pd.DataFrame(data = output, index = [''])\n",
+ "outputDf.T"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Opt-in diagnostics for better experience, quality, and security of future releases."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.telemetry import set_diagnostics_collection\n",
+ "set_diagnostics_collection(send_diagnostics = True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Data\n",
+ "\n",
+ "This uses scikit-learn's [load_digits](http://scikit-learn.org/stable/modules/generated/sklearn.datasets.load_digits.html) method."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "digits = datasets.load_digits()\n",
+ "\n",
+ "# Exclude the first 100 rows from training so that they can be used for test.\n",
+ "X_train = digits.data[100:,:]\n",
+ "y_train = digits.target[100:]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Train\n",
+ "\n",
+ "Instantiate an `AutoMLConfig` object to specify the settings and data used to run the experiment.\n",
+ "\n",
+ "|Property|Description|\n",
+ "|-|-|\n",
+ "|**task**|classification or regression|\n",
+ "|**primary_metric**|This is the metric that you want to optimize. Classification supports the following primary metrics:
accuracy
AUC_weighted
average_precision_score_weighted
norm_macro_recall
precision_score_weighted|\n",
+ "|**iteration_timeout_minutes**|Time limit in minutes for each iteration.|\n",
+ "|**iterations**|Number of iterations. In each iteration AutoML trains a specific pipeline with the data.|\n",
+ "|**n_cross_validations**|Number of cross validation splits.|\n",
+ "|**X**|(sparse) array-like, shape = [n_samples, n_features]|\n",
+ "|**y**|(sparse) array-like, shape = [n_samples, ], [n_samples, n_classes]
Multi-class targets. An indicator matrix turns on multilabel classification. This should be an array of integers.|\n",
+ "|**path**|Relative path to the project folder. AutoML stores configuration files for the experiment under this folder. You can specify a new empty folder.|"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "automl_config = AutoMLConfig(task = 'classification',\n",
+ " debug_log = 'automl_errors.log',\n",
+ " primary_metric = 'AUC_weighted',\n",
+ " iteration_timeout_minutes = 60,\n",
+ " iterations = 25,\n",
+ " n_cross_validations = 3,\n",
+ " verbosity = logging.INFO,\n",
+ " X = X_train, \n",
+ " y = y_train,\n",
+ " path = project_folder)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Call the `submit` method on the experiment object and pass the run configuration. Execution of local runs is synchronous. Depending on the data and the number of iterations this can run for a while.\n",
+ "In this example, we specify `show_output = True` to print currently running iterations to the console."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "local_run = experiment.submit(automl_config, show_output = True)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "local_run"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Optionally, you can continue an interrupted local run by calling `continue_experiment` without the `iterations` parameter, or run more iterations for a completed run by specifying the `iterations` parameter:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "local_run = local_run.continue_experiment(X = X_train, \n",
+ " y = y_train, \n",
+ " show_output = True,\n",
+ " iterations = 5)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Results"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Widget for Monitoring Runs\n",
+ "\n",
+ "The widget will first report a \"loading\" status while running the first iteration. After completing the first iteration, an auto-updating graph and table will be shown. The widget will refresh once per minute, so you should see the graph update as child runs complete.\n",
+ "\n",
+ "**Note:** The widget displays a link at the bottom. Use this link to open a web interface to explore the individual run details."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.widgets import RunDetails\n",
+ "RunDetails(local_run).show() "
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "\n",
+ "#### Retrieve All Child Runs\n",
+ "You can also use SDK methods to fetch all the child runs and see individual metrics that we log."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "children = list(local_run.get_children())\n",
+ "metricslist = {}\n",
+ "for run in children:\n",
+ " properties = run.get_properties()\n",
+ " metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)}\n",
+ " metricslist[int(properties['iteration'])] = metrics\n",
+ "\n",
+ "rundata = pd.DataFrame(metricslist).sort_index(1)\n",
+ "rundata"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Retrieve the Best Model\n",
+ "\n",
+ "Below we select the best pipeline from our iterations. The `get_output` method returns the best run and the fitted model. The Model includes the pipeline and any pre-processing. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "best_run, fitted_model = local_run.get_output()\n",
+ "print(best_run)\n",
+ "print(fitted_model)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Best Model Based on Any Other Metric\n",
+ "Show the run and the model that has the smallest `log_loss` value:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "lookup_metric = \"log_loss\"\n",
+ "best_run, fitted_model = local_run.get_output(metric = lookup_metric)\n",
+ "print(best_run)\n",
+ "print(fitted_model)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Model from a Specific Iteration\n",
+ "Show the run and the model from the third iteration:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "iteration = 3\n",
+ "third_run, third_model = local_run.get_output(iteration = iteration)\n",
+ "print(third_run)\n",
+ "print(third_model)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Test \n",
+ "\n",
+ "#### Load Test Data"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "digits = datasets.load_digits()\n",
+ "X_test = digits.data[:10, :]\n",
+ "y_test = digits.target[:10]\n",
+ "images = digits.images[:10]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Testing Our Best Fitted Model\n",
+ "We will try to predict 2 digits and see how our model works."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Randomly select digits and test.\n",
+ "for index in np.random.choice(len(y_test), 2, replace = False):\n",
+ " print(index)\n",
+ " predicted = fitted_model.predict(X_test[index:index + 1])[0]\n",
+ " label = y_test[index]\n",
+ " title = \"Label value = %d Predicted value = %d \" % (label, predicted)\n",
+ " fig = plt.figure(1, figsize = (3,3))\n",
+ " ax1 = fig.add_axes((0,0,.8,.8))\n",
+ " ax1.set_title(title)\n",
+ " plt.imshow(images[index], cmap = plt.cm.gray_r, interpolation = 'nearest')\n",
+ " plt.show()"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "savitam"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.6"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/automated-machine-learning/configuration.ipynb b/how-to-use-azureml/automated-machine-learning/configuration.ipynb
deleted file mode 100644
index 3b38870d..00000000
--- a/how-to-use-azureml/automated-machine-learning/configuration.ipynb
+++ /dev/null
@@ -1,154 +0,0 @@
-{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Automated Machine Learning Configuration\n",
- "\n",
- "In this example you will create an Azure Machine Learning `Workspace` object and initialize your notebook directory to easily reload this object from a configuration file. Typically you will only need to run this once per notebook directory, and all other notebooks in this directory or any sub-directories will automatically use the settings you indicate here.\n"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Check the Azure ML Core SDK Version to Validate Your Installation"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import azureml.core\n",
- "\n",
- "print(\"SDK Version:\", azureml.core.VERSION)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Initialize an Azure ML Workspace\n",
- "### What is an Azure ML Workspace and Why Do I Need One?\n",
- "\n",
- "An Azure ML workspace is an Azure resource that organizes and coordinates the actions of many other Azure resources to assist in executing and sharing machine learning workflows. In particular, an Azure ML workspace coordinates storage, databases, and compute resources providing added functionality for machine learning experimentation, operationalization, and the monitoring of operationalized models.\n",
- "\n",
- "\n",
- "### What do I Need?\n",
- "\n",
- "To create or access an Azure ML workspace, you will need to import the Azure ML library and specify following information:\n",
- "* A name for your workspace. You can choose one.\n",
- "* Your subscription id. Use the `id` value from the `az account show` command output above.\n",
- "* The resource group name. The resource group organizes Azure resources and provides a default region for the resources in the group. The resource group will be created if it doesn't exist. Resource groups can be created and viewed in the [Azure portal](https://portal.azure.com)\n",
- "* Supported regions include `eastus2`, `eastus`,`westcentralus`, `southeastasia`, `westeurope`, `australiaeast`, `westus2`, `southcentralus`."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "subscription_id = \"\"\n",
- "resource_group = \"myrg\"\n",
- "workspace_name = \"myws\"\n",
- "workspace_region = \"eastus2\""
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Creating a Workspace\n",
- "If you already have access to an Azure ML workspace you want to use, you can skip this cell. Otherwise, this cell will create an Azure ML workspace for you in the specified subscription, provided you have the correct permissions for the given `subscription_id`.\n",
- "\n",
- "This will fail when:\n",
- "1. The workspace already exists.\n",
- "2. You do not have permission to create a workspace in the resource group.\n",
- "3. You are not a subscription owner or contributor and no Azure ML workspaces have ever been created in this subscription.\n",
- "\n",
- "If workspace creation fails for any reason other than already existing, please work with your IT administrator to provide you with the appropriate permissions or to provision the required resources.\n",
- "\n",
- "**Note:** Creation of a new workspace can take several minutes."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Import the Workspace class and check the Azure ML SDK version.\n",
- "from azureml.core import Workspace\n",
- "\n",
- "ws = Workspace.create(name = workspace_name,\n",
- " subscription_id = subscription_id,\n",
- " resource_group = resource_group, \n",
- " location = workspace_region)\n",
- "ws.get_details()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Configuring Your Local Environment\n",
- "You can validate that you have access to the specified workspace and write a configuration file to the default configuration location, `./aml_config/config.json`."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core import Workspace\n",
- "\n",
- "ws = Workspace(workspace_name = workspace_name,\n",
- " subscription_id = subscription_id,\n",
- " resource_group = resource_group)\n",
- "\n",
- "# Persist the subscription id, resource group name, and workspace name in aml_config/config.json.\n",
- "ws.write_config()"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "savitam"
- }
- ],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
- },
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
\ No newline at end of file
diff --git a/how-to-use-azureml/automated-machine-learning/dataprep-remote-execution/auto-ml-dataprep-remote-execution.ipynb b/how-to-use-azureml/automated-machine-learning/dataprep-remote-execution/auto-ml-dataprep-remote-execution.ipynb
index 75576e68..427a475e 100644
--- a/how-to-use-azureml/automated-machine-learning/dataprep-remote-execution/auto-ml-dataprep-remote-execution.ipynb
+++ b/how-to-use-azureml/automated-machine-learning/dataprep-remote-execution/auto-ml-dataprep-remote-execution.ipynb
@@ -1,518 +1,515 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Automated Machine Learning\n",
- "_**Prepare Data using `azureml.dataprep` for Remote Execution (DSVM)**_\n",
- "\n",
- "## Contents\n",
- "1. [Introduction](#Introduction)\n",
- "1. [Setup](#Setup)\n",
- "1. [Data](#Data)\n",
- "1. [Train](#Train)\n",
- "1. [Results](#Results)\n",
- "1. [Test](#Test)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Introduction\n",
- "In this example we showcase how you can use the `azureml.dataprep` SDK to load and prepare data for AutoML. `azureml.dataprep` can also be used standalone; full documentation can be found [here](https://github.com/Microsoft/PendletonDocs).\n",
- "\n",
- "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
- "\n",
- "In this notebook you will learn how to:\n",
- "1. Define data loading and preparation steps in a `Dataflow` using `azureml.dataprep`.\n",
- "2. Pass the `Dataflow` to AutoML for a local run.\n",
- "3. Pass the `Dataflow` to AutoML for a remote run."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Setup\n",
- "\n",
- "Currently, Data Prep only supports __Ubuntu 16__ and __Red Hat Enterprise Linux 7__. We are working on supporting more linux distros."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Opt-in diagnostics for better experience, quality, and security of future releases."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.telemetry import set_diagnostics_collection\n",
- "set_diagnostics_collection(send_diagnostics = True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import logging\n",
- "import os\n",
- "import time\n",
- "\n",
- "import pandas as pd\n",
- "\n",
- "import azureml.core\n",
- "from azureml.core.compute import DsvmCompute\n",
- "from azureml.core.experiment import Experiment\n",
- "from azureml.core.workspace import Workspace\n",
- "import azureml.dataprep as dprep\n",
- "from azureml.train.automl import AutoMLConfig"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "ws = Workspace.from_config()\n",
- " \n",
- "# choose a name for experiment\n",
- "experiment_name = 'automl-dataprep-remote-dsvm'\n",
- "# project folder\n",
- "project_folder = './sample_projects/automl-dataprep-remote-dsvm'\n",
- " \n",
- "experiment = Experiment(ws, experiment_name)\n",
- " \n",
- "output = {}\n",
- "output['SDK version'] = azureml.core.VERSION\n",
- "output['Subscription ID'] = ws.subscription_id\n",
- "output['Workspace Name'] = ws.name\n",
- "output['Resource Group'] = ws.resource_group\n",
- "output['Location'] = ws.location\n",
- "output['Project Directory'] = project_folder\n",
- "output['Experiment Name'] = experiment.name\n",
- "pd.set_option('display.max_colwidth', -1)\n",
- "pd.DataFrame(data = output, index = ['']).T"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Data"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# You can use `auto_read_file` which intelligently figures out delimiters and datatypes of a file.\n",
- "# The data referenced here was pulled from `sklearn.datasets.load_digits()`.\n",
- "simple_example_data_root = 'https://dprepdata.blob.core.windows.net/automl-notebook-data/'\n",
- "X = dprep.auto_read_file(simple_example_data_root + 'X.csv').skip(1) # Remove the header row.\n",
- "\n",
- "# You can also use `read_csv` and `to_*` transformations to read (with overridable delimiter)\n",
- "# and convert column types manually.\n",
- "# Here we read a comma delimited file and convert all columns to integers.\n",
- "y = dprep.read_csv(simple_example_data_root + 'y.csv').to_long(dprep.ColumnSelector(term='.*', use_regex = True))"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "You can peek the result of a Dataflow at any range using `skip(i)` and `head(j)`. Doing so evaluates only `j` records for all the steps in the Dataflow, which makes it fast even against large datasets."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "X.skip(1).head(5)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Train\n",
- "\n",
- "This creates a general AutoML settings object applicable for both local and remote runs."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "automl_settings = {\n",
- " \"iteration_timeout_minutes\" : 10,\n",
- " \"iterations\" : 2,\n",
- " \"primary_metric\" : 'AUC_weighted',\n",
- " \"preprocess\" : False,\n",
- " \"verbosity\" : logging.INFO,\n",
- " \"n_cross_validations\": 3\n",
- "}"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create or Attach a Remote Linux DSVM"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "dsvm_name = 'mydsvmc'\n",
- "\n",
- "try:\n",
- " while ws.compute_targets[dsvm_name].provisioning_state == 'Creating':\n",
- " time.sleep(1)\n",
- " \n",
- " dsvm_compute = DsvmCompute(ws, dsvm_name)\n",
- " print('Found existing DVSM.')\n",
- "except:\n",
- " print('Creating a new DSVM.')\n",
- " dsvm_config = DsvmCompute.provisioning_configuration(vm_size = \"Standard_D2_v2\")\n",
- " dsvm_compute = DsvmCompute.create(ws, name = dsvm_name, provisioning_configuration = dsvm_config)\n",
- " dsvm_compute.wait_for_completion(show_output = True)\n",
- " print(\"Waiting one minute for ssh to be accessible\")\n",
- " time.sleep(60) # Wait for ssh to be accessible"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.runconfig import RunConfiguration\n",
- "from azureml.core.conda_dependencies import CondaDependencies\n",
- "\n",
- "conda_run_config = RunConfiguration(framework=\"python\")\n",
- "\n",
- "conda_run_config.target = dsvm_compute\n",
- "\n",
- "cd = CondaDependencies.create(pip_packages=['azureml-sdk[automl]'], conda_packages=['numpy'])\n",
- "conda_run_config.environment.python.conda_dependencies = cd"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Pass Data with `Dataflow` Objects\n",
- "\n",
- "The `Dataflow` objects captured above can also be passed to the `submit` method for a remote run. AutoML will serialize the `Dataflow` object and send it to the remote compute target. The `Dataflow` will not be evaluated locally."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "automl_config = AutoMLConfig(task = 'classification',\n",
- " debug_log = 'automl_errors.log',\n",
- " path = project_folder,\n",
- " run_configuration=conda_run_config,\n",
- " X = X,\n",
- " y = y,\n",
- " **automl_settings)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "remote_run = experiment.submit(automl_config, show_output = True)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "remote_run"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Results"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Widget for Monitoring Runs\n",
- "\n",
- "The widget will first report a \"loading\" status while running the first iteration. After completing the first iteration, an auto-updating graph and table will be shown. The widget will refresh once per minute, so you should see the graph update as child runs complete.\n",
- "\n",
- "**Note:** The widget displays a link at the bottom. Use this link to open a web interface to explore the individual run details."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.widgets import RunDetails\n",
- "RunDetails(remote_run).show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Retrieve All Child Runs\n",
- "You can also use SDK methods to fetch all the child runs and see individual metrics that we log."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "children = list(remote_run.get_children())\n",
- "metricslist = {}\n",
- "for run in children:\n",
- " properties = run.get_properties()\n",
- " metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)}\n",
- " metricslist[int(properties['iteration'])] = metrics\n",
- " \n",
- "import pandas as pd\n",
- "rundata = pd.DataFrame(metricslist).sort_index(1)\n",
- "rundata"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Retrieve the Best Model\n",
- "\n",
- "Below we select the best pipeline from our iterations. The `get_output` method returns the best run and the fitted model. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "best_run, fitted_model = remote_run.get_output()\n",
- "print(best_run)\n",
- "print(fitted_model)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Best Model Based on Any Other Metric\n",
- "Show the run and the model that has the smallest `log_loss` value:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "lookup_metric = \"log_loss\"\n",
- "best_run, fitted_model = remote_run.get_output(metric = lookup_metric)\n",
- "print(best_run)\n",
- "print(fitted_model)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Model from a Specific Iteration\n",
- "Show the run and the model from the first iteration:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "iteration = 0\n",
- "best_run, fitted_model = remote_run.get_output(iteration = iteration)\n",
- "print(best_run)\n",
- "print(fitted_model)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Test\n",
- "\n",
- "#### Load Test Data"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from sklearn import datasets\n",
- "\n",
- "digits = datasets.load_digits()\n",
- "X_test = digits.data[:10, :]\n",
- "y_test = digits.target[:10]\n",
- "images = digits.images[:10]"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Testing Our Best Fitted Model\n",
- "We will try to predict 2 digits and see how our model works."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "#Randomly select digits and test\n",
- "from matplotlib import pyplot as plt\n",
- "from matplotlib.pyplot import imshow\n",
- "import random\n",
- "import numpy as np\n",
- "\n",
- "for index in np.random.choice(len(y_test), 2, replace = False):\n",
- " print(index)\n",
- " predicted = fitted_model.predict(X_test[index:index + 1])[0]\n",
- " label = y_test[index]\n",
- " title = \"Label value = %d Predicted value = %d \" % (label, predicted)\n",
- " fig = plt.figure(1, figsize=(3,3))\n",
- " ax1 = fig.add_axes((0,0,.8,.8))\n",
- " ax1.set_title(title)\n",
- " plt.imshow(images[index], cmap = plt.cm.gray_r, interpolation = 'nearest')\n",
- " plt.show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Appendix"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Capture the `Dataflow` Objects for Later Use in AutoML\n",
- "\n",
- "`Dataflow` objects are immutable and are composed of a list of data preparation steps. A `Dataflow` object can be branched at any point for further usage."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# sklearn.digits.data + target\n",
- "digits_complete = dprep.auto_read_file('https://dprepdata.blob.core.windows.net/automl-notebook-data/digits-complete.csv')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "`digits_complete` (sourced from `sklearn.datasets.load_digits()`) is forked into `dflow_X` to capture all the feature columns and `dflow_y` to capture the label column."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "digits_complete.to_pandas_dataframe().shape\n",
- "labels_column = 'Column64'\n",
- "dflow_X = digits_complete.drop_columns(columns = [labels_column])\n",
- "dflow_y = digits_complete.keep_columns(columns = [labels_column])"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "savitam"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Automated Machine Learning\n",
+ "_**Prepare Data using `azureml.dataprep` for Remote Execution (DSVM)**_\n",
+ "\n",
+ "## Contents\n",
+ "1. [Introduction](#Introduction)\n",
+ "1. [Setup](#Setup)\n",
+ "1. [Data](#Data)\n",
+ "1. [Train](#Train)\n",
+ "1. [Results](#Results)\n",
+ "1. [Test](#Test)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Introduction\n",
+ "In this example we showcase how you can use the `azureml.dataprep` SDK to load and prepare data for AutoML. `azureml.dataprep` can also be used standalone; full documentation can be found [here](https://github.com/Microsoft/PendletonDocs).\n",
+ "\n",
+ "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
+ "\n",
+ "In this notebook you will learn how to:\n",
+ "1. Define data loading and preparation steps in a `Dataflow` using `azureml.dataprep`.\n",
+ "2. Pass the `Dataflow` to AutoML for a local run.\n",
+ "3. Pass the `Dataflow` to AutoML for a remote run."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Setup\n",
+ "\n",
+ "Currently, Data Prep only supports __Ubuntu 16__ and __Red Hat Enterprise Linux 7__. We are working on supporting more linux distros."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Opt-in diagnostics for better experience, quality, and security of future releases."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.telemetry import set_diagnostics_collection\n",
+ "set_diagnostics_collection(send_diagnostics = True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import logging\n",
+ "import time\n",
+ "\n",
+ "import pandas as pd\n",
+ "\n",
+ "import azureml.core\n",
+ "from azureml.core.compute import DsvmCompute\n",
+ "from azureml.core.experiment import Experiment\n",
+ "from azureml.core.workspace import Workspace\n",
+ "import azureml.dataprep as dprep\n",
+ "from azureml.train.automl import AutoMLConfig"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "ws = Workspace.from_config()\n",
+ " \n",
+ "# choose a name for experiment\n",
+ "experiment_name = 'automl-dataprep-remote-dsvm'\n",
+ "# project folder\n",
+ "project_folder = './sample_projects/automl-dataprep-remote-dsvm'\n",
+ " \n",
+ "experiment = Experiment(ws, experiment_name)\n",
+ " \n",
+ "output = {}\n",
+ "output['SDK version'] = azureml.core.VERSION\n",
+ "output['Subscription ID'] = ws.subscription_id\n",
+ "output['Workspace Name'] = ws.name\n",
+ "output['Resource Group'] = ws.resource_group\n",
+ "output['Location'] = ws.location\n",
+ "output['Project Directory'] = project_folder\n",
+ "output['Experiment Name'] = experiment.name\n",
+ "pd.set_option('display.max_colwidth', -1)\n",
+ "outputDf = pd.DataFrame(data = output, index = [''])\n",
+ "outputDf.T"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Data"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# You can use `auto_read_file` which intelligently figures out delimiters and datatypes of a file.\n",
+ "# The data referenced here was pulled from `sklearn.datasets.load_digits()`.\n",
+ "simple_example_data_root = 'https://dprepdata.blob.core.windows.net/automl-notebook-data/'\n",
+ "X = dprep.auto_read_file(simple_example_data_root + 'X.csv').skip(1) # Remove the header row.\n",
+ "\n",
+ "# You can also use `read_csv` and `to_*` transformations to read (with overridable delimiter)\n",
+ "# and convert column types manually.\n",
+ "# Here we read a comma delimited file and convert all columns to integers.\n",
+ "y = dprep.read_csv(simple_example_data_root + 'y.csv').to_long(dprep.ColumnSelector(term='.*', use_regex = True))"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "You can peek the result of a Dataflow at any range using `skip(i)` and `head(j)`. Doing so evaluates only `j` records for all the steps in the Dataflow, which makes it fast even against large datasets."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "X.skip(1).head(5)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Train\n",
+ "\n",
+ "This creates a general AutoML settings object applicable for both local and remote runs."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "automl_settings = {\n",
+ " \"iteration_timeout_minutes\" : 10,\n",
+ " \"iterations\" : 2,\n",
+ " \"primary_metric\" : 'AUC_weighted',\n",
+ " \"preprocess\" : False,\n",
+ " \"verbosity\" : logging.INFO,\n",
+ " \"n_cross_validations\": 3\n",
+ "}"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create or Attach a Remote Linux DSVM"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "dsvm_name = 'mydsvmc'\n",
+ "\n",
+ "try:\n",
+ " while ws.compute_targets[dsvm_name].provisioning_state == 'Creating':\n",
+ " time.sleep(1)\n",
+ " \n",
+ " dsvm_compute = DsvmCompute(ws, dsvm_name)\n",
+ " print('Found existing DVSM.')\n",
+ "except:\n",
+ " print('Creating a new DSVM.')\n",
+ " dsvm_config = DsvmCompute.provisioning_configuration(vm_size = \"Standard_D2_v2\")\n",
+ " dsvm_compute = DsvmCompute.create(ws, name = dsvm_name, provisioning_configuration = dsvm_config)\n",
+ " dsvm_compute.wait_for_completion(show_output = True)\n",
+ " print(\"Waiting one minute for ssh to be accessible\")\n",
+ " time.sleep(60) # Wait for ssh to be accessible"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.runconfig import RunConfiguration\n",
+ "from azureml.core.conda_dependencies import CondaDependencies\n",
+ "\n",
+ "conda_run_config = RunConfiguration(framework=\"python\")\n",
+ "\n",
+ "conda_run_config.target = dsvm_compute\n",
+ "\n",
+ "cd = CondaDependencies.create(pip_packages=['azureml-sdk[automl]'], conda_packages=['numpy'])\n",
+ "conda_run_config.environment.python.conda_dependencies = cd"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Pass Data with `Dataflow` Objects\n",
+ "\n",
+ "The `Dataflow` objects captured above can also be passed to the `submit` method for a remote run. AutoML will serialize the `Dataflow` object and send it to the remote compute target. The `Dataflow` will not be evaluated locally."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "automl_config = AutoMLConfig(task = 'classification',\n",
+ " debug_log = 'automl_errors.log',\n",
+ " path = project_folder,\n",
+ " run_configuration=conda_run_config,\n",
+ " X = X,\n",
+ " y = y,\n",
+ " **automl_settings)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "remote_run = experiment.submit(automl_config, show_output = True)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "remote_run"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Results"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Widget for Monitoring Runs\n",
+ "\n",
+ "The widget will first report a \"loading\" status while running the first iteration. After completing the first iteration, an auto-updating graph and table will be shown. The widget will refresh once per minute, so you should see the graph update as child runs complete.\n",
+ "\n",
+ "**Note:** The widget displays a link at the bottom. Use this link to open a web interface to explore the individual run details."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.widgets import RunDetails\n",
+ "RunDetails(remote_run).show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Retrieve All Child Runs\n",
+ "You can also use SDK methods to fetch all the child runs and see individual metrics that we log."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "children = list(remote_run.get_children())\n",
+ "metricslist = {}\n",
+ "for run in children:\n",
+ " properties = run.get_properties()\n",
+ " metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)}\n",
+ " metricslist[int(properties['iteration'])] = metrics\n",
+ " \n",
+ "rundata = pd.DataFrame(metricslist).sort_index(1)\n",
+ "rundata"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Retrieve the Best Model\n",
+ "\n",
+ "Below we select the best pipeline from our iterations. The `get_output` method returns the best run and the fitted model. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "best_run, fitted_model = remote_run.get_output()\n",
+ "print(best_run)\n",
+ "print(fitted_model)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Best Model Based on Any Other Metric\n",
+ "Show the run and the model that has the smallest `log_loss` value:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "lookup_metric = \"log_loss\"\n",
+ "best_run, fitted_model = remote_run.get_output(metric = lookup_metric)\n",
+ "print(best_run)\n",
+ "print(fitted_model)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Model from a Specific Iteration\n",
+ "Show the run and the model from the first iteration:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "iteration = 0\n",
+ "best_run, fitted_model = remote_run.get_output(iteration = iteration)\n",
+ "print(best_run)\n",
+ "print(fitted_model)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Test\n",
+ "\n",
+ "#### Load Test Data"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from sklearn import datasets\n",
+ "\n",
+ "digits = datasets.load_digits()\n",
+ "X_test = digits.data[:10, :]\n",
+ "y_test = digits.target[:10]\n",
+ "images = digits.images[:10]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Testing Our Best Fitted Model\n",
+ "We will try to predict 2 digits and see how our model works."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "#Randomly select digits and test\n",
+ "from matplotlib import pyplot as plt\n",
+ "import numpy as np\n",
+ "\n",
+ "for index in np.random.choice(len(y_test), 2, replace = False):\n",
+ " print(index)\n",
+ " predicted = fitted_model.predict(X_test[index:index + 1])[0]\n",
+ " label = y_test[index]\n",
+ " title = \"Label value = %d Predicted value = %d \" % (label, predicted)\n",
+ " fig = plt.figure(1, figsize=(3,3))\n",
+ " ax1 = fig.add_axes((0,0,.8,.8))\n",
+ " ax1.set_title(title)\n",
+ " plt.imshow(images[index], cmap = plt.cm.gray_r, interpolation = 'nearest')\n",
+ " plt.show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Appendix"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Capture the `Dataflow` Objects for Later Use in AutoML\n",
+ "\n",
+ "`Dataflow` objects are immutable and are composed of a list of data preparation steps. A `Dataflow` object can be branched at any point for further usage."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# sklearn.digits.data + target\n",
+ "digits_complete = dprep.auto_read_file('https://dprepdata.blob.core.windows.net/automl-notebook-data/digits-complete.csv')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "`digits_complete` (sourced from `sklearn.datasets.load_digits()`) is forked into `dflow_X` to capture all the feature columns and `dflow_y` to capture the label column."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "print(digits_complete.to_pandas_dataframe().shape)\n",
+ "labels_column = 'Column64'\n",
+ "dflow_X = digits_complete.drop_columns(columns = [labels_column])\n",
+ "dflow_y = digits_complete.keep_columns(columns = [labels_column])"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "savitam"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.5"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.5"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/automated-machine-learning/dataprep/auto-ml-dataprep.ipynb b/how-to-use-azureml/automated-machine-learning/dataprep/auto-ml-dataprep.ipynb
index 8431e81c..c81cdbad 100644
--- a/how-to-use-azureml/automated-machine-learning/dataprep/auto-ml-dataprep.ipynb
+++ b/how-to-use-azureml/automated-machine-learning/dataprep/auto-ml-dataprep.ipynb
@@ -1,469 +1,466 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Automated Machine Learning\n",
- "_**Prepare Data using `azureml.dataprep` for Local Execution**_\n",
- "\n",
- "## Contents\n",
- "1. [Introduction](#Introduction)\n",
- "1. [Setup](#Setup)\n",
- "1. [Data](#Data)\n",
- "1. [Train](#Train)\n",
- "1. [Results](#Results)\n",
- "1. [Test](#Test)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Introduction\n",
- "In this example we showcase how you can use the `azureml.dataprep` SDK to load and prepare data for AutoML. `azureml.dataprep` can also be used standalone; full documentation can be found [here](https://github.com/Microsoft/PendletonDocs).\n",
- "\n",
- "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
- "\n",
- "In this notebook you will learn how to:\n",
- "1. Define data loading and preparation steps in a `Dataflow` using `azureml.dataprep`.\n",
- "2. Pass the `Dataflow` to AutoML for a local run.\n",
- "3. Pass the `Dataflow` to AutoML for a remote run."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Setup\n",
- "\n",
- "Currently, Data Prep only supports __Ubuntu 16__ and __Red Hat Enterprise Linux 7__. We are working on supporting more linux distros."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Opt-in diagnostics for better experience, quality, and security of future releases."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.telemetry import set_diagnostics_collection\n",
- "set_diagnostics_collection(send_diagnostics = True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import logging\n",
- "import os\n",
- "\n",
- "import pandas as pd\n",
- "\n",
- "import azureml.core\n",
- "from azureml.core.experiment import Experiment\n",
- "from azureml.core.workspace import Workspace\n",
- "import azureml.dataprep as dprep\n",
- "from azureml.train.automl import AutoMLConfig"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "ws = Workspace.from_config()\n",
- " \n",
- "# choose a name for experiment\n",
- "experiment_name = 'automl-dataprep-local'\n",
- "# project folder\n",
- "project_folder = './sample_projects/automl-dataprep-local'\n",
- " \n",
- "experiment = Experiment(ws, experiment_name)\n",
- " \n",
- "output = {}\n",
- "output['SDK version'] = azureml.core.VERSION\n",
- "output['Subscription ID'] = ws.subscription_id\n",
- "output['Workspace Name'] = ws.name\n",
- "output['Resource Group'] = ws.resource_group\n",
- "output['Location'] = ws.location\n",
- "output['Project Directory'] = project_folder\n",
- "output['Experiment Name'] = experiment.name\n",
- "pd.set_option('display.max_colwidth', -1)\n",
- "pd.DataFrame(data = output, index = ['']).T"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Data"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# You can use `auto_read_file` which intelligently figures out delimiters and datatypes of a file.\n",
- "# The data referenced here was pulled from `sklearn.datasets.load_digits()`.\n",
- "simple_example_data_root = 'https://dprepdata.blob.core.windows.net/automl-notebook-data/'\n",
- "X = dprep.auto_read_file(simple_example_data_root + 'X.csv').skip(1) # Remove the header row.\n",
- "\n",
- "# You can also use `read_csv` and `to_*` transformations to read (with overridable delimiter)\n",
- "# and convert column types manually.\n",
- "# Here we read a comma delimited file and convert all columns to integers.\n",
- "y = dprep.read_csv(simple_example_data_root + 'y.csv').to_long(dprep.ColumnSelector(term='.*', use_regex = True))"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Review the Data Preparation Result\n",
- "\n",
- "You can peek the result of a Dataflow at any range using `skip(i)` and `head(j)`. Doing so evaluates only `j` records for all the steps in the Dataflow, which makes it fast even against large datasets."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "X.skip(1).head(5)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Train\n",
- "\n",
- "This creates a general AutoML settings object applicable for both local and remote runs."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "automl_settings = {\n",
- " \"iteration_timeout_minutes\" : 10,\n",
- " \"iterations\" : 2,\n",
- " \"primary_metric\" : 'AUC_weighted',\n",
- " \"preprocess\" : False,\n",
- " \"verbosity\" : logging.INFO,\n",
- " \"n_cross_validations\": 3\n",
- "}"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Pass Data with `Dataflow` Objects\n",
- "\n",
- "The `Dataflow` objects captured above can be passed to the `submit` method for a local run. AutoML will retrieve the results from the `Dataflow` for model training."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "automl_config = AutoMLConfig(task = 'classification',\n",
- " debug_log = 'automl_errors.log',\n",
- " X = X,\n",
- " y = y,\n",
- " **automl_settings)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "local_run = experiment.submit(automl_config, show_output = True)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "local_run"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Results"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Widget for Monitoring Runs\n",
- "\n",
- "The widget will first report a \"loading\" status while running the first iteration. After completing the first iteration, an auto-updating graph and table will be shown. The widget will refresh once per minute, so you should see the graph update as child runs complete.\n",
- "\n",
- "**Note:** The widget displays a link at the bottom. Use this link to open a web interface to explore the individual run details."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.widgets import RunDetails\n",
- "RunDetails(local_run).show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Retrieve All Child Runs\n",
- "You can also use SDK methods to fetch all the child runs and see individual metrics that we log."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "children = list(local_run.get_children())\n",
- "metricslist = {}\n",
- "for run in children:\n",
- " properties = run.get_properties()\n",
- " metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)}\n",
- " metricslist[int(properties['iteration'])] = metrics\n",
- " \n",
- "import pandas as pd\n",
- "rundata = pd.DataFrame(metricslist).sort_index(1)\n",
- "rundata"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Retrieve the Best Model\n",
- "\n",
- "Below we select the best pipeline from our iterations. The `get_output` method returns the best run and the fitted model. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "best_run, fitted_model = local_run.get_output()\n",
- "print(best_run)\n",
- "print(fitted_model)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Best Model Based on Any Other Metric\n",
- "Show the run and the model that has the smallest `log_loss` value:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "lookup_metric = \"log_loss\"\n",
- "best_run, fitted_model = local_run.get_output(metric = lookup_metric)\n",
- "print(best_run)\n",
- "print(fitted_model)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Model from a Specific Iteration\n",
- "Show the run and the model from the first iteration:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "iteration = 0\n",
- "best_run, fitted_model = local_run.get_output(iteration = iteration)\n",
- "print(best_run)\n",
- "print(fitted_model)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Test\n",
- "\n",
- "#### Load Test Data"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from sklearn import datasets\n",
- "\n",
- "digits = datasets.load_digits()\n",
- "X_test = digits.data[:10, :]\n",
- "y_test = digits.target[:10]\n",
- "images = digits.images[:10]"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Testing Our Best Fitted Model\n",
- "We will try to predict 2 digits and see how our model works."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "#Randomly select digits and test\n",
- "from matplotlib import pyplot as plt\n",
- "from matplotlib.pyplot import imshow\n",
- "import random\n",
- "import numpy as np\n",
- "\n",
- "for index in np.random.choice(len(y_test), 2, replace = False):\n",
- " print(index)\n",
- " predicted = fitted_model.predict(X_test[index:index + 1])[0]\n",
- " label = y_test[index]\n",
- " title = \"Label value = %d Predicted value = %d \" % (label, predicted)\n",
- " fig = plt.figure(1, figsize=(3,3))\n",
- " ax1 = fig.add_axes((0,0,.8,.8))\n",
- " ax1.set_title(title)\n",
- " plt.imshow(images[index], cmap = plt.cm.gray_r, interpolation = 'nearest')\n",
- " plt.show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Appendix"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Capture the `Dataflow` Objects for Later Use in AutoML\n",
- "\n",
- "`Dataflow` objects are immutable and are composed of a list of data preparation steps. A `Dataflow` object can be branched at any point for further usage."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# sklearn.digits.data + target\n",
- "digits_complete = dprep.auto_read_file('https://dprepdata.blob.core.windows.net/automl-notebook-data/digits-complete.csv')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "`digits_complete` (sourced from `sklearn.datasets.load_digits()`) is forked into `dflow_X` to capture all the feature columns and `dflow_y` to capture the label column."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "digits_complete.to_pandas_dataframe().shape\n",
- "labels_column = 'Column64'\n",
- "dflow_X = digits_complete.drop_columns(columns = [labels_column])\n",
- "dflow_y = digits_complete.keep_columns(columns = [labels_column])"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "savitam"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Automated Machine Learning\n",
+ "_**Prepare Data using `azureml.dataprep` for Local Execution**_\n",
+ "\n",
+ "## Contents\n",
+ "1. [Introduction](#Introduction)\n",
+ "1. [Setup](#Setup)\n",
+ "1. [Data](#Data)\n",
+ "1. [Train](#Train)\n",
+ "1. [Results](#Results)\n",
+ "1. [Test](#Test)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Introduction\n",
+ "In this example we showcase how you can use the `azureml.dataprep` SDK to load and prepare data for AutoML. `azureml.dataprep` can also be used standalone; full documentation can be found [here](https://github.com/Microsoft/PendletonDocs).\n",
+ "\n",
+ "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
+ "\n",
+ "In this notebook you will learn how to:\n",
+ "1. Define data loading and preparation steps in a `Dataflow` using `azureml.dataprep`.\n",
+ "2. Pass the `Dataflow` to AutoML for a local run.\n",
+ "3. Pass the `Dataflow` to AutoML for a remote run."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Setup\n",
+ "\n",
+ "Currently, Data Prep only supports __Ubuntu 16__ and __Red Hat Enterprise Linux 7__. We are working on supporting more linux distros."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Opt-in diagnostics for better experience, quality, and security of future releases."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.telemetry import set_diagnostics_collection\n",
+ "set_diagnostics_collection(send_diagnostics = True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import logging\n",
+ "\n",
+ "import pandas as pd\n",
+ "\n",
+ "import azureml.core\n",
+ "from azureml.core.experiment import Experiment\n",
+ "from azureml.core.workspace import Workspace\n",
+ "import azureml.dataprep as dprep\n",
+ "from azureml.train.automl import AutoMLConfig"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "ws = Workspace.from_config()\n",
+ " \n",
+ "# choose a name for experiment\n",
+ "experiment_name = 'automl-dataprep-local'\n",
+ "# project folder\n",
+ "project_folder = './sample_projects/automl-dataprep-local'\n",
+ " \n",
+ "experiment = Experiment(ws, experiment_name)\n",
+ " \n",
+ "output = {}\n",
+ "output['SDK version'] = azureml.core.VERSION\n",
+ "output['Subscription ID'] = ws.subscription_id\n",
+ "output['Workspace Name'] = ws.name\n",
+ "output['Resource Group'] = ws.resource_group\n",
+ "output['Location'] = ws.location\n",
+ "output['Project Directory'] = project_folder\n",
+ "output['Experiment Name'] = experiment.name\n",
+ "pd.set_option('display.max_colwidth', -1)\n",
+ "outputDf = pd.DataFrame(data = output, index = [''])\n",
+ "outputDf.T"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Data"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# You can use `auto_read_file` which intelligently figures out delimiters and datatypes of a file.\n",
+ "# The data referenced here was pulled from `sklearn.datasets.load_digits()`.\n",
+ "simple_example_data_root = 'https://dprepdata.blob.core.windows.net/automl-notebook-data/'\n",
+ "X = dprep.auto_read_file(simple_example_data_root + 'X.csv').skip(1) # Remove the header row.\n",
+ "\n",
+ "# You can also use `read_csv` and `to_*` transformations to read (with overridable delimiter)\n",
+ "# and convert column types manually.\n",
+ "# Here we read a comma delimited file and convert all columns to integers.\n",
+ "y = dprep.read_csv(simple_example_data_root + 'y.csv').to_long(dprep.ColumnSelector(term='.*', use_regex = True))"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Review the Data Preparation Result\n",
+ "\n",
+ "You can peek the result of a Dataflow at any range using `skip(i)` and `head(j)`. Doing so evaluates only `j` records for all the steps in the Dataflow, which makes it fast even against large datasets."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "X.skip(1).head(5)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Train\n",
+ "\n",
+ "This creates a general AutoML settings object applicable for both local and remote runs."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "automl_settings = {\n",
+ " \"iteration_timeout_minutes\" : 10,\n",
+ " \"iterations\" : 2,\n",
+ " \"primary_metric\" : 'AUC_weighted',\n",
+ " \"preprocess\" : False,\n",
+ " \"verbosity\" : logging.INFO,\n",
+ " \"n_cross_validations\": 3\n",
+ "}"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Pass Data with `Dataflow` Objects\n",
+ "\n",
+ "The `Dataflow` objects captured above can be passed to the `submit` method for a local run. AutoML will retrieve the results from the `Dataflow` for model training."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "automl_config = AutoMLConfig(task = 'classification',\n",
+ " debug_log = 'automl_errors.log',\n",
+ " X = X,\n",
+ " y = y,\n",
+ " **automl_settings)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "local_run = experiment.submit(automl_config, show_output = True)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "local_run"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Results"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Widget for Monitoring Runs\n",
+ "\n",
+ "The widget will first report a \"loading\" status while running the first iteration. After completing the first iteration, an auto-updating graph and table will be shown. The widget will refresh once per minute, so you should see the graph update as child runs complete.\n",
+ "\n",
+ "**Note:** The widget displays a link at the bottom. Use this link to open a web interface to explore the individual run details."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.widgets import RunDetails\n",
+ "RunDetails(local_run).show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Retrieve All Child Runs\n",
+ "You can also use SDK methods to fetch all the child runs and see individual metrics that we log."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "children = list(local_run.get_children())\n",
+ "metricslist = {}\n",
+ "for run in children:\n",
+ " properties = run.get_properties()\n",
+ " metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)}\n",
+ " metricslist[int(properties['iteration'])] = metrics\n",
+ " \n",
+ "rundata = pd.DataFrame(metricslist).sort_index(1)\n",
+ "rundata"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Retrieve the Best Model\n",
+ "\n",
+ "Below we select the best pipeline from our iterations. The `get_output` method returns the best run and the fitted model. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "best_run, fitted_model = local_run.get_output()\n",
+ "print(best_run)\n",
+ "print(fitted_model)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Best Model Based on Any Other Metric\n",
+ "Show the run and the model that has the smallest `log_loss` value:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "lookup_metric = \"log_loss\"\n",
+ "best_run, fitted_model = local_run.get_output(metric = lookup_metric)\n",
+ "print(best_run)\n",
+ "print(fitted_model)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Model from a Specific Iteration\n",
+ "Show the run and the model from the first iteration:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "iteration = 0\n",
+ "best_run, fitted_model = local_run.get_output(iteration = iteration)\n",
+ "print(best_run)\n",
+ "print(fitted_model)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Test\n",
+ "\n",
+ "#### Load Test Data"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from sklearn import datasets\n",
+ "\n",
+ "digits = datasets.load_digits()\n",
+ "X_test = digits.data[:10, :]\n",
+ "y_test = digits.target[:10]\n",
+ "images = digits.images[:10]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Testing Our Best Fitted Model\n",
+ "We will try to predict 2 digits and see how our model works."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "#Randomly select digits and test\n",
+ "from matplotlib import pyplot as plt\n",
+ "import numpy as np\n",
+ "\n",
+ "for index in np.random.choice(len(y_test), 2, replace = False):\n",
+ " print(index)\n",
+ " predicted = fitted_model.predict(X_test[index:index + 1])[0]\n",
+ " label = y_test[index]\n",
+ " title = \"Label value = %d Predicted value = %d \" % (label, predicted)\n",
+ " fig = plt.figure(1, figsize=(3,3))\n",
+ " ax1 = fig.add_axes((0,0,.8,.8))\n",
+ " ax1.set_title(title)\n",
+ " plt.imshow(images[index], cmap = plt.cm.gray_r, interpolation = 'nearest')\n",
+ " plt.show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Appendix"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Capture the `Dataflow` Objects for Later Use in AutoML\n",
+ "\n",
+ "`Dataflow` objects are immutable and are composed of a list of data preparation steps. A `Dataflow` object can be branched at any point for further usage."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# sklearn.digits.data + target\n",
+ "digits_complete = dprep.auto_read_file('https://dprepdata.blob.core.windows.net/automl-notebook-data/digits-complete.csv')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "`digits_complete` (sourced from `sklearn.datasets.load_digits()`) is forked into `dflow_X` to capture all the feature columns and `dflow_y` to capture the label column."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "print(digits_complete.to_pandas_dataframe().shape)\n",
+ "labels_column = 'Column64'\n",
+ "dflow_X = digits_complete.drop_columns(columns = [labels_column])\n",
+ "dflow_y = digits_complete.keep_columns(columns = [labels_column])"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "savitam"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.5"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.5"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/automated-machine-learning/exploring-previous-runs/auto-ml-exploring-previous-runs.ipynb b/how-to-use-azureml/automated-machine-learning/exploring-previous-runs/auto-ml-exploring-previous-runs.ipynb
index bc514883..8aa51733 100644
--- a/how-to-use-azureml/automated-machine-learning/exploring-previous-runs/auto-ml-exploring-previous-runs.ipynb
+++ b/how-to-use-azureml/automated-machine-learning/exploring-previous-runs/auto-ml-exploring-previous-runs.ipynb
@@ -1,370 +1,359 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Automated Machine Learning\n",
- "_**Exploring Previous Runs**_\n",
- "\n",
- "## Contents\n",
- "1. [Introduction](#Introduction)\n",
- "1. [Setup](#Setup)\n",
- "1. [Explore](#Explore)\n",
- "1. [Download](#Download)\n",
- "1. [Register](#Register)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Introduction\n",
- "In this example we present some examples on navigating previously executed runs. We also show how you can download a fitted model for any previous run.\n",
- "\n",
- "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
- "\n",
- "In this notebook you will learn how to:\n",
- "1. List all experiments in a workspace.\n",
- "2. List all AutoML runs in an experiment.\n",
- "3. Get details for an AutoML run, including settings, run widget, and all metrics.\n",
- "4. Download a fitted pipeline for any iteration."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Setup"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import logging\n",
- "import os\n",
- "import random\n",
- "import re\n",
- "\n",
- "from matplotlib import pyplot as plt\n",
- "from matplotlib.pyplot import imshow\n",
- "import numpy as np\n",
- "import pandas as pd\n",
- "from sklearn import datasets\n",
- "\n",
- "import azureml.core\n",
- "from azureml.core.experiment import Experiment\n",
- "from azureml.core.run import Run\n",
- "from azureml.core.workspace import Workspace\n",
- "from azureml.train.automl import AutoMLConfig\n",
- "from azureml.train.automl.run import AutoMLRun"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "ws = Workspace.from_config()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Opt-in diagnostics for better experience, quality, and security of future releases."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.telemetry import set_diagnostics_collection\n",
- "set_diagnostics_collection(send_diagnostics = True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Explore"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### List Experiments"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "experiment_list = Experiment.list(workspace=ws)\n",
- "\n",
- "summary_df = pd.DataFrame(index = ['No of Runs'])\n",
- "for experiment in experiment_list:\n",
- " automl_runs = list(experiment.get_runs(type='automl'))\n",
- " summary_df[experiment.name] = [len(automl_runs)]\n",
- " \n",
- "pd.set_option('display.max_colwidth', -1)\n",
- "summary_df.T"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### List runs for an experiment\n",
- "Set `experiment_name` to any experiment name from the result of the Experiment.list cell to load the AutoML runs."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "experiment_name = 'automl-local-classification' # Replace this with any project name from previous cell.\n",
- "\n",
- "proj = ws.experiments[experiment_name]\n",
- "summary_df = pd.DataFrame(index = ['Type', 'Status', 'Primary Metric', 'Iterations', 'Compute', 'Name'])\n",
- "automl_runs = list(proj.get_runs(type='automl'))\n",
- "automl_runs_project = []\n",
- "for run in automl_runs:\n",
- " properties = run.get_properties()\n",
- " tags = run.get_tags()\n",
- " amlsettings = eval(properties['RawAMLSettingsString'])\n",
- " if 'iterations' in tags:\n",
- " iterations = tags['iterations']\n",
- " else:\n",
- " iterations = properties['num_iterations']\n",
- " summary_df[run.id] = [amlsettings['task_type'], run.get_details()['status'], properties['primary_metric'], iterations, properties['target'], amlsettings['name']]\n",
- " if run.get_details()['status'] == 'Completed':\n",
- " automl_runs_project.append(run.id)\n",
- " \n",
- "from IPython.display import HTML\n",
- "projname_html = HTML(\"{}
\".format(proj.name))\n",
- "\n",
- "from IPython.display import display\n",
- "display(projname_html)\n",
- "display(summary_df.T)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Get details for a run\n",
- "\n",
- "Copy the project name and run id from the previous cell output to find more details on a particular run."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run_id = automl_runs_project[0] # Replace with your own run_id from above run ids\n",
- "assert (run_id in summary_df.keys()), \"Run id not found! Please set run id to a value from above run ids\"\n",
- "\n",
- "from azureml.widgets import RunDetails\n",
- "\n",
- "experiment = Experiment(ws, experiment_name)\n",
- "ml_run = AutoMLRun(experiment = experiment, run_id = run_id)\n",
- "\n",
- "summary_df = pd.DataFrame(index = ['Type', 'Status', 'Primary Metric', 'Iterations', 'Compute', 'Name', 'Start Time', 'End Time'])\n",
- "properties = ml_run.get_properties()\n",
- "tags = ml_run.get_tags()\n",
- "status = ml_run.get_details()\n",
- "amlsettings = eval(properties['RawAMLSettingsString'])\n",
- "if 'iterations' in tags:\n",
- " iterations = tags['iterations']\n",
- "else:\n",
- " iterations = properties['num_iterations']\n",
- "start_time = None\n",
- "if 'startTimeUtc' in status:\n",
- " start_time = status['startTimeUtc']\n",
- "end_time = None\n",
- "if 'endTimeUtc' in status:\n",
- " end_time = status['endTimeUtc']\n",
- "summary_df[ml_run.id] = [amlsettings['task_type'], status['status'], properties['primary_metric'], iterations, properties['target'], amlsettings['name'], start_time, end_time]\n",
- "display(HTML('Runtime Details
'))\n",
- "display(summary_df)\n",
- "\n",
- "#settings_df = pd.DataFrame(data = amlsettings, index = [''])\n",
- "display(HTML('AutoML Settings
'))\n",
- "display(amlsettings)\n",
- "\n",
- "display(HTML('Iterations
'))\n",
- "RunDetails(ml_run).show() \n",
- "\n",
- "children = list(ml_run.get_children())\n",
- "metricslist = {}\n",
- "for run in children:\n",
- " properties = run.get_properties()\n",
- " metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)}\n",
- " metricslist[int(properties['iteration'])] = metrics\n",
- "\n",
- "rundata = pd.DataFrame(metricslist).sort_index(1)\n",
- "display(HTML('Metrics
'))\n",
- "display(rundata)\n"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Download"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Download the Best Model for Any Given Metric"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "metric = 'AUC_weighted' # Replace with a metric name.\n",
- "best_run, fitted_model = ml_run.get_output(metric = metric)\n",
- "fitted_model"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Download the Model for Any Given Iteration"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "iteration = 1 # Replace with an iteration number.\n",
- "best_run, fitted_model = ml_run.get_output(iteration = iteration)\n",
- "fitted_model"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Register"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Register fitted model for deployment\n",
- "If neither `metric` nor `iteration` are specified in the `register_model` call, the iteration with the best primary metric is registered."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "description = 'AutoML Model'\n",
- "tags = None\n",
- "ml_run.register_model(description = description, tags = tags)\n",
- "ml_run.model_id # Use this id to deploy the model as a web service in Azure."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Register the Best Model for Any Given Metric"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "metric = 'AUC_weighted' # Replace with a metric name.\n",
- "description = 'AutoML Model'\n",
- "tags = None\n",
- "ml_run.register_model(description = description, tags = tags, metric = metric)\n",
- "print(ml_run.model_id) # Use this id to deploy the model as a web service in Azure."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Register the Model for Any Given Iteration"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "iteration = 1 # Replace with an iteration number.\n",
- "description = 'AutoML Model'\n",
- "tags = None\n",
- "ml_run.register_model(description = description, tags = tags, iteration = iteration)\n",
- "print(ml_run.model_id) # Use this id to deploy the model as a web service in Azure."
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "savitam"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Automated Machine Learning\n",
+ "_**Exploring Previous Runs**_\n",
+ "\n",
+ "## Contents\n",
+ "1. [Introduction](#Introduction)\n",
+ "1. [Setup](#Setup)\n",
+ "1. [Explore](#Explore)\n",
+ "1. [Download](#Download)\n",
+ "1. [Register](#Register)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Introduction\n",
+ "In this example we present some examples on navigating previously executed runs. We also show how you can download a fitted model for any previous run.\n",
+ "\n",
+ "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
+ "\n",
+ "In this notebook you will learn how to:\n",
+ "1. List all experiments in a workspace.\n",
+ "2. List all AutoML runs in an experiment.\n",
+ "3. Get details for an AutoML run, including settings, run widget, and all metrics.\n",
+ "4. Download a fitted pipeline for any iteration."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Setup"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import pandas as pd\n",
+ "import json\n",
+ "\n",
+ "from azureml.core.experiment import Experiment\n",
+ "from azureml.core.workspace import Workspace\n",
+ "from azureml.train.automl.run import AutoMLRun"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "ws = Workspace.from_config()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Opt-in diagnostics for better experience, quality, and security of future releases."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.telemetry import set_diagnostics_collection\n",
+ "set_diagnostics_collection(send_diagnostics = True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Explore"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### List Experiments"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "experiment_list = Experiment.list(workspace=ws)\n",
+ "\n",
+ "summary_df = pd.DataFrame(index = ['No of Runs'])\n",
+ "for experiment in experiment_list:\n",
+ " automl_runs = list(experiment.get_runs(type='automl'))\n",
+ " summary_df[experiment.name] = [len(automl_runs)]\n",
+ " \n",
+ "pd.set_option('display.max_colwidth', -1)\n",
+ "summary_df.T"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### List runs for an experiment\n",
+ "Set `experiment_name` to any experiment name from the result of the Experiment.list cell to load the AutoML runs."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "experiment_name = 'automl-local-classification' # Replace this with any project name from previous cell.\n",
+ "\n",
+ "proj = ws.experiments[experiment_name]\n",
+ "summary_df = pd.DataFrame(index = ['Type', 'Status', 'Primary Metric', 'Iterations', 'Compute', 'Name'])\n",
+ "automl_runs = list(proj.get_runs(type='automl'))\n",
+ "automl_runs_project = []\n",
+ "for run in automl_runs:\n",
+ " properties = run.get_properties()\n",
+ " tags = run.get_tags()\n",
+ " amlsettings = json.loads(properties['AMLSettingsJsonString'])\n",
+ " if 'iterations' in tags:\n",
+ " iterations = tags['iterations']\n",
+ " else:\n",
+ " iterations = properties['num_iterations']\n",
+ " summary_df[run.id] = [amlsettings['task_type'], run.get_details()['status'], properties['primary_metric'], iterations, properties['target'], amlsettings['name']]\n",
+ " if run.get_details()['status'] == 'Completed':\n",
+ " automl_runs_project.append(run.id)\n",
+ " \n",
+ "from IPython.display import HTML\n",
+ "projname_html = HTML(\"{}
\".format(proj.name))\n",
+ "\n",
+ "from IPython.display import display\n",
+ "display(projname_html)\n",
+ "display(summary_df.T)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Get details for a run\n",
+ "\n",
+ "Copy the project name and run id from the previous cell output to find more details on a particular run."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run_id = automl_runs_project[0] # Replace with your own run_id from above run ids\n",
+ "assert (run_id in summary_df.keys()), \"Run id not found! Please set run id to a value from above run ids\"\n",
+ "\n",
+ "from azureml.widgets import RunDetails\n",
+ "\n",
+ "experiment = Experiment(ws, experiment_name)\n",
+ "ml_run = AutoMLRun(experiment = experiment, run_id = run_id)\n",
+ "\n",
+ "summary_df = pd.DataFrame(index = ['Type', 'Status', 'Primary Metric', 'Iterations', 'Compute', 'Name', 'Start Time', 'End Time'])\n",
+ "properties = ml_run.get_properties()\n",
+ "tags = ml_run.get_tags()\n",
+ "status = ml_run.get_details()\n",
+ "amlsettings = json.loads(properties['AMLSettingsJsonString'])\n",
+ "if 'iterations' in tags:\n",
+ " iterations = tags['iterations']\n",
+ "else:\n",
+ " iterations = properties['num_iterations']\n",
+ "start_time = None\n",
+ "if 'startTimeUtc' in status:\n",
+ " start_time = status['startTimeUtc']\n",
+ "end_time = None\n",
+ "if 'endTimeUtc' in status:\n",
+ " end_time = status['endTimeUtc']\n",
+ "summary_df[ml_run.id] = [amlsettings['task_type'], status['status'], properties['primary_metric'], iterations, properties['target'], amlsettings['name'], start_time, end_time]\n",
+ "display(HTML('Runtime Details
'))\n",
+ "display(summary_df)\n",
+ "\n",
+ "#settings_df = pd.DataFrame(data = amlsettings, index = [''])\n",
+ "display(HTML('AutoML Settings
'))\n",
+ "display(amlsettings)\n",
+ "\n",
+ "display(HTML('Iterations
'))\n",
+ "RunDetails(ml_run).show() \n",
+ "\n",
+ "children = list(ml_run.get_children())\n",
+ "metricslist = {}\n",
+ "for run in children:\n",
+ " properties = run.get_properties()\n",
+ " metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)}\n",
+ " metricslist[int(properties['iteration'])] = metrics\n",
+ "\n",
+ "rundata = pd.DataFrame(metricslist).sort_index(1)\n",
+ "display(HTML('Metrics
'))\n",
+ "display(rundata)\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Download"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Download the Best Model for Any Given Metric"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "metric = 'AUC_weighted' # Replace with a metric name.\n",
+ "best_run, fitted_model = ml_run.get_output(metric = metric)\n",
+ "fitted_model"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Download the Model for Any Given Iteration"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "iteration = 1 # Replace with an iteration number.\n",
+ "best_run, fitted_model = ml_run.get_output(iteration = iteration)\n",
+ "fitted_model"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Register"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Register fitted model for deployment\n",
+ "If neither `metric` nor `iteration` are specified in the `register_model` call, the iteration with the best primary metric is registered."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "description = 'AutoML Model'\n",
+ "tags = None\n",
+ "ml_run.register_model(description = description, tags = tags)\n",
+ "print(ml_run.model_id) # Use this id to deploy the model as a web service in Azure."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Register the Best Model for Any Given Metric"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "metric = 'AUC_weighted' # Replace with a metric name.\n",
+ "description = 'AutoML Model'\n",
+ "tags = None\n",
+ "ml_run.register_model(description = description, tags = tags, metric = metric)\n",
+ "print(ml_run.model_id) # Use this id to deploy the model as a web service in Azure."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Register the Model for Any Given Iteration"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "iteration = 1 # Replace with an iteration number.\n",
+ "description = 'AutoML Model'\n",
+ "tags = None\n",
+ "ml_run.register_model(description = description, tags = tags, iteration = iteration)\n",
+ "print(ml_run.model_id) # Use this id to deploy the model as a web service in Azure."
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "savitam"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.6"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/automated-machine-learning/forecasting-energy-demand/auto-ml-forecasting-energy-demand.ipynb b/how-to-use-azureml/automated-machine-learning/forecasting-energy-demand/auto-ml-forecasting-energy-demand.ipynb
index 52c959bd..72bc6dc2 100644
--- a/how-to-use-azureml/automated-machine-learning/forecasting-energy-demand/auto-ml-forecasting-energy-demand.ipynb
+++ b/how-to-use-azureml/automated-machine-learning/forecasting-energy-demand/auto-ml-forecasting-energy-demand.ipynb
@@ -1,418 +1,376 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Automated Machine Learning\n",
- "_**Energy Demand Forecasting**_\n",
- "\n",
- "## Contents\n",
- "1. [Introduction](#Introduction)\n",
- "1. [Setup](#Setup)\n",
- "1. [Data](#Data)\n",
- "1. [Train](#Train)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Introduction\n",
- "In this example, we show how AutoML can be used for energy demand forecasting.\n",
- "\n",
- "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
- "\n",
- "In this notebook you would see\n",
- "1. Creating an Experiment in an existing Workspace\n",
- "2. Instantiating AutoMLConfig with new task type \"forecasting\" for timeseries data training, and other timeseries related settings: for this dataset we use the basic one: \"time_column_name\" \n",
- "3. Training the Model using local compute\n",
- "4. Exploring the results\n",
- "5. Testing the fitted model"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Setup\n",
- "\n",
- "As part of the setup you have already created a Workspace. For AutoML you would need to create an Experiment. An Experiment is a named object in a Workspace, which is used to run experiments."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import azureml.core\n",
- "import pandas as pd\n",
- "import numpy as np\n",
- "import os\n",
- "import logging\n",
- "import warnings\n",
- "# Squash warning messages for cleaner output in the notebook\n",
- "warnings.showwarning = lambda *args, **kwargs: None\n",
- "\n",
- "\n",
- "from azureml.core.workspace import Workspace\n",
- "from azureml.core.experiment import Experiment\n",
- "from azureml.train.automl import AutoMLConfig\n",
- "from azureml.train.automl.run import AutoMLRun\n",
- "from matplotlib import pyplot as plt\n",
- "from matplotlib.pyplot import imshow\n",
- "from sklearn.metrics import mean_absolute_error, mean_squared_error, r2_score"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "ws = Workspace.from_config()\n",
- "\n",
- "# choose a name for the run history container in the workspace\n",
- "experiment_name = 'automl-energydemandforecasting'\n",
- "# project folder\n",
- "project_folder = './sample_projects/automl-local-energydemandforecasting'\n",
- "\n",
- "experiment = Experiment(ws, experiment_name)\n",
- "\n",
- "output = {}\n",
- "output['SDK version'] = azureml.core.VERSION\n",
- "output['Subscription ID'] = ws.subscription_id\n",
- "output['Workspace'] = ws.name\n",
- "output['Resource Group'] = ws.resource_group\n",
- "output['Location'] = ws.location\n",
- "output['Project Directory'] = project_folder\n",
- "output['Run History Name'] = experiment_name\n",
- "pd.set_option('display.max_colwidth', -1)\n",
- "pd.DataFrame(data=output, index=['']).T"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Data\n",
- "Read energy demanding data from file, and preview data."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "data = pd.read_csv(\"nyc_energy.csv\", parse_dates=['timeStamp'])\n",
- "data.head()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Split the data to train and test\n",
- "\n"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "train = data[data['timeStamp'] < '2017-02-01']\n",
- "test = data[data['timeStamp'] >= '2017-02-01']\n"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Prepare the test data, we will feed X_test to the fitted model and get prediction"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "y_test = test.pop('demand').values\n",
- "X_test = test"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Split the train data to train and valid\n",
- "\n",
- "Use one month's data as valid data\n"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "X_train = train[train['timeStamp'] < '2017-01-01']\n",
- "X_valid = train[train['timeStamp'] >= '2017-01-01']\n",
- "y_train = X_train.pop('demand').values\n",
- "y_valid = X_valid.pop('demand').values\n",
- "print(X_train.shape)\n",
- "print(y_train.shape)\n",
- "print(X_valid.shape)\n",
- "print(y_valid.shape)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Train\n",
- "\n",
- "Instantiate a AutoMLConfig object. This defines the settings and data used to run the experiment.\n",
- "\n",
- "|Property|Description|\n",
- "|-|-|\n",
- "|**task**|forecasting|\n",
- "|**primary_metric**|This is the metric that you want to optimize.
Forecasting supports the following primary metrics
spearman_correlation
normalized_root_mean_squared_error
r2_score
normalized_mean_absolute_error\n",
- "|**iterations**|Number of iterations. In each iteration, Auto ML trains a specific pipeline on the given data|\n",
- "|**iteration_timeout_minutes**|Time limit in minutes for each iteration.|\n",
- "|**X**|(sparse) array-like, shape = [n_samples, n_features]|\n",
- "|**y**|(sparse) array-like, shape = [n_samples, ], [n_samples, n_classes]
Multi-class targets. An indicator matrix turns on multilabel classification. This should be an array of integers. |\n",
- "|**X_valid**|Data used to evaluate a model in a iteration. (sparse) array-like, shape = [n_samples, n_features]|\n",
- "|**y_valid**|Data used to evaluate a model in a iteration. (sparse) array-like, shape = [n_samples, ], [n_samples, n_classes]
Multi-class targets. An indicator matrix turns on multilabel classification. This should be an array of integers. |\n",
- "|**path**|Relative path to the project folder. AutoML stores configuration files for the experiment under this folder. You can specify a new empty folder. "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "time_column_name = 'timeStamp'\n",
- "automl_settings = {\n",
- " \"time_column_name\": time_column_name,\n",
- "}\n",
- "\n",
- "\n",
- "automl_config = AutoMLConfig(task = 'forecasting',\n",
- " debug_log = 'automl_nyc_energy_errors.log',\n",
- " primary_metric='normalized_root_mean_squared_error',\n",
- " iterations = 10,\n",
- " iteration_timeout_minutes = 5,\n",
- " X = X_train,\n",
- " y = y_train,\n",
- " X_valid = X_valid,\n",
- " y_valid = y_valid,\n",
- " path=project_folder,\n",
- " verbosity = logging.INFO,\n",
- " **automl_settings)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "You can call the submit method on the experiment object and pass the run configuration. For Local runs the execution is synchronous. Depending on the data and number of iterations this can run for while.\n",
- "You will see the currently running iterations printing to the console."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "local_run = experiment.submit(automl_config, show_output=True)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "local_run"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Retrieve the Best Model\n",
- "Below we select the best pipeline from our iterations. The get_output method on automl_classifier returns the best run and the fitted model for the last fit invocation. There are overloads on get_output that allow you to retrieve the best run and fitted model for any logged metric or a particular iteration."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "best_run, fitted_model = local_run.get_output()\n",
- "fitted_model.steps"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Test the Best Fitted Model\n",
- "\n",
- "Predict on training and test set, and calculate residual values."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "y_pred = fitted_model.predict(X_test)\n",
- "y_pred"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Define a Check Data Function\n",
- "\n",
- "Remove the nan values from y_test to avoid error when calculate metrics "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "def _check_calc_input(y_true, y_pred, rm_na=True):\n",
- " \"\"\"\n",
- " Check that 'y_true' and 'y_pred' are non-empty and\n",
- " have equal length.\n",
- "\n",
- " :param y_true: Vector of actual values\n",
- " :type y_true: array-like\n",
- "\n",
- " :param y_pred: Vector of predicted values\n",
- " :type y_pred: array-like\n",
- "\n",
- " :param rm_na:\n",
- " If rm_na=True, remove entries where y_true=NA and y_pred=NA.\n",
- " :type rm_na: boolean\n",
- "\n",
- " :return:\n",
- " Tuple (y_true, y_pred). if rm_na=True,\n",
- " the returned vectors may differ from their input values.\n",
- " :rtype: Tuple with 2 entries\n",
- " \"\"\"\n",
- " if len(y_true) != len(y_pred):\n",
- " raise ValueError(\n",
- " 'the true values and prediction values do not have equal length.')\n",
- " elif len(y_true) == 0:\n",
- " raise ValueError(\n",
- " 'y_true and y_pred are empty.')\n",
- " # if there is any non-numeric element in the y_true or y_pred,\n",
- " # the ValueError exception will be thrown.\n",
- " y_true = np.array(y_true).astype(float)\n",
- " y_pred = np.array(y_pred).astype(float)\n",
- " if rm_na:\n",
- " # remove entries both in y_true and y_pred where at least\n",
- " # one element in y_true or y_pred is missing\n",
- " y_true_rm_na = y_true[~(np.isnan(y_true) | np.isnan(y_pred))]\n",
- " y_pred_rm_na = y_pred[~(np.isnan(y_true) | np.isnan(y_pred))]\n",
- " return (y_true_rm_na, y_pred_rm_na)\n",
- " else:\n",
- " return y_true, y_pred"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Use the Check Data Function to remove the nan values from y_test to avoid error when calculate metrics "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "y_test,y_pred = _check_calc_input(y_test,y_pred)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Calculate metrics for the prediction\n"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "print(\"[Test Data] \\nRoot Mean squared error: %.2f\" % np.sqrt(mean_squared_error(y_test, y_pred)))\n",
- "# Explained variance score: 1 is perfect prediction\n",
- "print('mean_absolute_error score: %.2f' % mean_absolute_error(y_test, y_pred))\n",
- "print('R2 score: %.2f' % r2_score(y_test, y_pred))\n",
- "\n",
- "\n",
- "\n",
- "# Plot outputs\n",
- "%matplotlib notebook\n",
- "test_pred = plt.scatter(y_test, y_pred, color='b')\n",
- "test_test = plt.scatter(y_test, y_test, color='g')\n",
- "plt.legend((test_pred, test_test), ('prediction', 'truth'), loc='upper left', fontsize=8)\n",
- "plt.show()"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "xiaga"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Automated Machine Learning\n",
+ "_**Energy Demand Forecasting**_\n",
+ "\n",
+ "## Contents\n",
+ "1. [Introduction](#Introduction)\n",
+ "1. [Setup](#Setup)\n",
+ "1. [Data](#Data)\n",
+ "1. [Train](#Train)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Introduction\n",
+ "In this example, we show how AutoML can be used for energy demand forecasting.\n",
+ "\n",
+ "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
+ "\n",
+ "In this notebook you would see\n",
+ "1. Creating an Experiment in an existing Workspace\n",
+ "2. Instantiating AutoMLConfig with new task type \"forecasting\" for timeseries data training, and other timeseries related settings: for this dataset we use the basic one: \"time_column_name\" \n",
+ "3. Training the Model using local compute\n",
+ "4. Exploring the results\n",
+ "5. Testing the fitted model"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Setup\n",
+ "\n",
+ "As part of the setup you have already created a Workspace. For AutoML you would need to create an Experiment. An Experiment is a named object in a Workspace, which is used to run experiments."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import azureml.core\n",
+ "import pandas as pd\n",
+ "import numpy as np\n",
+ "import logging\n",
+ "import warnings\n",
+ "# Squash warning messages for cleaner output in the notebook\n",
+ "warnings.showwarning = lambda *args, **kwargs: None\n",
+ "\n",
+ "\n",
+ "from azureml.core.workspace import Workspace\n",
+ "from azureml.core.experiment import Experiment\n",
+ "from azureml.train.automl import AutoMLConfig\n",
+ "from matplotlib import pyplot as plt\n",
+ "from sklearn.metrics import mean_absolute_error, mean_squared_error, r2_score"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "ws = Workspace.from_config()\n",
+ "\n",
+ "# choose a name for the run history container in the workspace\n",
+ "experiment_name = 'automl-energydemandforecasting'\n",
+ "# project folder\n",
+ "project_folder = './sample_projects/automl-local-energydemandforecasting'\n",
+ "\n",
+ "experiment = Experiment(ws, experiment_name)\n",
+ "\n",
+ "output = {}\n",
+ "output['SDK version'] = azureml.core.VERSION\n",
+ "output['Subscription ID'] = ws.subscription_id\n",
+ "output['Workspace'] = ws.name\n",
+ "output['Resource Group'] = ws.resource_group\n",
+ "output['Location'] = ws.location\n",
+ "output['Project Directory'] = project_folder\n",
+ "output['Run History Name'] = experiment_name\n",
+ "pd.set_option('display.max_colwidth', -1)\n",
+ "outputDf = pd.DataFrame(data = output, index = [''])\n",
+ "outputDf.T"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Data\n",
+ "Read energy demanding data from file, and preview data."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "data = pd.read_csv(\"nyc_energy.csv\", parse_dates=['timeStamp'])\n",
+ "data.head()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Split the data to train and test\n",
+ "\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "train = data[data['timeStamp'] < '2017-02-01']\n",
+ "test = data[data['timeStamp'] >= '2017-02-01']\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Prepare the test data, we will feed X_test to the fitted model and get prediction"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "y_test = test.pop('demand').values\n",
+ "X_test = test"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Split the train data to train and valid\n",
+ "\n",
+ "Use one month's data as valid data\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "X_train = train[train['timeStamp'] < '2017-01-01']\n",
+ "X_valid = train[train['timeStamp'] >= '2017-01-01']\n",
+ "y_train = X_train.pop('demand').values\n",
+ "y_valid = X_valid.pop('demand').values\n",
+ "print(X_train.shape)\n",
+ "print(y_train.shape)\n",
+ "print(X_valid.shape)\n",
+ "print(y_valid.shape)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Train\n",
+ "\n",
+ "Instantiate a AutoMLConfig object. This defines the settings and data used to run the experiment.\n",
+ "\n",
+ "|Property|Description|\n",
+ "|-|-|\n",
+ "|**task**|forecasting|\n",
+ "|**primary_metric**|This is the metric that you want to optimize.
Forecasting supports the following primary metrics
spearman_correlation
normalized_root_mean_squared_error
r2_score
normalized_mean_absolute_error\n",
+ "|**iterations**|Number of iterations. In each iteration, Auto ML trains a specific pipeline on the given data|\n",
+ "|**iteration_timeout_minutes**|Time limit in minutes for each iteration.|\n",
+ "|**X**|(sparse) array-like, shape = [n_samples, n_features]|\n",
+ "|**y**|(sparse) array-like, shape = [n_samples, ], [n_samples, n_classes]
Multi-class targets. An indicator matrix turns on multilabel classification. This should be an array of integers. |\n",
+ "|**X_valid**|Data used to evaluate a model in a iteration. (sparse) array-like, shape = [n_samples, n_features]|\n",
+ "|**y_valid**|Data used to evaluate a model in a iteration. (sparse) array-like, shape = [n_samples, ], [n_samples, n_classes]
Multi-class targets. An indicator matrix turns on multilabel classification. This should be an array of integers. |\n",
+ "|**path**|Relative path to the project folder. AutoML stores configuration files for the experiment under this folder. You can specify a new empty folder. "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "time_column_name = 'timeStamp'\n",
+ "automl_settings = {\n",
+ " \"time_column_name\": time_column_name,\n",
+ "}\n",
+ "\n",
+ "\n",
+ "automl_config = AutoMLConfig(task = 'forecasting',\n",
+ " debug_log = 'automl_nyc_energy_errors.log',\n",
+ " primary_metric='normalized_root_mean_squared_error',\n",
+ " iterations = 10,\n",
+ " iteration_timeout_minutes = 5,\n",
+ " X = X_train,\n",
+ " y = y_train,\n",
+ " X_valid = X_valid,\n",
+ " y_valid = y_valid,\n",
+ " path=project_folder,\n",
+ " verbosity = logging.INFO,\n",
+ " **automl_settings)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "You can call the submit method on the experiment object and pass the run configuration. For Local runs the execution is synchronous. Depending on the data and number of iterations this can run for while.\n",
+ "You will see the currently running iterations printing to the console."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "local_run = experiment.submit(automl_config, show_output=True)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "local_run"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Retrieve the Best Model\n",
+ "Below we select the best pipeline from our iterations. The get_output method on automl_classifier returns the best run and the fitted model for the last fit invocation. There are overloads on get_output that allow you to retrieve the best run and fitted model for any logged metric or a particular iteration."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "best_run, fitted_model = local_run.get_output()\n",
+ "fitted_model.steps"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Test the Best Fitted Model\n",
+ "\n",
+ "Predict on training and test set, and calculate residual values."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "y_pred = fitted_model.predict(X_test)\n",
+ "y_pred"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Use the Check Data Function to remove the nan values from y_test to avoid error when calculate metrics "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "if len(y_test) != len(y_pred):\n",
+ " raise ValueError(\n",
+ " 'the true values and prediction values do not have equal length.')\n",
+ "elif len(y_test) == 0:\n",
+ " raise ValueError(\n",
+ " 'y_true and y_pred are empty.')\n",
+ "\n",
+ "# if there is any non-numeric element in the y_true or y_pred,\n",
+ "# the ValueError exception will be thrown.\n",
+ "y_test_f = np.array(y_test).astype(float)\n",
+ "y_pred_f = np.array(y_pred).astype(float)\n",
+ "\n",
+ "# remove entries both in y_true and y_pred where at least\n",
+ "# one element in y_true or y_pred is missing\n",
+ "y_test = y_test_f[~(np.isnan(y_test_f) | np.isnan(y_pred_f))]\n",
+ "y_pred = y_pred_f[~(np.isnan(y_test_f) | np.isnan(y_pred_f))]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Calculate metrics for the prediction\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "print(\"[Test Data] \\nRoot Mean squared error: %.2f\" % np.sqrt(mean_squared_error(y_test, y_pred)))\n",
+ "# Explained variance score: 1 is perfect prediction\n",
+ "print('mean_absolute_error score: %.2f' % mean_absolute_error(y_test, y_pred))\n",
+ "print('R2 score: %.2f' % r2_score(y_test, y_pred))\n",
+ "\n",
+ "\n",
+ "\n",
+ "# Plot outputs\n",
+ "%matplotlib notebook\n",
+ "test_pred = plt.scatter(y_test, y_pred, color='b')\n",
+ "test_test = plt.scatter(y_test, y_test, color='g')\n",
+ "plt.legend((test_pred, test_test), ('prediction', 'truth'), loc='upper left', fontsize=8)\n",
+ "plt.show()"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "xiaga"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.6"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/automated-machine-learning/forecasting-orange-juice-sales/auto-ml-forecasting-orange-juice-sales.ipynb b/how-to-use-azureml/automated-machine-learning/forecasting-orange-juice-sales/auto-ml-forecasting-orange-juice-sales.ipynb
index 00084daf..45cdc291 100644
--- a/how-to-use-azureml/automated-machine-learning/forecasting-orange-juice-sales/auto-ml-forecasting-orange-juice-sales.ipynb
+++ b/how-to-use-azureml/automated-machine-learning/forecasting-orange-juice-sales/auto-ml-forecasting-orange-juice-sales.ipynb
@@ -1,413 +1,412 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Automated Machine Learning\n",
- "_**Orange Juice Sales Forecasting**_\n",
- "\n",
- "## Contents\n",
- "1. [Introduction](#Introduction)\n",
- "1. [Setup](#Setup)\n",
- "1. [Data](#Data)\n",
- "1. [Train](#Train)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Introduction\n",
- "In this example, we use AutoML to find and tune a time-series forecasting model.\n",
- "\n",
- "Make sure you have executed the [configuration notebook](../../../configuration.ipynb) before running this notebook.\n",
- "\n",
- "In this notebook, you will:\n",
- "1. Create an Experiment in an existing Workspace\n",
- "2. Instantiate an AutoMLConfig \n",
- "3. Find and train a forecasting model using local compute\n",
- "4. Evaluate the performance of the model\n",
- "\n",
- "The examples in the follow code samples use the [University of Chicago's Dominick's Finer Foods dataset](https://research.chicagobooth.edu/kilts/marketing-databases/dominicks) to forecast orange juice sales. Dominick's was a grocery chain in the Chicago metropolitan area."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Setup\n",
- "\n",
- "As part of the setup you have already created a Workspace. To run AutoML, you also need to create an Experiment. An Experiment is a named object in a Workspace which represents a predictive task, the output of which is a trained model and a set of evaluation metrics for the model. "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import azureml.core\n",
- "import pandas as pd\n",
- "import numpy as np\n",
- "import os\n",
- "import logging\n",
- "import warnings\n",
- "# Squash warning messages for cleaner output in the notebook\n",
- "warnings.showwarning = lambda *args, **kwargs: None\n",
- "\n",
- "\n",
- "from azureml.core.workspace import Workspace\n",
- "from azureml.core.experiment import Experiment\n",
- "from azureml.train.automl import AutoMLConfig\n",
- "from azureml.train.automl.run import AutoMLRun\n",
- "from sklearn.metrics import mean_absolute_error, mean_squared_error"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "ws = Workspace.from_config()\n",
- "\n",
- "# choose a name for the run history container in the workspace\n",
- "experiment_name = 'automl-ojsalesforecasting'\n",
- "# project folder\n",
- "project_folder = './sample_projects/automl-local-ojsalesforecasting'\n",
- "\n",
- "experiment = Experiment(ws, experiment_name)\n",
- "\n",
- "output = {}\n",
- "output['SDK version'] = azureml.core.VERSION\n",
- "output['Subscription ID'] = ws.subscription_id\n",
- "output['Workspace'] = ws.name\n",
- "output['Resource Group'] = ws.resource_group\n",
- "output['Location'] = ws.location\n",
- "output['Project Directory'] = project_folder\n",
- "output['Run History Name'] = experiment_name\n",
- "pd.set_option('display.max_colwidth', -1)\n",
- "pd.DataFrame(data=output, index=['']).T"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Data\n",
- "You are now ready to load the historical orange juice sales data. We will load the CSV file into a plain pandas DataFrame; the time column in the CSV is called _WeekStarting_, so it will be specially parsed into the datetime type."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "time_column_name = 'WeekStarting'\n",
- "data = pd.read_csv(\"dominicks_OJ.csv\", parse_dates=[time_column_name])\n",
- "data.head()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Each row in the DataFrame holds a quantity of weekly sales for an OJ brand at a single store. The data also includes the sales price, a flag indicating if the OJ brand was advertised in the store that week, and some customer demographic information based on the store location. For historical reasons, the data also include the logarithm of the sales quantity. The Dominick's grocery data is commonly used to illustrate econometric modeling techniques where logarithms of quantities are generally preferred. \n",
- "\n",
- "The task is now to build a time-series model for the _Quantity_ column. It is important to note that this dataset is comprised of many individual time-series - one for each unique combination of _Store_ and _Brand_. To distinguish the individual time-series, we thus define the **grain** - the columns whose values determine the boundaries between time-series: "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "grain_column_names = ['Store', 'Brand']\n",
- "nseries = data.groupby(grain_column_names).ngroups\n",
- "print('Data contains {0} individual time-series.'.format(nseries))"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Data Splitting\n",
- "For the purposes of demonstration and later forecast evaluation, we now split the data into a training and a testing set. The test set will contain the final 20 weeks of observed sales for each time-series."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "ntest_periods = 20\n",
- "\n",
- "def split_last_n_by_grain(df, n):\n",
- " \"\"\"\n",
- " Group df by grain and split on last n rows for each group\n",
- " \"\"\"\n",
- " df_grouped = (df.sort_values(time_column_name) # Sort by ascending time\n",
- " .groupby(grain_column_names, group_keys=False))\n",
- " df_head = df_grouped.apply(lambda dfg: dfg.iloc[:-n])\n",
- " df_tail = df_grouped.apply(lambda dfg: dfg.iloc[-n:])\n",
- " return df_head, df_tail\n",
- "\n",
- "X_train, X_test = split_last_n_by_grain(data, ntest_periods)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Modeling\n",
- "\n",
- "For forecasting tasks, AutoML uses pre-processing and estimation steps that are specific to time-series. AutoML will undertake the following pre-processing steps:\n",
- "* Detect time-series sample frequency (e.g. hourly, daily, weekly) and create new records for absent time points to make the series regular. A regular time series has a well-defined frequency and has a value at every sample point in a contiguous time span \n",
- "* Impute missing values in the target (via forward-fill) and feature columns (using median column values) \n",
- "* Create grain-based features to enable fixed effects across different series\n",
- "* Create time-based features to assist in learning seasonal patterns\n",
- "* Encode categorical variables to numeric quantities\n",
- "\n",
- "AutoML will currently train a single, regression-type model across **all** time-series in a given training set. This allows the model to generalize across related series.\n",
- "\n",
- "You are almost ready to start an AutoML training job. We will first need to create a validation set from the existing training set (i.e. for hyper-parameter tuning): "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "nvalidation_periods = 20\n",
- "X_train, X_validate = split_last_n_by_grain(X_train, nvalidation_periods)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "We also need to separate the target column from the rest of the DataFrame: "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "target_column_name = 'Quantity'\n",
- "y_train = X_train.pop(target_column_name).values\n",
- "y_validate = X_validate.pop(target_column_name).values "
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Train\n",
- "\n",
- "The AutoMLConfig object defines the settings and data for an AutoML training job. Here, we set necessary inputs like the task type, the number of AutoML iterations to try, and the training and validation data. \n",
- "\n",
- "For forecasting tasks, there are some additional parameters that can be set: the name of the column holding the date/time and the grain column names. A time column is required for forecasting, while the grain is optional. If a grain is not given, the forecaster assumes that the whole dataset is a single time-series. We also pass a list of columns to drop prior to modeling. The _logQuantity_ column is completely correlated with the target quantity, so it must be removed to prevent a target leak. \n",
- "\n",
- "|Property|Description|\n",
- "|-|-|\n",
- "|**task**|forecasting|\n",
- "|**primary_metric**|This is the metric that you want to optimize.
Forecasting supports the following primary metrics
spearman_correlation
normalized_root_mean_squared_error
r2_score
normalized_mean_absolute_error\n",
- "|**iterations**|Number of iterations. In each iteration, Auto ML trains a specific pipeline on the given data|\n",
- "|**X**|Training matrix of features, shape = [n_training_samples, n_features]|\n",
- "|**y**|Target values, shape = [n_training_samples, ]|\n",
- "|**X_valid**|Validation matrix of features, shape = [n_validation_samples, n_features]|\n",
- "|**y_valid**|Target values for validation, shape = [n_validation_samples, ]\n",
- "|**enable_ensembling**|Allow AutoML to create ensembles of the best performing models\n",
- "|**debug_log**|Log file path for writing debugging information\n",
- "|**path**|Relative path to the project folder. AutoML stores configuration files for the experiment under this folder. You can specify a new empty folder. "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "automl_settings = {\n",
- " 'time_column_name': time_column_name,\n",
- " 'grain_column_names': grain_column_names,\n",
- " 'drop_column_names': ['logQuantity']\n",
- "}\n",
- "\n",
- "automl_config = AutoMLConfig(task='forecasting',\n",
- " debug_log='automl_oj_sales_errors.log',\n",
- " primary_metric='normalized_root_mean_squared_error',\n",
- " iterations=10,\n",
- " X=X_train,\n",
- " y=y_train,\n",
- " X_valid=X_validate,\n",
- " y_valid=y_validate,\n",
- " enable_ensembling=False,\n",
- " path=project_folder,\n",
- " verbosity=logging.INFO,\n",
- " **automl_settings)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "You can now submit a new training run. For local runs, the execution is synchronous. Depending on the data and number of iterations this operation may take several minutes.\n",
- "Information from each iteration will be printed to the console."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "local_run = experiment.submit(automl_config, show_output=True)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "local_run"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Retrieve the Best Model\n",
- "Each run within an Experiment stores serialized (i.e. pickled) pipelines from the AutoML iterations. We can now retrieve the pipeline with the best performance on the validation dataset:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "best_run, fitted_pipeline = local_run.get_output()\n",
- "fitted_pipeline.steps"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Make Predictions from the Best Fitted Model\n",
- "Now that we have retrieved the best pipeline/model, it can be used to make predictions on test data. First, we remove the target values from the test set:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "y_test = X_test.pop(target_column_name).values"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "X_test.head()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "To produce predictions on the test set, we need to know the feature values at all dates in the test set. This requirement is somewhat reasonable for the OJ sales data since the features mainly consist of price, which is usually set in advance, and customer demographics which are approximately constant for each store over the 20 week forecast horizon in the testing data. \n",
- "\n",
- "The target predictions can be retrieved by calling the `predict` method on the best model:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "y_pred = fitted_pipeline.predict(X_test)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Calculate evaluation metrics for the prediction\n",
- "To evaluate the accuracy of the forecast, we'll compare against the actual sales quantities for some select metrics, included the mean absolute percentage error (MAPE)."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "def MAPE(actual, pred):\n",
- " \"\"\"\n",
- " Calculate mean absolute percentage error.\n",
- " Remove NA and values where actual is close to zero\n",
- " \"\"\"\n",
- " not_na = ~(np.isnan(actual) | np.isnan(pred))\n",
- " not_zero = ~np.isclose(actual, 0.0)\n",
- " actual_safe = actual[not_na & not_zero]\n",
- " pred_safe = pred[not_na & not_zero]\n",
- " APE = 100*np.abs((actual_safe - pred_safe)/actual_safe)\n",
- " return np.mean(APE)\n",
- "\n",
- "print(\"[Test Data] \\nRoot Mean squared error: %.2f\" % np.sqrt(mean_squared_error(y_test, y_pred)))\n",
- "print('mean_absolute_error score: %.2f' % mean_absolute_error(y_test, y_pred))\n",
- "print('MAPE: %.2f' % MAPE(y_test, y_pred))"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "erwright"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Automated Machine Learning\n",
+ "_**Orange Juice Sales Forecasting**_\n",
+ "\n",
+ "## Contents\n",
+ "1. [Introduction](#Introduction)\n",
+ "1. [Setup](#Setup)\n",
+ "1. [Data](#Data)\n",
+ "1. [Train](#Train)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Introduction\n",
+ "In this example, we use AutoML to find and tune a time-series forecasting model.\n",
+ "\n",
+ "Make sure you have executed the [configuration notebook](../../../configuration.ipynb) before running this notebook.\n",
+ "\n",
+ "In this notebook, you will:\n",
+ "1. Create an Experiment in an existing Workspace\n",
+ "2. Instantiate an AutoMLConfig \n",
+ "3. Find and train a forecasting model using local compute\n",
+ "4. Evaluate the performance of the model\n",
+ "\n",
+ "The examples in the follow code samples use the [University of Chicago's Dominick's Finer Foods dataset](https://research.chicagobooth.edu/kilts/marketing-databases/dominicks) to forecast orange juice sales. Dominick's was a grocery chain in the Chicago metropolitan area."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Setup\n",
+ "\n",
+ "As part of the setup you have already created a Workspace. To run AutoML, you also need to create an Experiment. An Experiment is a named object in a Workspace which represents a predictive task, the output of which is a trained model and a set of evaluation metrics for the model. "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import azureml.core\n",
+ "import pandas as pd\n",
+ "import numpy as np\n",
+ "import logging\n",
+ "import warnings\n",
+ "# Squash warning messages for cleaner output in the notebook\n",
+ "warnings.showwarning = lambda *args, **kwargs: None\n",
+ "\n",
+ "\n",
+ "from azureml.core.workspace import Workspace\n",
+ "from azureml.core.experiment import Experiment\n",
+ "from azureml.train.automl import AutoMLConfig\n",
+ "from sklearn.metrics import mean_absolute_error, mean_squared_error"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "ws = Workspace.from_config()\n",
+ "\n",
+ "# choose a name for the run history container in the workspace\n",
+ "experiment_name = 'automl-ojsalesforecasting'\n",
+ "# project folder\n",
+ "project_folder = './sample_projects/automl-local-ojsalesforecasting'\n",
+ "\n",
+ "experiment = Experiment(ws, experiment_name)\n",
+ "\n",
+ "output = {}\n",
+ "output['SDK version'] = azureml.core.VERSION\n",
+ "output['Subscription ID'] = ws.subscription_id\n",
+ "output['Workspace'] = ws.name\n",
+ "output['Resource Group'] = ws.resource_group\n",
+ "output['Location'] = ws.location\n",
+ "output['Project Directory'] = project_folder\n",
+ "output['Run History Name'] = experiment_name\n",
+ "pd.set_option('display.max_colwidth', -1)\n",
+ "outputDf = pd.DataFrame(data = output, index = [''])\n",
+ "outputDf.T"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Data\n",
+ "You are now ready to load the historical orange juice sales data. We will load the CSV file into a plain pandas DataFrame; the time column in the CSV is called _WeekStarting_, so it will be specially parsed into the datetime type."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "time_column_name = 'WeekStarting'\n",
+ "data = pd.read_csv(\"dominicks_OJ.csv\", parse_dates=[time_column_name])\n",
+ "data.head()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Each row in the DataFrame holds a quantity of weekly sales for an OJ brand at a single store. The data also includes the sales price, a flag indicating if the OJ brand was advertised in the store that week, and some customer demographic information based on the store location. For historical reasons, the data also include the logarithm of the sales quantity. The Dominick's grocery data is commonly used to illustrate econometric modeling techniques where logarithms of quantities are generally preferred. \n",
+ "\n",
+ "The task is now to build a time-series model for the _Quantity_ column. It is important to note that this dataset is comprised of many individual time-series - one for each unique combination of _Store_ and _Brand_. To distinguish the individual time-series, we thus define the **grain** - the columns whose values determine the boundaries between time-series: "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "grain_column_names = ['Store', 'Brand']\n",
+ "nseries = data.groupby(grain_column_names).ngroups\n",
+ "print('Data contains {0} individual time-series.'.format(nseries))"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Data Splitting\n",
+ "For the purposes of demonstration and later forecast evaluation, we now split the data into a training and a testing set. The test set will contain the final 20 weeks of observed sales for each time-series."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "ntest_periods = 20\n",
+ "\n",
+ "def split_last_n_by_grain(df, n):\n",
+ " \"\"\"\n",
+ " Group df by grain and split on last n rows for each group\n",
+ " \"\"\"\n",
+ " df_grouped = (df.sort_values(time_column_name) # Sort by ascending time\n",
+ " .groupby(grain_column_names, group_keys=False))\n",
+ " df_head = df_grouped.apply(lambda dfg: dfg.iloc[:-n])\n",
+ " df_tail = df_grouped.apply(lambda dfg: dfg.iloc[-n:])\n",
+ " return df_head, df_tail\n",
+ "\n",
+ "X_train, X_test = split_last_n_by_grain(data, ntest_periods)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Modeling\n",
+ "\n",
+ "For forecasting tasks, AutoML uses pre-processing and estimation steps that are specific to time-series. AutoML will undertake the following pre-processing steps:\n",
+ "* Detect time-series sample frequency (e.g. hourly, daily, weekly) and create new records for absent time points to make the series regular. A regular time series has a well-defined frequency and has a value at every sample point in a contiguous time span \n",
+ "* Impute missing values in the target (via forward-fill) and feature columns (using median column values) \n",
+ "* Create grain-based features to enable fixed effects across different series\n",
+ "* Create time-based features to assist in learning seasonal patterns\n",
+ "* Encode categorical variables to numeric quantities\n",
+ "\n",
+ "AutoML will currently train a single, regression-type model across **all** time-series in a given training set. This allows the model to generalize across related series.\n",
+ "\n",
+ "You are almost ready to start an AutoML training job. We will first need to create a validation set from the existing training set (i.e. for hyper-parameter tuning): "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "nvalidation_periods = 20\n",
+ "X_train, X_validate = split_last_n_by_grain(X_train, nvalidation_periods)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "We also need to separate the target column from the rest of the DataFrame: "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "target_column_name = 'Quantity'\n",
+ "y_train = X_train.pop(target_column_name).values\n",
+ "y_validate = X_validate.pop(target_column_name).values "
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Train\n",
+ "\n",
+ "The AutoMLConfig object defines the settings and data for an AutoML training job. Here, we set necessary inputs like the task type, the number of AutoML iterations to try, and the training and validation data. \n",
+ "\n",
+ "For forecasting tasks, there are some additional parameters that can be set: the name of the column holding the date/time and the grain column names. A time column is required for forecasting, while the grain is optional. If a grain is not given, the forecaster assumes that the whole dataset is a single time-series. We also pass a list of columns to drop prior to modeling. The _logQuantity_ column is completely correlated with the target quantity, so it must be removed to prevent a target leak. \n",
+ "\n",
+ "|Property|Description|\n",
+ "|-|-|\n",
+ "|**task**|forecasting|\n",
+ "|**primary_metric**|This is the metric that you want to optimize.
Forecasting supports the following primary metrics
spearman_correlation
normalized_root_mean_squared_error
r2_score
normalized_mean_absolute_error\n",
+ "|**iterations**|Number of iterations. In each iteration, Auto ML trains a specific pipeline on the given data|\n",
+ "|**X**|Training matrix of features, shape = [n_training_samples, n_features]|\n",
+ "|**y**|Target values, shape = [n_training_samples, ]|\n",
+ "|**X_valid**|Validation matrix of features, shape = [n_validation_samples, n_features]|\n",
+ "|**y_valid**|Target values for validation, shape = [n_validation_samples, ]\n",
+ "|**enable_ensembling**|Allow AutoML to create ensembles of the best performing models\n",
+ "|**debug_log**|Log file path for writing debugging information\n",
+ "|**path**|Relative path to the project folder. AutoML stores configuration files for the experiment under this folder. You can specify a new empty folder. "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "automl_settings = {\n",
+ " 'time_column_name': time_column_name,\n",
+ " 'grain_column_names': grain_column_names,\n",
+ " 'drop_column_names': ['logQuantity']\n",
+ "}\n",
+ "\n",
+ "automl_config = AutoMLConfig(task='forecasting',\n",
+ " debug_log='automl_oj_sales_errors.log',\n",
+ " primary_metric='normalized_root_mean_squared_error',\n",
+ " iterations=10,\n",
+ " X=X_train,\n",
+ " y=y_train,\n",
+ " X_valid=X_validate,\n",
+ " y_valid=y_validate,\n",
+ " enable_ensembling=False,\n",
+ " path=project_folder,\n",
+ " verbosity=logging.INFO,\n",
+ " **automl_settings)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "You can now submit a new training run. For local runs, the execution is synchronous. Depending on the data and number of iterations this operation may take several minutes.\n",
+ "Information from each iteration will be printed to the console."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "local_run = experiment.submit(automl_config, show_output=True)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "local_run"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Retrieve the Best Model\n",
+ "Each run within an Experiment stores serialized (i.e. pickled) pipelines from the AutoML iterations. We can now retrieve the pipeline with the best performance on the validation dataset:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "best_run, fitted_pipeline = local_run.get_output()\n",
+ "fitted_pipeline.steps"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Make Predictions from the Best Fitted Model\n",
+ "Now that we have retrieved the best pipeline/model, it can be used to make predictions on test data. First, we remove the target values from the test set:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "y_test = X_test.pop(target_column_name).values"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "X_test.head()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "To produce predictions on the test set, we need to know the feature values at all dates in the test set. This requirement is somewhat reasonable for the OJ sales data since the features mainly consist of price, which is usually set in advance, and customer demographics which are approximately constant for each store over the 20 week forecast horizon in the testing data. \n",
+ "\n",
+ "The target predictions can be retrieved by calling the `predict` method on the best model:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "y_pred = fitted_pipeline.predict(X_test)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Calculate evaluation metrics for the prediction\n",
+ "To evaluate the accuracy of the forecast, we'll compare against the actual sales quantities for some select metrics, included the mean absolute percentage error (MAPE)."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "def MAPE(actual, pred):\n",
+ " \"\"\"\n",
+ " Calculate mean absolute percentage error.\n",
+ " Remove NA and values where actual is close to zero\n",
+ " \"\"\"\n",
+ " not_na = ~(np.isnan(actual) | np.isnan(pred))\n",
+ " not_zero = ~np.isclose(actual, 0.0)\n",
+ " actual_safe = actual[not_na & not_zero]\n",
+ " pred_safe = pred[not_na & not_zero]\n",
+ " APE = 100*np.abs((actual_safe - pred_safe)/actual_safe)\n",
+ " return np.mean(APE)\n",
+ "\n",
+ "print(\"[Test Data] \\nRoot Mean squared error: %.2f\" % np.sqrt(mean_squared_error(y_test, y_pred)))\n",
+ "print('mean_absolute_error score: %.2f' % mean_absolute_error(y_test, y_pred))\n",
+ "print('MAPE: %.2f' % MAPE(y_test, y_pred))"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "erwright"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.6"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/automated-machine-learning/missing-data-blacklist-early-termination/auto-ml-missing-data-blacklist-early-termination.ipynb b/how-to-use-azureml/automated-machine-learning/missing-data-blacklist-early-termination/auto-ml-missing-data-blacklist-early-termination.ipynb
index 73bd51a5..2a4959d6 100644
--- a/how-to-use-azureml/automated-machine-learning/missing-data-blacklist-early-termination/auto-ml-missing-data-blacklist-early-termination.ipynb
+++ b/how-to-use-azureml/automated-machine-learning/missing-data-blacklist-early-termination/auto-ml-missing-data-blacklist-early-termination.ipynb
@@ -1,401 +1,396 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Automated Machine Learning\n",
- "_**Blacklisting Models, Early Termination, and Handling Missing Data**_\n",
- "\n",
- "## Contents\n",
- "1. [Introduction](#Introduction)\n",
- "1. [Setup](#Setup)\n",
- "1. [Data](#Data)\n",
- "1. [Train](#Train)\n",
- "1. [Results](#Results)\n",
- "1. [Test](#Test)\n"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Introduction\n",
- "In this example we use the scikit-learn's [digit dataset](http://scikit-learn.org/stable/datasets/index.html#optical-recognition-of-handwritten-digits-dataset) to showcase how you can use AutoML for handling missing values in data. We also provide a stopping metric indicating a target for the primary metrics so that AutoML can terminate the run without necessarly going through all the iterations. Finally, if you want to avoid a certain pipeline, we allow you to specify a blacklist of algorithms that AutoML will ignore for this run.\n",
- "\n",
- "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
- "\n",
- "In this notebook you will learn how to:\n",
- "1. Create an `Experiment` in an existing `Workspace`.\n",
- "2. Configure AutoML using `AutoMLConfig`.\n",
- "4. Train the model.\n",
- "5. Explore the results.\n",
- "6. Test the best fitted model.\n",
- "\n",
- "In addition this notebook showcases the following features\n",
- "- **Blacklisting** certain pipelines\n",
- "- Specifying **target metrics** to indicate stopping criteria\n",
- "- Handling **missing data** in the input"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Setup\n",
- "\n",
- "As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import logging\n",
- "import os\n",
- "import random\n",
- "\n",
- "from matplotlib import pyplot as plt\n",
- "from matplotlib.pyplot import imshow\n",
- "import numpy as np\n",
- "import pandas as pd\n",
- "from sklearn import datasets\n",
- "\n",
- "import azureml.core\n",
- "from azureml.core.experiment import Experiment\n",
- "from azureml.core.workspace import Workspace\n",
- "from azureml.train.automl import AutoMLConfig\n",
- "from azureml.train.automl.run import AutoMLRun"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "ws = Workspace.from_config()\n",
- "\n",
- "# Choose a name for the experiment.\n",
- "experiment_name = 'automl-local-missing-data'\n",
- "project_folder = './sample_projects/automl-local-missing-data'\n",
- "\n",
- "experiment = Experiment(ws, experiment_name)\n",
- "\n",
- "output = {}\n",
- "output['SDK version'] = azureml.core.VERSION\n",
- "output['Subscription ID'] = ws.subscription_id\n",
- "output['Workspace'] = ws.name\n",
- "output['Resource Group'] = ws.resource_group\n",
- "output['Location'] = ws.location\n",
- "output['Project Directory'] = project_folder\n",
- "output['Experiment Name'] = experiment.name\n",
- "pd.set_option('display.max_colwidth', -1)\n",
- "pd.DataFrame(data=output, index=['']).T"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Opt-in diagnostics for better experience, quality, and security of future releases."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.telemetry import set_diagnostics_collection\n",
- "set_diagnostics_collection(send_diagnostics = True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Data"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from scipy import sparse\n",
- "\n",
- "digits = datasets.load_digits()\n",
- "X_train = digits.data[10:,:]\n",
- "y_train = digits.target[10:]\n",
- "\n",
- "# Add missing values in 75% of the lines.\n",
- "missing_rate = 0.75\n",
- "n_missing_samples = int(np.floor(X_train.shape[0] * missing_rate))\n",
- "missing_samples = np.hstack((np.zeros(X_train.shape[0] - n_missing_samples, dtype=np.bool), np.ones(n_missing_samples, dtype=np.bool)))\n",
- "rng = np.random.RandomState(0)\n",
- "rng.shuffle(missing_samples)\n",
- "missing_features = rng.randint(0, X_train.shape[1], n_missing_samples)\n",
- "X_train[np.where(missing_samples)[0], missing_features] = np.nan"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "df = pd.DataFrame(data = X_train)\n",
- "df['Label'] = pd.Series(y_train, index=df.index)\n",
- "df.head()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Train\n",
- "\n",
- "Instantiate an `AutoMLConfig` object to specify the settings and data used to run the experiment. This includes setting `experiment_exit_score`, which should cause the run to complete before the `iterations` count is reached.\n",
- "\n",
- "|Property|Description|\n",
- "|-|-|\n",
- "|**task**|classification or regression|\n",
- "|**primary_metric**|This is the metric that you want to optimize. Classification supports the following primary metrics:
accuracy
AUC_weighted
average_precision_score_weighted
norm_macro_recall
precision_score_weighted|\n",
- "|**iteration_timeout_minutes**|Time limit in minutes for each iteration.|\n",
- "|**iterations**|Number of iterations. In each iteration AutoML trains a specific pipeline with the data.|\n",
- "|**n_cross_validations**|Number of cross validation splits.|\n",
- "|**preprocess**|Setting this to *True* enables AutoML to perform preprocessing on the input to handle *missing data*, and to perform some common *feature extraction*.|\n",
- "|**experiment_exit_score**|*double* value indicating the target for *primary_metric*.
Once the target is surpassed the run terminates.|\n",
- "|**blacklist_models**|*List* of *strings* indicating machine learning algorithms for AutoML to avoid in this run.
Allowed values for **Classification**
LogisticRegression
SGD
MultinomialNaiveBayes
BernoulliNaiveBayes
SVM
LinearSVM
KNN
DecisionTree
RandomForest
ExtremeRandomTrees
LightGBM
GradientBoosting
TensorFlowDNN
TensorFlowLinearClassifier
Allowed values for **Regression**
ElasticNet
GradientBoosting
DecisionTree
KNN
LassoLars
SGD
RandomForest
ExtremeRandomTrees
LightGBM
TensorFlowLinearRegressor
TensorFlowDNN|\n",
- "|**X**|(sparse) array-like, shape = [n_samples, n_features]|\n",
- "|**y**|(sparse) array-like, shape = [n_samples, ], [n_samples, n_classes]
Multi-class targets. An indicator matrix turns on multilabel classification. This should be an array of integers.|\n",
- "|**path**|Relative path to the project folder. AutoML stores configuration files for the experiment under this folder. You can specify a new empty folder.|"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "automl_config = AutoMLConfig(task = 'classification',\n",
- " debug_log = 'automl_errors.log',\n",
- " primary_metric = 'AUC_weighted',\n",
- " iteration_timeout_minutes = 60,\n",
- " iterations = 20,\n",
- " n_cross_validations = 5,\n",
- " preprocess = True,\n",
- " experiment_exit_score = 0.9984,\n",
- " blacklist_models = ['KNN','LinearSVM'],\n",
- " verbosity = logging.INFO,\n",
- " X = X_train, \n",
- " y = y_train,\n",
- " path = project_folder)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Call the `submit` method on the experiment object and pass the run configuration. Execution of local runs is synchronous. Depending on the data and the number of iterations this can run for a while.\n",
- "In this example, we specify `show_output = True` to print currently running iterations to the console."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "local_run = experiment.submit(automl_config, show_output = True)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "local_run"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Results"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Widget for Monitoring Runs\n",
- "\n",
- "The widget will first report a \"loading\" status while running the first iteration. After completing the first iteration, an auto-updating graph and table will be shown. The widget will refresh once per minute, so you should see the graph update as child runs complete.\n",
- "\n",
- "**Note:** The widget displays a link at the bottom. Use this link to open a web interface to explore the individual run details."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.widgets import RunDetails\n",
- "RunDetails(local_run).show() "
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "\n",
- "#### Retrieve All Child Runs\n",
- "You can also use SDK methods to fetch all the child runs and see individual metrics that we log."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "children = list(local_run.get_children())\n",
- "metricslist = {}\n",
- "for run in children:\n",
- " properties = run.get_properties()\n",
- " metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)}\n",
- " metricslist[int(properties['iteration'])] = metrics\n",
- "\n",
- "rundata = pd.DataFrame(metricslist).sort_index(1)\n",
- "rundata"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Retrieve the Best Model\n",
- "\n",
- "Below we select the best pipeline from our iterations. The `get_output` method returns the best run and the fitted model. The Model includes the pipeline and any pre-processing. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "best_run, fitted_model = local_run.get_output()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Best Model Based on Any Other Metric\n",
- "Show the run and the model which has the smallest `accuracy` value:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# lookup_metric = \"accuracy\"\n",
- "# best_run, fitted_model = local_run.get_output(metric = lookup_metric)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Model from a Specific Iteration\n",
- "Show the run and the model from the third iteration:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# iteration = 3\n",
- "# best_run, fitted_model = local_run.get_output(iteration = iteration)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Test"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "digits = datasets.load_digits()\n",
- "X_test = digits.data[:10, :]\n",
- "y_test = digits.target[:10]\n",
- "images = digits.images[:10]\n",
- "\n",
- "# Randomly select digits and test.\n",
- "for index in np.random.choice(len(y_test), 2, replace = False):\n",
- " print(index)\n",
- " predicted = fitted_model.predict(X_test[index:index + 1])[0]\n",
- " label = y_test[index]\n",
- " title = \"Label value = %d Predicted value = %d \" % (label, predicted)\n",
- " fig = plt.figure(1, figsize=(3,3))\n",
- " ax1 = fig.add_axes((0,0,.8,.8))\n",
- " ax1.set_title(title)\n",
- " plt.imshow(images[index], cmap = plt.cm.gray_r, interpolation = 'nearest')\n",
- " plt.show()\n"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "savitam"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Automated Machine Learning\n",
+ "_**Blacklisting Models, Early Termination, and Handling Missing Data**_\n",
+ "\n",
+ "## Contents\n",
+ "1. [Introduction](#Introduction)\n",
+ "1. [Setup](#Setup)\n",
+ "1. [Data](#Data)\n",
+ "1. [Train](#Train)\n",
+ "1. [Results](#Results)\n",
+ "1. [Test](#Test)\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Introduction\n",
+ "In this example we use the scikit-learn's [digit dataset](http://scikit-learn.org/stable/datasets/index.html#optical-recognition-of-handwritten-digits-dataset) to showcase how you can use AutoML for handling missing values in data. We also provide a stopping metric indicating a target for the primary metrics so that AutoML can terminate the run without necessarly going through all the iterations. Finally, if you want to avoid a certain pipeline, we allow you to specify a blacklist of algorithms that AutoML will ignore for this run.\n",
+ "\n",
+ "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
+ "\n",
+ "In this notebook you will learn how to:\n",
+ "1. Create an `Experiment` in an existing `Workspace`.\n",
+ "2. Configure AutoML using `AutoMLConfig`.\n",
+ "4. Train the model.\n",
+ "5. Explore the results.\n",
+ "6. Test the best fitted model.\n",
+ "\n",
+ "In addition this notebook showcases the following features\n",
+ "- **Blacklisting** certain pipelines\n",
+ "- Specifying **target metrics** to indicate stopping criteria\n",
+ "- Handling **missing data** in the input"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Setup\n",
+ "\n",
+ "As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import logging\n",
+ "\n",
+ "from matplotlib import pyplot as plt\n",
+ "import numpy as np\n",
+ "import pandas as pd\n",
+ "from sklearn import datasets\n",
+ "\n",
+ "import azureml.core\n",
+ "from azureml.core.experiment import Experiment\n",
+ "from azureml.core.workspace import Workspace\n",
+ "from azureml.train.automl import AutoMLConfig"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "ws = Workspace.from_config()\n",
+ "\n",
+ "# Choose a name for the experiment.\n",
+ "experiment_name = 'automl-local-missing-data'\n",
+ "project_folder = './sample_projects/automl-local-missing-data'\n",
+ "\n",
+ "experiment = Experiment(ws, experiment_name)\n",
+ "\n",
+ "output = {}\n",
+ "output['SDK version'] = azureml.core.VERSION\n",
+ "output['Subscription ID'] = ws.subscription_id\n",
+ "output['Workspace'] = ws.name\n",
+ "output['Resource Group'] = ws.resource_group\n",
+ "output['Location'] = ws.location\n",
+ "output['Project Directory'] = project_folder\n",
+ "output['Experiment Name'] = experiment.name\n",
+ "pd.set_option('display.max_colwidth', -1)\n",
+ "outputDf = pd.DataFrame(data = output, index = [''])\n",
+ "outputDf.T"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Opt-in diagnostics for better experience, quality, and security of future releases."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.telemetry import set_diagnostics_collection\n",
+ "set_diagnostics_collection(send_diagnostics = True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Data"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "digits = datasets.load_digits()\n",
+ "X_train = digits.data[10:,:]\n",
+ "y_train = digits.target[10:]\n",
+ "\n",
+ "# Add missing values in 75% of the lines.\n",
+ "missing_rate = 0.75\n",
+ "n_missing_samples = int(np.floor(X_train.shape[0] * missing_rate))\n",
+ "missing_samples = np.hstack((np.zeros(X_train.shape[0] - n_missing_samples, dtype=np.bool), np.ones(n_missing_samples, dtype=np.bool)))\n",
+ "rng = np.random.RandomState(0)\n",
+ "rng.shuffle(missing_samples)\n",
+ "missing_features = rng.randint(0, X_train.shape[1], n_missing_samples)\n",
+ "X_train[np.where(missing_samples)[0], missing_features] = np.nan"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "df = pd.DataFrame(data = X_train)\n",
+ "df['Label'] = pd.Series(y_train, index=df.index)\n",
+ "df.head()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Train\n",
+ "\n",
+ "Instantiate an `AutoMLConfig` object to specify the settings and data used to run the experiment. This includes setting `experiment_exit_score`, which should cause the run to complete before the `iterations` count is reached.\n",
+ "\n",
+ "|Property|Description|\n",
+ "|-|-|\n",
+ "|**task**|classification or regression|\n",
+ "|**primary_metric**|This is the metric that you want to optimize. Classification supports the following primary metrics:
accuracy
AUC_weighted
average_precision_score_weighted
norm_macro_recall
precision_score_weighted|\n",
+ "|**iteration_timeout_minutes**|Time limit in minutes for each iteration.|\n",
+ "|**iterations**|Number of iterations. In each iteration AutoML trains a specific pipeline with the data.|\n",
+ "|**n_cross_validations**|Number of cross validation splits.|\n",
+ "|**preprocess**|Setting this to *True* enables AutoML to perform preprocessing on the input to handle *missing data*, and to perform some common *feature extraction*.|\n",
+ "|**experiment_exit_score**|*double* value indicating the target for *primary_metric*.
Once the target is surpassed the run terminates.|\n",
+ "|**blacklist_models**|*List* of *strings* indicating machine learning algorithms for AutoML to avoid in this run.
Allowed values for **Classification**
LogisticRegression
SGD
MultinomialNaiveBayes
BernoulliNaiveBayes
SVM
LinearSVM
KNN
DecisionTree
RandomForest
ExtremeRandomTrees
LightGBM
GradientBoosting
TensorFlowDNN
TensorFlowLinearClassifier
Allowed values for **Regression**
ElasticNet
GradientBoosting
DecisionTree
KNN
LassoLars
SGD
RandomForest
ExtremeRandomTrees
LightGBM
TensorFlowLinearRegressor
TensorFlowDNN|\n",
+ "|**X**|(sparse) array-like, shape = [n_samples, n_features]|\n",
+ "|**y**|(sparse) array-like, shape = [n_samples, ], [n_samples, n_classes]
Multi-class targets. An indicator matrix turns on multilabel classification. This should be an array of integers.|\n",
+ "|**path**|Relative path to the project folder. AutoML stores configuration files for the experiment under this folder. You can specify a new empty folder.|"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "automl_config = AutoMLConfig(task = 'classification',\n",
+ " debug_log = 'automl_errors.log',\n",
+ " primary_metric = 'AUC_weighted',\n",
+ " iteration_timeout_minutes = 60,\n",
+ " iterations = 20,\n",
+ " n_cross_validations = 5,\n",
+ " preprocess = True,\n",
+ " experiment_exit_score = 0.9984,\n",
+ " blacklist_models = ['KNN','LinearSVM'],\n",
+ " verbosity = logging.INFO,\n",
+ " X = X_train, \n",
+ " y = y_train,\n",
+ " path = project_folder)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Call the `submit` method on the experiment object and pass the run configuration. Execution of local runs is synchronous. Depending on the data and the number of iterations this can run for a while.\n",
+ "In this example, we specify `show_output = True` to print currently running iterations to the console."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "local_run = experiment.submit(automl_config, show_output = True)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "local_run"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Results"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Widget for Monitoring Runs\n",
+ "\n",
+ "The widget will first report a \"loading\" status while running the first iteration. After completing the first iteration, an auto-updating graph and table will be shown. The widget will refresh once per minute, so you should see the graph update as child runs complete.\n",
+ "\n",
+ "**Note:** The widget displays a link at the bottom. Use this link to open a web interface to explore the individual run details."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.widgets import RunDetails\n",
+ "RunDetails(local_run).show() "
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "\n",
+ "#### Retrieve All Child Runs\n",
+ "You can also use SDK methods to fetch all the child runs and see individual metrics that we log."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "children = list(local_run.get_children())\n",
+ "metricslist = {}\n",
+ "for run in children:\n",
+ " properties = run.get_properties()\n",
+ " metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)}\n",
+ " metricslist[int(properties['iteration'])] = metrics\n",
+ "\n",
+ "rundata = pd.DataFrame(metricslist).sort_index(1)\n",
+ "rundata"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Retrieve the Best Model\n",
+ "\n",
+ "Below we select the best pipeline from our iterations. The `get_output` method returns the best run and the fitted model. The Model includes the pipeline and any pre-processing. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "best_run, fitted_model = local_run.get_output()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Best Model Based on Any Other Metric\n",
+ "Show the run and the model which has the smallest `accuracy` value:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# lookup_metric = \"accuracy\"\n",
+ "# best_run, fitted_model = local_run.get_output(metric = lookup_metric)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Model from a Specific Iteration\n",
+ "Show the run and the model from the third iteration:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# iteration = 3\n",
+ "# best_run, fitted_model = local_run.get_output(iteration = iteration)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Test"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "digits = datasets.load_digits()\n",
+ "X_test = digits.data[:10, :]\n",
+ "y_test = digits.target[:10]\n",
+ "images = digits.images[:10]\n",
+ "\n",
+ "# Randomly select digits and test.\n",
+ "for index in np.random.choice(len(y_test), 2, replace = False):\n",
+ " print(index)\n",
+ " predicted = fitted_model.predict(X_test[index:index + 1])[0]\n",
+ " label = y_test[index]\n",
+ " title = \"Label value = %d Predicted value = %d \" % (label, predicted)\n",
+ " fig = plt.figure(1, figsize=(3,3))\n",
+ " ax1 = fig.add_axes((0,0,.8,.8))\n",
+ " ax1.set_title(title)\n",
+ " plt.imshow(images[index], cmap = plt.cm.gray_r, interpolation = 'nearest')\n",
+ " plt.show()\n"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "savitam"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.6"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/automated-machine-learning/model-explanation/auto-ml-model-explanation.ipynb b/how-to-use-azureml/automated-machine-learning/model-explanation/auto-ml-model-explanation.ipynb
index 94568b15..e4df9d6b 100644
--- a/how-to-use-azureml/automated-machine-learning/model-explanation/auto-ml-model-explanation.ipynb
+++ b/how-to-use-azureml/automated-machine-learning/model-explanation/auto-ml-model-explanation.ipynb
@@ -1,367 +1,365 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Automated Machine Learning\n",
- "_**Explain classification model and visualize the explanation**_\n",
- "\n",
- "## Contents\n",
- "1. [Introduction](#Introduction)\n",
- "1. [Setup](#Setup)\n",
- "1. [Data](#Data)\n",
- "1. [Train](#Train)\n",
- "1. [Results](#Results)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Introduction\n",
- "In this example we use the sklearn's [iris dataset](http://scikit-learn.org/stable/modules/generated/sklearn.datasets.load_iris.html) to showcase how you can use the AutoML Classifier for a simple classification problem.\n",
- "\n",
- "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
- "\n",
- "In this notebook you would see\n",
- "1. Creating an Experiment in an existing Workspace\n",
- "2. Instantiating AutoMLConfig\n",
- "3. Training the Model using local compute and explain the model\n",
- "4. Visualization model's feature importance in widget\n",
- "5. Explore best model's explanation"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Setup\n",
- "\n",
- "As part of the setup you have already created a Workspace. For AutoML you would need to create an Experiment. An Experiment is a named object in a Workspace, which is used to run experiments."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import logging\n",
- "import os\n",
- "import random\n",
- "\n",
- "import pandas as pd\n",
- "import azureml.core\n",
- "from azureml.core.experiment import Experiment\n",
- "from azureml.core.workspace import Workspace\n",
- "from azureml.train.automl import AutoMLConfig\n",
- "from azureml.train.automl.run import AutoMLRun"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "ws = Workspace.from_config()\n",
- "\n",
- "# choose a name for experiment\n",
- "experiment_name = 'automl-local-classification'\n",
- "# project folder\n",
- "project_folder = './sample_projects/automl-local-classification-model-explanation'\n",
- "\n",
- "experiment=Experiment(ws, experiment_name)\n",
- "\n",
- "output = {}\n",
- "output['SDK version'] = azureml.core.VERSION\n",
- "output['Subscription ID'] = ws.subscription_id\n",
- "output['Workspace Name'] = ws.name\n",
- "output['Resource Group'] = ws.resource_group\n",
- "output['Location'] = ws.location\n",
- "output['Project Directory'] = project_folder\n",
- "output['Experiment Name'] = experiment.name\n",
- "pd.set_option('display.max_colwidth', -1)\n",
- "pd.DataFrame(data = output, index = ['']).T"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Opt-in diagnostics for better experience, quality, and security of future releases"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.telemetry import set_diagnostics_collection\n",
- "set_diagnostics_collection(send_diagnostics=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Data"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from sklearn import datasets\n",
- "\n",
- "iris = datasets.load_iris()\n",
- "y = iris.target\n",
- "X = iris.data\n",
- "\n",
- "features = iris.feature_names\n",
- "\n",
- "from sklearn.model_selection import train_test_split\n",
- "X_train, X_test, y_train, y_test = train_test_split(X,\n",
- " y,\n",
- " test_size=0.1,\n",
- " random_state=100,\n",
- " stratify=y)\n",
- "\n",
- "X_train = pd.DataFrame(X_train, columns=features)\n",
- "X_test = pd.DataFrame(X_test, columns=features)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Train\n",
- "\n",
- "Instantiate a AutoMLConfig object. This defines the settings and data used to run the experiment.\n",
- "\n",
- "|Property|Description|\n",
- "|-|-|\n",
- "|**task**|classification or regression|\n",
- "|**primary_metric**|This is the metric that you want to optimize. Classification supports the following primary metrics:
accuracy
AUC_weighted
average_precision_score_weighted
norm_macro_recall
precision_score_weighted|\n",
- "|**max_time_sec**|Time limit in minutes for each iterations|\n",
- "|**iterations**|Number of iterations. In each iteration Auto ML trains the data with a specific pipeline|\n",
- "|**X**|(sparse) array-like, shape = [n_samples, n_features]|\n",
- "|**y**|(sparse) array-like, shape = [n_samples, ], [n_samples, n_classes]
Multi-class targets. An indicator matrix turns on multilabel classification. This should be an array of integers. |\n",
- "|**X_valid**|(sparse) array-like, shape = [n_samples, n_features]|\n",
- "|**y_valid**|(sparse) array-like, shape = [n_samples, ], [n_samples, n_classes]|\n",
- "|**model_explainability**|Indicate to explain each trained pipeline or not |\n",
- "|**path**|Relative path to the project folder. AutoML stores configuration files for the experiment under this folder. You can specify a new empty folder. |"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "automl_config = AutoMLConfig(task = 'classification',\n",
- " debug_log = 'automl_errors.log',\n",
- " primary_metric = 'AUC_weighted',\n",
- " iteration_timeout_minutes = 200,\n",
- " iterations = 10,\n",
- " verbosity = logging.INFO,\n",
- " X = X_train, \n",
- " y = y_train,\n",
- " X_valid = X_test,\n",
- " y_valid = y_test,\n",
- " model_explainability=True,\n",
- " path=project_folder)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "You can call the submit method on the experiment object and pass the run configuration. For Local runs the execution is synchronous. Depending on the data and number of iterations this can run for while.\n",
- "You will see the currently running iterations printing to the console."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "local_run = experiment.submit(automl_config, show_output=True)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "local_run"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Results"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Widget for monitoring runs\n",
- "\n",
- "The widget will sit on \"loading\" until the first iteration completed, then you will see an auto-updating graph and table show up. It refreshed once per minute, so you should see the graph update as child runs complete.\n",
- "\n",
- "NOTE: The widget displays a link at the bottom. This links to a web-ui to explore the individual run details."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.widgets import RunDetails\n",
- "RunDetails(local_run).show() "
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Retrieve the Best Model\n",
- "\n",
- "Below we select the best pipeline from our iterations. The *get_output* method on automl_classifier returns the best run and the fitted model for the last *fit* invocation. There are overloads on *get_output* that allow you to retrieve the best run and fitted model for *any* logged metric or a particular *iteration*."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "best_run, fitted_model = local_run.get_output()\n",
- "print(best_run)\n",
- "print(fitted_model)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Best Model 's explanation\n",
- "\n",
- "Retrieve the explanation from the best_run. And explanation information includes:\n",
- "\n",
- "1.\tshap_values: The explanation information generated by shap lib\n",
- "2.\texpected_values: The expected value of the model applied to set of X_train data.\n",
- "3.\toverall_summary: The model level feature importance values sorted in descending order\n",
- "4.\toverall_imp: The feature names sorted in the same order as in overall_summary\n",
- "5.\tper_class_summary: The class level feature importance values sorted in descending order. Only available for the classification case\n",
- "6.\tper_class_imp: The feature names sorted in the same order as in per_class_summary. Only available for the classification case"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.train.automl.automlexplainer import retrieve_model_explanation\n",
- "\n",
- "shap_values, expected_values, overall_summary, overall_imp, per_class_summary, per_class_imp = \\\n",
- " retrieve_model_explanation(best_run)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "print(overall_summary)\n",
- "print(overall_imp)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "print(per_class_summary)\n",
- "print(per_class_imp)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Beside retrieve the existed model explanation information, explain the model with different train/test data"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.train.automl.automlexplainer import explain_model\n",
- "\n",
- "shap_values, expected_values, overall_summary, overall_imp, per_class_summary, per_class_imp = \\\n",
- " explain_model(fitted_model, X_train, X_test)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "print(overall_summary)\n",
- "print(overall_imp)"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "xif"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Automated Machine Learning\n",
+ "_**Explain classification model and visualize the explanation**_\n",
+ "\n",
+ "## Contents\n",
+ "1. [Introduction](#Introduction)\n",
+ "1. [Setup](#Setup)\n",
+ "1. [Data](#Data)\n",
+ "1. [Train](#Train)\n",
+ "1. [Results](#Results)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Introduction\n",
+ "In this example we use the sklearn's [iris dataset](http://scikit-learn.org/stable/modules/generated/sklearn.datasets.load_iris.html) to showcase how you can use the AutoML Classifier for a simple classification problem.\n",
+ "\n",
+ "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
+ "\n",
+ "In this notebook you would see\n",
+ "1. Creating an Experiment in an existing Workspace\n",
+ "2. Instantiating AutoMLConfig\n",
+ "3. Training the Model using local compute and explain the model\n",
+ "4. Visualization model's feature importance in widget\n",
+ "5. Explore best model's explanation"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Setup\n",
+ "\n",
+ "As part of the setup you have already created a Workspace. For AutoML you would need to create an Experiment. An Experiment is a named object in a Workspace, which is used to run experiments."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import logging\n",
+ "\n",
+ "import pandas as pd\n",
+ "import azureml.core\n",
+ "from azureml.core.experiment import Experiment\n",
+ "from azureml.core.workspace import Workspace\n",
+ "from azureml.train.automl import AutoMLConfig"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "ws = Workspace.from_config()\n",
+ "\n",
+ "# choose a name for experiment\n",
+ "experiment_name = 'automl-local-classification'\n",
+ "# project folder\n",
+ "project_folder = './sample_projects/automl-local-classification-model-explanation'\n",
+ "\n",
+ "experiment=Experiment(ws, experiment_name)\n",
+ "\n",
+ "output = {}\n",
+ "output['SDK version'] = azureml.core.VERSION\n",
+ "output['Subscription ID'] = ws.subscription_id\n",
+ "output['Workspace Name'] = ws.name\n",
+ "output['Resource Group'] = ws.resource_group\n",
+ "output['Location'] = ws.location\n",
+ "output['Project Directory'] = project_folder\n",
+ "output['Experiment Name'] = experiment.name\n",
+ "pd.set_option('display.max_colwidth', -1)\n",
+ "outputDf = pd.DataFrame(data = output, index = [''])\n",
+ "outputDf.T"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Opt-in diagnostics for better experience, quality, and security of future releases"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.telemetry import set_diagnostics_collection\n",
+ "set_diagnostics_collection(send_diagnostics=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Data"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from sklearn import datasets\n",
+ "\n",
+ "iris = datasets.load_iris()\n",
+ "y = iris.target\n",
+ "X = iris.data\n",
+ "\n",
+ "features = iris.feature_names\n",
+ "\n",
+ "from sklearn.model_selection import train_test_split\n",
+ "X_train, X_test, y_train, y_test = train_test_split(X,\n",
+ " y,\n",
+ " test_size=0.1,\n",
+ " random_state=100,\n",
+ " stratify=y)\n",
+ "\n",
+ "X_train = pd.DataFrame(X_train, columns=features)\n",
+ "X_test = pd.DataFrame(X_test, columns=features)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Train\n",
+ "\n",
+ "Instantiate a AutoMLConfig object. This defines the settings and data used to run the experiment.\n",
+ "\n",
+ "|Property|Description|\n",
+ "|-|-|\n",
+ "|**task**|classification or regression|\n",
+ "|**primary_metric**|This is the metric that you want to optimize. Classification supports the following primary metrics:
accuracy
AUC_weighted
average_precision_score_weighted
norm_macro_recall
precision_score_weighted|\n",
+ "|**max_time_sec**|Time limit in minutes for each iterations|\n",
+ "|**iterations**|Number of iterations. In each iteration Auto ML trains the data with a specific pipeline|\n",
+ "|**X**|(sparse) array-like, shape = [n_samples, n_features]|\n",
+ "|**y**|(sparse) array-like, shape = [n_samples, ], [n_samples, n_classes]
Multi-class targets. An indicator matrix turns on multilabel classification. This should be an array of integers. |\n",
+ "|**X_valid**|(sparse) array-like, shape = [n_samples, n_features]|\n",
+ "|**y_valid**|(sparse) array-like, shape = [n_samples, ], [n_samples, n_classes]|\n",
+ "|**model_explainability**|Indicate to explain each trained pipeline or not |\n",
+ "|**path**|Relative path to the project folder. AutoML stores configuration files for the experiment under this folder. You can specify a new empty folder. |"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "automl_config = AutoMLConfig(task = 'classification',\n",
+ " debug_log = 'automl_errors.log',\n",
+ " primary_metric = 'AUC_weighted',\n",
+ " iteration_timeout_minutes = 200,\n",
+ " iterations = 10,\n",
+ " verbosity = logging.INFO,\n",
+ " X = X_train, \n",
+ " y = y_train,\n",
+ " X_valid = X_test,\n",
+ " y_valid = y_test,\n",
+ " model_explainability=True,\n",
+ " path=project_folder)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "You can call the submit method on the experiment object and pass the run configuration. For Local runs the execution is synchronous. Depending on the data and number of iterations this can run for while.\n",
+ "You will see the currently running iterations printing to the console."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "local_run = experiment.submit(automl_config, show_output=True)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "local_run"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Results"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Widget for monitoring runs\n",
+ "\n",
+ "The widget will sit on \"loading\" until the first iteration completed, then you will see an auto-updating graph and table show up. It refreshed once per minute, so you should see the graph update as child runs complete.\n",
+ "\n",
+ "NOTE: The widget displays a link at the bottom. This links to a web-ui to explore the individual run details."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.widgets import RunDetails\n",
+ "RunDetails(local_run).show() "
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Retrieve the Best Model\n",
+ "\n",
+ "Below we select the best pipeline from our iterations. The *get_output* method on automl_classifier returns the best run and the fitted model for the last *fit* invocation. There are overloads on *get_output* that allow you to retrieve the best run and fitted model for *any* logged metric or a particular *iteration*."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "best_run, fitted_model = local_run.get_output()\n",
+ "print(best_run)\n",
+ "print(fitted_model)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Best Model 's explanation\n",
+ "\n",
+ "Retrieve the explanation from the best_run. And explanation information includes:\n",
+ "\n",
+ "1.\tshap_values: The explanation information generated by shap lib\n",
+ "2.\texpected_values: The expected value of the model applied to set of X_train data.\n",
+ "3.\toverall_summary: The model level feature importance values sorted in descending order\n",
+ "4.\toverall_imp: The feature names sorted in the same order as in overall_summary\n",
+ "5.\tper_class_summary: The class level feature importance values sorted in descending order. Only available for the classification case\n",
+ "6.\tper_class_imp: The feature names sorted in the same order as in per_class_summary. Only available for the classification case"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.train.automl.automlexplainer import retrieve_model_explanation\n",
+ "\n",
+ "shap_values, expected_values, overall_summary, overall_imp, per_class_summary, per_class_imp = \\\n",
+ " retrieve_model_explanation(best_run)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "print(overall_summary)\n",
+ "print(overall_imp)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "print(per_class_summary)\n",
+ "print(per_class_imp)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Beside retrieve the existed model explanation information, explain the model with different train/test data"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.train.automl.automlexplainer import explain_model\n",
+ "\n",
+ "shap_values, expected_values, overall_summary, overall_imp, per_class_summary, per_class_imp = \\\n",
+ " explain_model(fitted_model, X_train, X_test)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "print(overall_summary)\n",
+ "print(overall_imp)"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "xif"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.6"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/automated-machine-learning/regression/auto-ml-regression.ipynb b/how-to-use-azureml/automated-machine-learning/regression/auto-ml-regression.ipynb
index b8ba4fac..4b7750cf 100644
--- a/how-to-use-azureml/automated-machine-learning/regression/auto-ml-regression.ipynb
+++ b/how-to-use-azureml/automated-machine-learning/regression/auto-ml-regression.ipynb
@@ -1,424 +1,417 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Automated Machine Learning\n",
- "_**Regression with Local Compute**_\n",
- "\n",
- "## Contents\n",
- "1. [Introduction](#Introduction)\n",
- "1. [Setup](#Setup)\n",
- "1. [Data](#Data)\n",
- "1. [Train](#Train)\n",
- "1. [Results](#Results)\n",
- "1. [Test](#Test)\n"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Introduction\n",
- "In this example we use the scikit-learn's [diabetes dataset](http://scikit-learn.org/stable/datasets/index.html#diabetes-dataset) to showcase how you can use AutoML for a simple regression problem.\n",
- "\n",
- "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
- "\n",
- "In this notebook you will learn how to:\n",
- "1. Create an `Experiment` in an existing `Workspace`.\n",
- "2. Configure AutoML using `AutoMLConfig`.\n",
- "3. Train the model using local compute.\n",
- "4. Explore the results.\n",
- "5. Test the best fitted model."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Setup\n",
- "\n",
- "As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import logging\n",
- "import os\n",
- "import random\n",
- "\n",
- "from matplotlib import pyplot as plt\n",
- "from matplotlib.pyplot import imshow\n",
- "import numpy as np\n",
- "import pandas as pd\n",
- "from sklearn import datasets\n",
- "\n",
- "import azureml.core\n",
- "from azureml.core.experiment import Experiment\n",
- "from azureml.core.workspace import Workspace\n",
- "from azureml.train.automl import AutoMLConfig\n",
- "from azureml.train.automl.run import AutoMLRun"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "ws = Workspace.from_config()\n",
- "\n",
- "# Choose a name for the experiment and specify the project folder.\n",
- "experiment_name = 'automl-local-regression'\n",
- "project_folder = './sample_projects/automl-local-regression'\n",
- "\n",
- "experiment = Experiment(ws, experiment_name)\n",
- "\n",
- "output = {}\n",
- "output['SDK version'] = azureml.core.VERSION\n",
- "output['Subscription ID'] = ws.subscription_id\n",
- "output['Workspace Name'] = ws.name\n",
- "output['Resource Group'] = ws.resource_group\n",
- "output['Location'] = ws.location\n",
- "output['Project Directory'] = project_folder\n",
- "output['Experiment Name'] = experiment.name\n",
- "pd.set_option('display.max_colwidth', -1)\n",
- "pd.DataFrame(data = output, index = ['']).T"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Opt-in diagnostics for better experience, quality, and security of future releases."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.telemetry import set_diagnostics_collection\n",
- "set_diagnostics_collection(send_diagnostics = True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Data\n",
- "This uses scikit-learn's [load_diabetes](http://scikit-learn.org/stable/modules/generated/sklearn.datasets.load_diabetes.html) method."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Load the diabetes dataset, a well-known built-in small dataset that comes with scikit-learn.\n",
- "from sklearn.datasets import load_diabetes\n",
- "from sklearn.model_selection import train_test_split\n",
- "\n",
- "X, y = load_diabetes(return_X_y = True)\n",
- "\n",
- "columns = ['age', 'gender', 'bmi', 'bp', 's1', 's2', 's3', 's4', 's5', 's6']\n",
- "\n",
- "X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2, random_state = 0)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Train\n",
- "\n",
- "Instantiate an `AutoMLConfig` object to specify the settings and data used to run the experiment.\n",
- "\n",
- "|Property|Description|\n",
- "|-|-|\n",
- "|**task**|classification or regression|\n",
- "|**primary_metric**|This is the metric that you want to optimize. Regression supports the following primary metrics:
spearman_correlation
normalized_root_mean_squared_error
r2_score
normalized_mean_absolute_error|\n",
- "|**iteration_timeout_minutes**|Time limit in minutes for each iteration.|\n",
- "|**iterations**|Number of iterations. In each iteration AutoML trains a specific pipeline with the data.|\n",
- "|**n_cross_validations**|Number of cross validation splits.|\n",
- "|**X**|(sparse) array-like, shape = [n_samples, n_features]|\n",
- "|**y**|(sparse) array-like, shape = [n_samples, ], [n_samples, n_classes]
Multi-class targets. An indicator matrix turns on multilabel classification. This should be an array of integers.|\n",
- "|**path**|Relative path to the project folder. AutoML stores configuration files for the experiment under this folder. You can specify a new empty folder.|"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "automl_config = AutoMLConfig(task = 'regression',\n",
- " iteration_timeout_minutes = 10,\n",
- " iterations = 10,\n",
- " primary_metric = 'spearman_correlation',\n",
- " n_cross_validations = 5,\n",
- " debug_log = 'automl.log',\n",
- " verbosity = logging.INFO,\n",
- " X = X_train, \n",
- " y = y_train,\n",
- " path = project_folder)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Call the `submit` method on the experiment object and pass the run configuration. Execution of local runs is synchronous. Depending on the data and the number of iterations this can run for a while.\n",
- "In this example, we specify `show_output = True` to print currently running iterations to the console."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "local_run = experiment.submit(automl_config, show_output = True)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "local_run"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Results"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Widget for Monitoring Runs\n",
- "\n",
- "The widget will first report a \"loading\" status while running the first iteration. After completing the first iteration, an auto-updating graph and table will be shown. The widget will refresh once per minute, so you should see the graph update as child runs complete.\n",
- "\n",
- "**Note:** The widget displays a link at the bottom. Use this link to open a web interface to explore the individual run details."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.widgets import RunDetails\n",
- "RunDetails(local_run).show() "
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "\n",
- "#### Retrieve All Child Runs\n",
- "You can also use SDK methods to fetch all the child runs and see individual metrics that we log."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "children = list(local_run.get_children())\n",
- "metricslist = {}\n",
- "for run in children:\n",
- " properties = run.get_properties()\n",
- " metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)}\n",
- " metricslist[int(properties['iteration'])] = metrics\n",
- "\n",
- "rundata = pd.DataFrame(metricslist).sort_index(1)\n",
- "rundata"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Retrieve the Best Model\n",
- "\n",
- "Below we select the best pipeline from our iterations. The `get_output` method returns the best run and the fitted model. The Model includes the pipeline and any pre-processing. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "best_run, fitted_model = local_run.get_output()\n",
- "print(best_run)\n",
- "print(fitted_model)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Best Model Based on Any Other Metric\n",
- "Show the run and the model that has the smallest `root_mean_squared_error` value (which turned out to be the same as the one with largest `spearman_correlation` value):"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "lookup_metric = \"root_mean_squared_error\"\n",
- "best_run, fitted_model = local_run.get_output(metric = lookup_metric)\n",
- "print(best_run)\n",
- "print(fitted_model)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Model from a Specific Iteration\n",
- "Show the run and the model from the third iteration:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "iteration = 3\n",
- "third_run, third_model = local_run.get_output(iteration = iteration)\n",
- "print(third_run)\n",
- "print(third_model)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Test"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Predict on training and test set, and calculate residual values."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "y_pred_train = fitted_model.predict(X_train)\n",
- "y_residual_train = y_train - y_pred_train\n",
- "\n",
- "y_pred_test = fitted_model.predict(X_test)\n",
- "y_residual_test = y_test - y_pred_test"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%matplotlib inline\n",
- "import matplotlib.pyplot as plt\n",
- "import numpy as np\n",
- "from sklearn import datasets\n",
- "from sklearn.metrics import mean_squared_error, r2_score\n",
- "\n",
- "# Set up a multi-plot chart.\n",
- "f, (a0, a1) = plt.subplots(1, 2, gridspec_kw = {'width_ratios':[1, 1], 'wspace':0, 'hspace': 0})\n",
- "f.suptitle('Regression Residual Values', fontsize = 18)\n",
- "f.set_figheight(6)\n",
- "f.set_figwidth(16)\n",
- "\n",
- "# Plot residual values of training set.\n",
- "a0.axis([0, 360, -200, 200])\n",
- "a0.plot(y_residual_train, 'bo', alpha = 0.5)\n",
- "a0.plot([-10,360],[0,0], 'r-', lw = 3)\n",
- "a0.text(16,170,'RMSE = {0:.2f}'.format(np.sqrt(mean_squared_error(y_train, y_pred_train))), fontsize = 12)\n",
- "a0.text(16,140,'R2 score = {0:.2f}'.format(r2_score(y_train, y_pred_train)), fontsize = 12)\n",
- "a0.set_xlabel('Training samples', fontsize = 12)\n",
- "a0.set_ylabel('Residual Values', fontsize = 12)\n",
- "\n",
- "# Plot a histogram.\n",
- "a0.hist(y_residual_train, orientation = 'horizontal', color = 'b', bins = 10, histtype = 'step');\n",
- "a0.hist(y_residual_train, orientation = 'horizontal', color = 'b', alpha = 0.2, bins = 10);\n",
- "\n",
- "# Plot residual values of test set.\n",
- "a1.axis([0, 90, -200, 200])\n",
- "a1.plot(y_residual_test, 'bo', alpha = 0.5)\n",
- "a1.plot([-10,360],[0,0], 'r-', lw = 3)\n",
- "a1.text(5,170,'RMSE = {0:.2f}'.format(np.sqrt(mean_squared_error(y_test, y_pred_test))), fontsize = 12)\n",
- "a1.text(5,140,'R2 score = {0:.2f}'.format(r2_score(y_test, y_pred_test)), fontsize = 12)\n",
- "a1.set_xlabel('Test samples', fontsize = 12)\n",
- "a1.set_yticklabels([])\n",
- "\n",
- "# Plot a histogram.\n",
- "a1.hist(y_residual_test, orientation = 'horizontal', color = 'b', bins = 10, histtype = 'step')\n",
- "a1.hist(y_residual_test, orientation = 'horizontal', color = 'b', alpha = 0.2, bins = 10)\n",
- "\n",
- "plt.show()"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "savitam"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Automated Machine Learning\n",
+ "_**Regression with Local Compute**_\n",
+ "\n",
+ "## Contents\n",
+ "1. [Introduction](#Introduction)\n",
+ "1. [Setup](#Setup)\n",
+ "1. [Data](#Data)\n",
+ "1. [Train](#Train)\n",
+ "1. [Results](#Results)\n",
+ "1. [Test](#Test)\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Introduction\n",
+ "In this example we use the scikit-learn's [diabetes dataset](http://scikit-learn.org/stable/datasets/index.html#diabetes-dataset) to showcase how you can use AutoML for a simple regression problem.\n",
+ "\n",
+ "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
+ "\n",
+ "In this notebook you will learn how to:\n",
+ "1. Create an `Experiment` in an existing `Workspace`.\n",
+ "2. Configure AutoML using `AutoMLConfig`.\n",
+ "3. Train the model using local compute.\n",
+ "4. Explore the results.\n",
+ "5. Test the best fitted model."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Setup\n",
+ "\n",
+ "As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import logging\n",
+ "\n",
+ "from matplotlib import pyplot as plt\n",
+ "import numpy as np\n",
+ "import pandas as pd\n",
+ "\n",
+ "import azureml.core\n",
+ "from azureml.core.experiment import Experiment\n",
+ "from azureml.core.workspace import Workspace\n",
+ "from azureml.train.automl import AutoMLConfig"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "ws = Workspace.from_config()\n",
+ "\n",
+ "# Choose a name for the experiment and specify the project folder.\n",
+ "experiment_name = 'automl-local-regression'\n",
+ "project_folder = './sample_projects/automl-local-regression'\n",
+ "\n",
+ "experiment = Experiment(ws, experiment_name)\n",
+ "\n",
+ "output = {}\n",
+ "output['SDK version'] = azureml.core.VERSION\n",
+ "output['Subscription ID'] = ws.subscription_id\n",
+ "output['Workspace Name'] = ws.name\n",
+ "output['Resource Group'] = ws.resource_group\n",
+ "output['Location'] = ws.location\n",
+ "output['Project Directory'] = project_folder\n",
+ "output['Experiment Name'] = experiment.name\n",
+ "pd.set_option('display.max_colwidth', -1)\n",
+ "outputDf = pd.DataFrame(data = output, index = [''])\n",
+ "outputDf.T"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Opt-in diagnostics for better experience, quality, and security of future releases."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.telemetry import set_diagnostics_collection\n",
+ "set_diagnostics_collection(send_diagnostics = True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Data\n",
+ "This uses scikit-learn's [load_diabetes](http://scikit-learn.org/stable/modules/generated/sklearn.datasets.load_diabetes.html) method."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Load the diabetes dataset, a well-known built-in small dataset that comes with scikit-learn.\n",
+ "from sklearn.datasets import load_diabetes\n",
+ "from sklearn.model_selection import train_test_split\n",
+ "\n",
+ "X, y = load_diabetes(return_X_y = True)\n",
+ "\n",
+ "columns = ['age', 'gender', 'bmi', 'bp', 's1', 's2', 's3', 's4', 's5', 's6']\n",
+ "\n",
+ "X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2, random_state = 0)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Train\n",
+ "\n",
+ "Instantiate an `AutoMLConfig` object to specify the settings and data used to run the experiment.\n",
+ "\n",
+ "|Property|Description|\n",
+ "|-|-|\n",
+ "|**task**|classification or regression|\n",
+ "|**primary_metric**|This is the metric that you want to optimize. Regression supports the following primary metrics:
spearman_correlation
normalized_root_mean_squared_error
r2_score
normalized_mean_absolute_error|\n",
+ "|**iteration_timeout_minutes**|Time limit in minutes for each iteration.|\n",
+ "|**iterations**|Number of iterations. In each iteration AutoML trains a specific pipeline with the data.|\n",
+ "|**n_cross_validations**|Number of cross validation splits.|\n",
+ "|**X**|(sparse) array-like, shape = [n_samples, n_features]|\n",
+ "|**y**|(sparse) array-like, shape = [n_samples, ], [n_samples, n_classes]
Multi-class targets. An indicator matrix turns on multilabel classification. This should be an array of integers.|\n",
+ "|**path**|Relative path to the project folder. AutoML stores configuration files for the experiment under this folder. You can specify a new empty folder.|"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "automl_config = AutoMLConfig(task = 'regression',\n",
+ " iteration_timeout_minutes = 10,\n",
+ " iterations = 10,\n",
+ " primary_metric = 'spearman_correlation',\n",
+ " n_cross_validations = 5,\n",
+ " debug_log = 'automl.log',\n",
+ " verbosity = logging.INFO,\n",
+ " X = X_train, \n",
+ " y = y_train,\n",
+ " path = project_folder)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Call the `submit` method on the experiment object and pass the run configuration. Execution of local runs is synchronous. Depending on the data and the number of iterations this can run for a while.\n",
+ "In this example, we specify `show_output = True` to print currently running iterations to the console."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "local_run = experiment.submit(automl_config, show_output = True)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "local_run"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Results"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Widget for Monitoring Runs\n",
+ "\n",
+ "The widget will first report a \"loading\" status while running the first iteration. After completing the first iteration, an auto-updating graph and table will be shown. The widget will refresh once per minute, so you should see the graph update as child runs complete.\n",
+ "\n",
+ "**Note:** The widget displays a link at the bottom. Use this link to open a web interface to explore the individual run details."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.widgets import RunDetails\n",
+ "RunDetails(local_run).show() "
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "\n",
+ "#### Retrieve All Child Runs\n",
+ "You can also use SDK methods to fetch all the child runs and see individual metrics that we log."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "children = list(local_run.get_children())\n",
+ "metricslist = {}\n",
+ "for run in children:\n",
+ " properties = run.get_properties()\n",
+ " metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)}\n",
+ " metricslist[int(properties['iteration'])] = metrics\n",
+ "\n",
+ "rundata = pd.DataFrame(metricslist).sort_index(1)\n",
+ "rundata"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Retrieve the Best Model\n",
+ "\n",
+ "Below we select the best pipeline from our iterations. The `get_output` method returns the best run and the fitted model. The Model includes the pipeline and any pre-processing. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "best_run, fitted_model = local_run.get_output()\n",
+ "print(best_run)\n",
+ "print(fitted_model)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Best Model Based on Any Other Metric\n",
+ "Show the run and the model that has the smallest `root_mean_squared_error` value (which turned out to be the same as the one with largest `spearman_correlation` value):"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "lookup_metric = \"root_mean_squared_error\"\n",
+ "best_run, fitted_model = local_run.get_output(metric = lookup_metric)\n",
+ "print(best_run)\n",
+ "print(fitted_model)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Model from a Specific Iteration\n",
+ "Show the run and the model from the third iteration:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "iteration = 3\n",
+ "third_run, third_model = local_run.get_output(iteration = iteration)\n",
+ "print(third_run)\n",
+ "print(third_model)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Test"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Predict on training and test set, and calculate residual values."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "y_pred_train = fitted_model.predict(X_train)\n",
+ "y_residual_train = y_train - y_pred_train\n",
+ "\n",
+ "y_pred_test = fitted_model.predict(X_test)\n",
+ "y_residual_test = y_test - y_pred_test"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%matplotlib inline\n",
+ "from sklearn.metrics import mean_squared_error, r2_score\n",
+ "\n",
+ "# Set up a multi-plot chart.\n",
+ "f, (a0, a1) = plt.subplots(1, 2, gridspec_kw = {'width_ratios':[1, 1], 'wspace':0, 'hspace': 0})\n",
+ "f.suptitle('Regression Residual Values', fontsize = 18)\n",
+ "f.set_figheight(6)\n",
+ "f.set_figwidth(16)\n",
+ "\n",
+ "# Plot residual values of training set.\n",
+ "a0.axis([0, 360, -200, 200])\n",
+ "a0.plot(y_residual_train, 'bo', alpha = 0.5)\n",
+ "a0.plot([-10,360],[0,0], 'r-', lw = 3)\n",
+ "a0.text(16,170,'RMSE = {0:.2f}'.format(np.sqrt(mean_squared_error(y_train, y_pred_train))), fontsize = 12)\n",
+ "a0.text(16,140,'R2 score = {0:.2f}'.format(r2_score(y_train, y_pred_train)), fontsize = 12)\n",
+ "a0.set_xlabel('Training samples', fontsize = 12)\n",
+ "a0.set_ylabel('Residual Values', fontsize = 12)\n",
+ "\n",
+ "# Plot a histogram.\n",
+ "a0.hist(y_residual_train, orientation = 'horizontal', color = 'b', bins = 10, histtype = 'step')\n",
+ "a0.hist(y_residual_train, orientation = 'horizontal', color = 'b', alpha = 0.2, bins = 10)\n",
+ "\n",
+ "# Plot residual values of test set.\n",
+ "a1.axis([0, 90, -200, 200])\n",
+ "a1.plot(y_residual_test, 'bo', alpha = 0.5)\n",
+ "a1.plot([-10,360],[0,0], 'r-', lw = 3)\n",
+ "a1.text(5,170,'RMSE = {0:.2f}'.format(np.sqrt(mean_squared_error(y_test, y_pred_test))), fontsize = 12)\n",
+ "a1.text(5,140,'R2 score = {0:.2f}'.format(r2_score(y_test, y_pred_test)), fontsize = 12)\n",
+ "a1.set_xlabel('Test samples', fontsize = 12)\n",
+ "a1.set_yticklabels([])\n",
+ "\n",
+ "# Plot a histogram.\n",
+ "a1.hist(y_residual_test, orientation = 'horizontal', color = 'b', bins = 10, histtype = 'step')\n",
+ "a1.hist(y_residual_test, orientation = 'horizontal', color = 'b', alpha = 0.2, bins = 10)\n",
+ "\n",
+ "plt.show()"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "savitam"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.6"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/automated-machine-learning/remote-attach/auto-ml-remote-attach.ipynb b/how-to-use-azureml/automated-machine-learning/remote-attach/auto-ml-remote-attach.ipynb
index 55b72531..9466826c 100644
--- a/how-to-use-azureml/automated-machine-learning/remote-attach/auto-ml-remote-attach.ipynb
+++ b/how-to-use-azureml/automated-machine-learning/remote-attach/auto-ml-remote-attach.ipynb
@@ -1,537 +1,532 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Automated Machine Learning\n",
- "_**Remote Execution using attach**_\n",
- "\n",
- "## Contents\n",
- "1. [Introduction](#Introduction)\n",
- "1. [Setup](#Setup)\n",
- "1. [Data](#Data)\n",
- "1. [Train](#Train)\n",
- "1. [Results](#Results)\n",
- "1. [Test](#Test)\n"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Introduction\n",
- "In this example we use the scikit-learn's [20newsgroup](http://scikit-learn.org/stable/modules/generated/sklearn.datasets.fetch_20newsgroups.html) to showcase how you can use AutoML to handle text data with remote attach.\n",
- "\n",
- "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
- "\n",
- "In this notebook you will learn how to:\n",
- "1. Create an `Experiment` in an existing `Workspace`.\n",
- "2. Attach an existing DSVM to a workspace.\n",
- "3. Configure AutoML using `AutoMLConfig`.\n",
- "4. Train the model using the DSVM.\n",
- "5. Explore the results.\n",
- "6. Test the best fitted model.\n",
- "\n",
- "In addition this notebook showcases the following features\n",
- "- **Parallel** executions for iterations\n",
- "- **Asynchronous** tracking of progress\n",
- "- **Cancellation** of individual iterations or the entire run\n",
- "- Retrieving models for any iteration or logged metric\n",
- "- Specifying AutoML settings as `**kwargs`\n",
- "- Handling **text** data using the `preprocess` flag"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Setup\n",
- "\n",
- "As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import logging\n",
- "import os\n",
- "import random\n",
- "\n",
- "from matplotlib import pyplot as plt\n",
- "from matplotlib.pyplot import imshow\n",
- "import numpy as np\n",
- "import pandas as pd\n",
- "from sklearn import datasets\n",
- "\n",
- "import azureml.core\n",
- "from azureml.core.experiment import Experiment\n",
- "from azureml.core.workspace import Workspace\n",
- "from azureml.train.automl import AutoMLConfig\n",
- "from azureml.train.automl.run import AutoMLRun"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "ws = Workspace.from_config()\n",
- "\n",
- "# Choose a name for the run history container in the workspace.\n",
- "experiment_name = 'automl-remote-attach'\n",
- "project_folder = './sample_projects/automl-remote-attach'\n",
- "\n",
- "experiment = Experiment(ws, experiment_name)\n",
- "\n",
- "output = {}\n",
- "output['SDK version'] = azureml.core.VERSION\n",
- "output['Subscription ID'] = ws.subscription_id\n",
- "output['Workspace'] = ws.name\n",
- "output['Resource Group'] = ws.resource_group\n",
- "output['Location'] = ws.location\n",
- "output['Project Directory'] = project_folder\n",
- "output['Experiment Name'] = experiment.name\n",
- "pd.set_option('display.max_colwidth', -1)\n",
- "pd.DataFrame(data=output, index=['']).T"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Opt-in diagnostics for better experience, quality, and security of future releases."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.telemetry import set_diagnostics_collection\n",
- "set_diagnostics_collection(send_diagnostics = True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Attach a Remote Linux DSVM\n",
- "To use a remote Docker compute target:\n",
- "1. Create a Linux DSVM in Azure, following these [quick instructions](https://docs.microsoft.com/en-us/azure/machine-learning/desktop-workbench/how-to-create-dsvm-hdi). Make sure you use the Ubuntu flavor (not CentOS). Make sure that disk space is available under `/tmp` because AutoML creates files under `/tmp/azureml_run`s. The DSVM should have more cores than the number of parallel runs that you plan to enable. It should also have at least 4GB per core.\n",
- "2. Enter the IP address, user name and password below.\n",
- "\n",
- "**Note:** By default, SSH runs on port 22 and you don't need to change the port number below. If you've configured SSH to use a different port, change `dsvm_ssh_port` accordinglyaddress. [Read more](https://render.githubusercontent.com/documentation/sdk/ssh-issue.md) on changing SSH ports for security reasons."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.compute import ComputeTarget, RemoteCompute\n",
- "import time\n",
- "\n",
- "# Add your VM information below\n",
- "# If a compute with the specified compute_name already exists, it will be used and the dsvm_ip_addr, dsvm_ssh_port, \n",
- "# dsvm_username and dsvm_password will be ignored.\n",
- "compute_name = 'mydsvmb'\n",
- "dsvm_ip_addr = '<>'\n",
- "dsvm_ssh_port = 22\n",
- "dsvm_username = '<>'\n",
- "dsvm_password = '<>'\n",
- "\n",
- "if compute_name in ws.compute_targets:\n",
- " print('Using existing compute.')\n",
- " dsvm_compute = ws.compute_targets[compute_name]\n",
- "else:\n",
- " attach_config = RemoteCompute.attach_configuration(address=dsvm_ip_addr, username=dsvm_username, password=dsvm_password, ssh_port=dsvm_ssh_port)\n",
- " ComputeTarget.attach(workspace=ws, name=compute_name, attach_configuration=attach_config)\n",
- "\n",
- " while ws.compute_targets[compute_name].provisioning_state == 'Creating':\n",
- " time.sleep(1)\n",
- "\n",
- " dsvm_compute = ws.compute_targets[compute_name]\n",
- " \n",
- " if dsvm_compute.provisioning_state == 'Failed':\n",
- " print('Attached failed.')\n",
- " print(dsvm_compute.provisioning_errors)\n",
- " dsvm_compute.detach()"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.runconfig import RunConfiguration\n",
- "from azureml.core.conda_dependencies import CondaDependencies\n",
- "\n",
- "# create a new RunConfig object\n",
- "conda_run_config = RunConfiguration(framework=\"python\")\n",
- "\n",
- "# Set compute target to the Linux DSVM\n",
- "conda_run_config.target = dsvm_compute\n",
- "\n",
- "cd = CondaDependencies.create(pip_packages=['azureml-sdk[automl]'], conda_packages=['numpy'])\n",
- "conda_run_config.environment.python.conda_dependencies = cd"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Data\n",
- "For remote executions you should author a `get_data.py` file containing a `get_data()` function. This file should be in the root directory of the project. You can encapsulate code to read data either from a blob storage or local disk in this file.\n",
- "In this example, the `get_data()` function returns a [dictionary](README.md#getdata)."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "if not os.path.exists(project_folder):\n",
- " os.makedirs(project_folder)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%writefile $project_folder/get_data.py\n",
- "\n",
- "import numpy as np\n",
- "from sklearn.datasets import fetch_20newsgroups\n",
- "\n",
- "def get_data():\n",
- " remove = ('headers', 'footers', 'quotes')\n",
- " categories = [\n",
- " 'alt.atheism',\n",
- " 'talk.religion.misc',\n",
- " 'comp.graphics',\n",
- " 'sci.space',\n",
- " ]\n",
- " data_train = fetch_20newsgroups(subset = 'train', categories = categories,\n",
- " shuffle = True, random_state = 42,\n",
- " remove = remove)\n",
- " \n",
- " X_train = np.array(data_train.data).reshape((len(data_train.data),1))\n",
- " y_train = np.array(data_train.target)\n",
- " \n",
- " return { \"X\" : X_train, \"y\" : y_train }"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Train\n",
- "\n",
- "You can specify `automl_settings` as `**kwargs` as well. Also note that you can use a `get_data()` function for local excutions too.\n",
- "\n",
- "**Note:** When using Remote DSVM, you can't pass Numpy arrays directly to the fit method.\n",
- "\n",
- "|Property|Description|\n",
- "|-|-|\n",
- "|**primary_metric**|This is the metric that you want to optimize. Classification supports the following primary metrics:
accuracy
AUC_weighted
average_precision_score_weighted
norm_macro_recall
precision_score_weighted|\n",
- "|**iteration_timeout_minutes**|Time limit in minutes for each iteration.|\n",
- "|**iterations**|Number of iterations. In each iteration AutoML trains a specific pipeline with the data.|\n",
- "|**n_cross_validations**|Number of cross validation splits.|\n",
- "|**max_concurrent_iterations**|Maximum number of iterations that would be executed in parallel. This should be less than the number of cores on the DSVM.|\n",
- "|**preprocess**|Setting this to *True* enables AutoML to perform preprocessing on the input to handle *missing data*, and to perform some common *feature extraction*.|\n",
- "|**enable_cache**|Setting this to *True* enables preprocess done once and reuse the same preprocessed data for all the iterations. Default value is True.\n",
- "|**max_cores_per_iteration**|Indicates how many cores on the compute target would be used to train a single pipeline.
Default is *1*; you can set it to *-1* to use all cores.|"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "automl_settings = {\n",
- " \"iteration_timeout_minutes\": 60,\n",
- " \"iterations\": 4,\n",
- " \"n_cross_validations\": 5,\n",
- " \"primary_metric\": 'AUC_weighted',\n",
- " \"preprocess\": True,\n",
- " \"max_cores_per_iteration\": 2\n",
- "}\n",
- "\n",
- "automl_config = AutoMLConfig(task = 'classification',\n",
- " path = project_folder,\n",
- " run_configuration=conda_run_config,\n",
- " data_script = project_folder + \"/get_data.py\",\n",
- " **automl_settings\n",
- " )\n"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Call the `submit` method on the experiment object and pass the run configuration. For remote runs the execution is asynchronous, so you will see the iterations get populated as they complete. You can interact with the widgets and models even when the experiment is running to retrieve the best model up to that point. Once you are satisfied with the model, you can cancel a particular iteration or the whole run."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "remote_run = experiment.submit(automl_config)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "remote_run"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Results\n",
- "#### Widget for Monitoring Runs\n",
- "\n",
- "The widget will first report a \"loading\" status while running the first iteration. After completing the first iteration, an auto-updating graph and table will be shown. The widget will refresh once per minute, so you should see the graph update as child runs complete.\n",
- "\n",
- "You can click on a pipeline to see run properties and output logs. Logs are also available on the DSVM under `/tmp/azureml_run/{iterationid}/azureml-logs`\n",
- "\n",
- "**Note:** The widget displays a link at the bottom. Use this link to open a web interface to explore the individual run details."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.widgets import RunDetails\n",
- "RunDetails(remote_run).show() "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Wait until the run finishes.\n",
- "remote_run.wait_for_completion(show_output = True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Pre-process cache cleanup\n",
- "The preprocess data gets cache at user default file store. When the run is completed the cache can be cleaned by running below cell"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "remote_run.clean_preprocessor_cache()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "\n",
- "#### Retrieve All Child Runs\n",
- "You can also use SDK methods to fetch all the child runs and see individual metrics that we log. "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "children = list(remote_run.get_children())\n",
- "metricslist = {}\n",
- "for run in children:\n",
- " properties = run.get_properties()\n",
- " metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)}\n",
- " metricslist[int(properties['iteration'])] = metrics\n",
- "\n",
- "rundata = pd.DataFrame(metricslist).sort_index(1)\n",
- "rundata"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Cancelling Runs\n",
- "You can cancel ongoing remote runs using the `cancel` and `cancel_iteration` functions."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Cancel the ongoing experiment and stop scheduling new iterations.\n",
- "# remote_run.cancel()\n",
- "\n",
- "# Cancel iteration 1 and move onto iteration 2.\n",
- "# remote_run.cancel_iteration(1)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Retrieve the Best Model\n",
- "\n",
- "Below we select the best pipeline from our iterations. The `get_output` method returns the best run and the fitted model. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "best_run, fitted_model = remote_run.get_output()\n",
- "print(best_run)\n",
- "print(fitted_model)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Best Model Based on Any Other Metric\n",
- "Show the run and the model which has the smallest `accuracy` value:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# lookup_metric = \"accuracy\"\n",
- "# best_run, fitted_model = remote_run.get_output(metric = lookup_metric)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Model from a Specific Iteration"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "iteration = 0\n",
- "zero_run, zero_model = remote_run.get_output(iteration = iteration)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Test"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Load test data.\n",
- "from pandas_ml import ConfusionMatrix\n",
- "from sklearn.datasets import fetch_20newsgroups\n",
- "\n",
- "remove = ('headers', 'footers', 'quotes')\n",
- "categories = [\n",
- " 'alt.atheism',\n",
- " 'talk.religion.misc',\n",
- " 'comp.graphics',\n",
- " 'sci.space',\n",
- " ]\n",
- "\n",
- "data_test = fetch_20newsgroups(subset = 'test', categories = categories,\n",
- " shuffle = True, random_state = 42,\n",
- " remove = remove)\n",
- "\n",
- "X_test = np.array(data_test.data).reshape((len(data_test.data),1))\n",
- "y_test = data_test.target\n",
- "\n",
- "# Test our best pipeline.\n",
- "\n",
- "y_pred = fitted_model.predict(X_test)\n",
- "y_pred_strings = [data_test.target_names[i] for i in y_pred]\n",
- "y_test_strings = [data_test.target_names[i] for i in y_test]\n",
- "\n",
- "cm = ConfusionMatrix(y_test_strings, y_pred_strings)\n",
- "print(cm)\n",
- "cm.plot()"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "savitam"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Automated Machine Learning\n",
+ "_**Remote Execution using attach**_\n",
+ "\n",
+ "## Contents\n",
+ "1. [Introduction](#Introduction)\n",
+ "1. [Setup](#Setup)\n",
+ "1. [Data](#Data)\n",
+ "1. [Train](#Train)\n",
+ "1. [Results](#Results)\n",
+ "1. [Test](#Test)\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Introduction\n",
+ "In this example we use the scikit-learn's [20newsgroup](http://scikit-learn.org/stable/modules/generated/sklearn.datasets.fetch_20newsgroups.html) to showcase how you can use AutoML to handle text data with remote attach.\n",
+ "\n",
+ "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
+ "\n",
+ "In this notebook you will learn how to:\n",
+ "1. Create an `Experiment` in an existing `Workspace`.\n",
+ "2. Attach an existing DSVM to a workspace.\n",
+ "3. Configure AutoML using `AutoMLConfig`.\n",
+ "4. Train the model using the DSVM.\n",
+ "5. Explore the results.\n",
+ "6. Test the best fitted model.\n",
+ "\n",
+ "In addition this notebook showcases the following features\n",
+ "- **Parallel** executions for iterations\n",
+ "- **Asynchronous** tracking of progress\n",
+ "- **Cancellation** of individual iterations or the entire run\n",
+ "- Retrieving models for any iteration or logged metric\n",
+ "- Specifying AutoML settings as `**kwargs`\n",
+ "- Handling **text** data using the `preprocess` flag"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Setup\n",
+ "\n",
+ "As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import os\n",
+ "\n",
+ "import numpy as np\n",
+ "import pandas as pd\n",
+ "\n",
+ "import azureml.core\n",
+ "from azureml.core.experiment import Experiment\n",
+ "from azureml.core.workspace import Workspace\n",
+ "from azureml.train.automl import AutoMLConfig"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "ws = Workspace.from_config()\n",
+ "\n",
+ "# Choose a name for the run history container in the workspace.\n",
+ "experiment_name = 'automl-remote-attach'\n",
+ "project_folder = './sample_projects/automl-remote-attach'\n",
+ "\n",
+ "experiment = Experiment(ws, experiment_name)\n",
+ "\n",
+ "output = {}\n",
+ "output['SDK version'] = azureml.core.VERSION\n",
+ "output['Subscription ID'] = ws.subscription_id\n",
+ "output['Workspace'] = ws.name\n",
+ "output['Resource Group'] = ws.resource_group\n",
+ "output['Location'] = ws.location\n",
+ "output['Project Directory'] = project_folder\n",
+ "output['Experiment Name'] = experiment.name\n",
+ "pd.set_option('display.max_colwidth', -1)\n",
+ "outputDf = pd.DataFrame(data = output, index = [''])\n",
+ "outputDf.T"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Opt-in diagnostics for better experience, quality, and security of future releases."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.telemetry import set_diagnostics_collection\n",
+ "set_diagnostics_collection(send_diagnostics = True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Attach a Remote Linux DSVM\n",
+ "To use a remote Docker compute target:\n",
+ "1. Create a Linux DSVM in Azure, following these [quick instructions](https://docs.microsoft.com/en-us/azure/machine-learning/desktop-workbench/how-to-create-dsvm-hdi). Make sure you use the Ubuntu flavor (not CentOS). Make sure that disk space is available under `/tmp` because AutoML creates files under `/tmp/azureml_run`s. The DSVM should have more cores than the number of parallel runs that you plan to enable. It should also have at least 4GB per core.\n",
+ "2. Enter the IP address, user name and password below.\n",
+ "\n",
+ "**Note:** By default, SSH runs on port 22 and you don't need to change the port number below. If you've configured SSH to use a different port, change `dsvm_ssh_port` accordinglyaddress. [Read more](https://render.githubusercontent.com/documentation/sdk/ssh-issue.md) on changing SSH ports for security reasons."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.compute import ComputeTarget, RemoteCompute\n",
+ "import time\n",
+ "\n",
+ "# Add your VM information below\n",
+ "# If a compute with the specified compute_name already exists, it will be used and the dsvm_ip_addr, dsvm_ssh_port, \n",
+ "# dsvm_username and dsvm_password will be ignored.\n",
+ "compute_name = 'mydsvmb'\n",
+ "dsvm_ip_addr = '<>'\n",
+ "dsvm_ssh_port = 22\n",
+ "dsvm_username = '<>'\n",
+ "dsvm_password = '<>'\n",
+ "\n",
+ "if compute_name in ws.compute_targets:\n",
+ " print('Using existing compute.')\n",
+ " dsvm_compute = ws.compute_targets[compute_name]\n",
+ "else:\n",
+ " attach_config = RemoteCompute.attach_configuration(address=dsvm_ip_addr, username=dsvm_username, password=dsvm_password, ssh_port=dsvm_ssh_port)\n",
+ " ComputeTarget.attach(workspace=ws, name=compute_name, attach_configuration=attach_config)\n",
+ "\n",
+ " while ws.compute_targets[compute_name].provisioning_state == 'Creating':\n",
+ " time.sleep(1)\n",
+ "\n",
+ " dsvm_compute = ws.compute_targets[compute_name]\n",
+ " \n",
+ " if dsvm_compute.provisioning_state == 'Failed':\n",
+ " print('Attached failed.')\n",
+ " print(dsvm_compute.provisioning_errors)\n",
+ " dsvm_compute.detach()"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.runconfig import RunConfiguration\n",
+ "from azureml.core.conda_dependencies import CondaDependencies\n",
+ "\n",
+ "# create a new RunConfig object\n",
+ "conda_run_config = RunConfiguration(framework=\"python\")\n",
+ "\n",
+ "# Set compute target to the Linux DSVM\n",
+ "conda_run_config.target = dsvm_compute\n",
+ "\n",
+ "cd = CondaDependencies.create(pip_packages=['azureml-sdk[automl]'], conda_packages=['numpy'])\n",
+ "conda_run_config.environment.python.conda_dependencies = cd"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Data\n",
+ "For remote executions you should author a `get_data.py` file containing a `get_data()` function. This file should be in the root directory of the project. You can encapsulate code to read data either from a blob storage or local disk in this file.\n",
+ "In this example, the `get_data()` function returns a [dictionary](README.md#getdata)."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "if not os.path.exists(project_folder):\n",
+ " os.makedirs(project_folder)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%writefile $project_folder/get_data.py\n",
+ "\n",
+ "import numpy as np\n",
+ "from sklearn.datasets import fetch_20newsgroups\n",
+ "\n",
+ "def get_data():\n",
+ " remove = ('headers', 'footers', 'quotes')\n",
+ " categories = [\n",
+ " 'alt.atheism',\n",
+ " 'talk.religion.misc',\n",
+ " 'comp.graphics',\n",
+ " 'sci.space',\n",
+ " ]\n",
+ " data_train = fetch_20newsgroups(subset = 'train', categories = categories,\n",
+ " shuffle = True, random_state = 42,\n",
+ " remove = remove)\n",
+ " \n",
+ " X_train = np.array(data_train.data).reshape((len(data_train.data),1))\n",
+ " y_train = np.array(data_train.target)\n",
+ " \n",
+ " return { \"X\" : X_train, \"y\" : y_train }"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Train\n",
+ "\n",
+ "You can specify `automl_settings` as `**kwargs` as well. Also note that you can use a `get_data()` function for local excutions too.\n",
+ "\n",
+ "**Note:** When using Remote DSVM, you can't pass Numpy arrays directly to the fit method.\n",
+ "\n",
+ "|Property|Description|\n",
+ "|-|-|\n",
+ "|**primary_metric**|This is the metric that you want to optimize. Classification supports the following primary metrics:
accuracy
AUC_weighted
average_precision_score_weighted
norm_macro_recall
precision_score_weighted|\n",
+ "|**iteration_timeout_minutes**|Time limit in minutes for each iteration.|\n",
+ "|**iterations**|Number of iterations. In each iteration AutoML trains a specific pipeline with the data.|\n",
+ "|**n_cross_validations**|Number of cross validation splits.|\n",
+ "|**max_concurrent_iterations**|Maximum number of iterations that would be executed in parallel. This should be less than the number of cores on the DSVM.|\n",
+ "|**preprocess**|Setting this to *True* enables AutoML to perform preprocessing on the input to handle *missing data*, and to perform some common *feature extraction*.|\n",
+ "|**enable_cache**|Setting this to *True* enables preprocess done once and reuse the same preprocessed data for all the iterations. Default value is True.\n",
+ "|**max_cores_per_iteration**|Indicates how many cores on the compute target would be used to train a single pipeline.
Default is *1*; you can set it to *-1* to use all cores.|"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "automl_settings = {\n",
+ " \"iteration_timeout_minutes\": 60,\n",
+ " \"iterations\": 4,\n",
+ " \"n_cross_validations\": 5,\n",
+ " \"primary_metric\": 'AUC_weighted',\n",
+ " \"preprocess\": True,\n",
+ " \"max_cores_per_iteration\": 2\n",
+ "}\n",
+ "\n",
+ "automl_config = AutoMLConfig(task = 'classification',\n",
+ " path = project_folder,\n",
+ " run_configuration=conda_run_config,\n",
+ " data_script = project_folder + \"/get_data.py\",\n",
+ " **automl_settings\n",
+ " )\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Call the `submit` method on the experiment object and pass the run configuration. For remote runs the execution is asynchronous, so you will see the iterations get populated as they complete. You can interact with the widgets and models even when the experiment is running to retrieve the best model up to that point. Once you are satisfied with the model, you can cancel a particular iteration or the whole run."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "remote_run = experiment.submit(automl_config)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "remote_run"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Results\n",
+ "#### Widget for Monitoring Runs\n",
+ "\n",
+ "The widget will first report a \"loading\" status while running the first iteration. After completing the first iteration, an auto-updating graph and table will be shown. The widget will refresh once per minute, so you should see the graph update as child runs complete.\n",
+ "\n",
+ "You can click on a pipeline to see run properties and output logs. Logs are also available on the DSVM under `/tmp/azureml_run/{iterationid}/azureml-logs`\n",
+ "\n",
+ "**Note:** The widget displays a link at the bottom. Use this link to open a web interface to explore the individual run details."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.widgets import RunDetails\n",
+ "RunDetails(remote_run).show() "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Wait until the run finishes.\n",
+ "remote_run.wait_for_completion(show_output = True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Pre-process cache cleanup\n",
+ "The preprocess data gets cache at user default file store. When the run is completed the cache can be cleaned by running below cell"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "remote_run.clean_preprocessor_cache()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "\n",
+ "#### Retrieve All Child Runs\n",
+ "You can also use SDK methods to fetch all the child runs and see individual metrics that we log. "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "children = list(remote_run.get_children())\n",
+ "metricslist = {}\n",
+ "for run in children:\n",
+ " properties = run.get_properties()\n",
+ " metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)}\n",
+ " metricslist[int(properties['iteration'])] = metrics\n",
+ "\n",
+ "rundata = pd.DataFrame(metricslist).sort_index(1)\n",
+ "rundata"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Cancelling Runs\n",
+ "You can cancel ongoing remote runs using the `cancel` and `cancel_iteration` functions."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Cancel the ongoing experiment and stop scheduling new iterations.\n",
+ "# remote_run.cancel()\n",
+ "\n",
+ "# Cancel iteration 1 and move onto iteration 2.\n",
+ "# remote_run.cancel_iteration(1)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Retrieve the Best Model\n",
+ "\n",
+ "Below we select the best pipeline from our iterations. The `get_output` method returns the best run and the fitted model. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "best_run, fitted_model = remote_run.get_output()\n",
+ "print(best_run)\n",
+ "print(fitted_model)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Best Model Based on Any Other Metric\n",
+ "Show the run and the model which has the smallest `accuracy` value:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# lookup_metric = \"accuracy\"\n",
+ "# best_run, fitted_model = remote_run.get_output(metric = lookup_metric)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Model from a Specific Iteration"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "iteration = 0\n",
+ "zero_run, zero_model = remote_run.get_output(iteration = iteration)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Test"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Load test data.\n",
+ "from pandas_ml import ConfusionMatrix\n",
+ "from sklearn.datasets import fetch_20newsgroups\n",
+ "\n",
+ "remove = ('headers', 'footers', 'quotes')\n",
+ "categories = [\n",
+ " 'alt.atheism',\n",
+ " 'talk.religion.misc',\n",
+ " 'comp.graphics',\n",
+ " 'sci.space',\n",
+ " ]\n",
+ "\n",
+ "data_test = fetch_20newsgroups(subset = 'test', categories = categories,\n",
+ " shuffle = True, random_state = 42,\n",
+ " remove = remove)\n",
+ "\n",
+ "X_test = np.array(data_test.data).reshape((len(data_test.data),1))\n",
+ "y_test = data_test.target\n",
+ "\n",
+ "# Test our best pipeline.\n",
+ "\n",
+ "y_pred = fitted_model.predict(X_test)\n",
+ "y_pred_strings = [data_test.target_names[i] for i in y_pred]\n",
+ "y_test_strings = [data_test.target_names[i] for i in y_test]\n",
+ "\n",
+ "cm = ConfusionMatrix(y_test_strings, y_pred_strings)\n",
+ "print(cm)\n",
+ "cm.plot()"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "savitam"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.6"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/automated-machine-learning/remote-batchai/auto-ml-remote-batchai.ipynb b/how-to-use-azureml/automated-machine-learning/remote-batchai/auto-ml-remote-batchai.ipynb
index c2f22728..551a624e 100644
--- a/how-to-use-azureml/automated-machine-learning/remote-batchai/auto-ml-remote-batchai.ipynb
+++ b/how-to-use-azureml/automated-machine-learning/remote-batchai/auto-ml-remote-batchai.ipynb
@@ -1,548 +1,546 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Automated Machine Learning\n",
- "_**Remote Execution using AmlCompute**_\n",
- "\n",
- "## Contents\n",
- "1. [Introduction](#Introduction)\n",
- "1. [Setup](#Setup)\n",
- "1. [Data](#Data)\n",
- "1. [Train](#Train)\n",
- "1. [Results](#Results)\n",
- "1. [Test](#Test)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Introduction\n",
- "In this example we use the scikit-learn's [digit dataset](http://scikit-learn.org/stable/datasets/index.html#optical-recognition-of-handwritten-digits-dataset) to showcase how you can use AutoML for a simple classification problem.\n",
- "\n",
- "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
- "\n",
- "In this notebook you would see\n",
- "1. Create an `Experiment` in an existing `Workspace`.\n",
- "2. Create or Attach existing AmlCompute to a workspace.\n",
- "3. Configure AutoML using `AutoMLConfig`.\n",
- "4. Train the model using AmlCompute\n",
- "5. Explore the results.\n",
- "6. Test the best fitted model.\n",
- "\n",
- "In addition this notebook showcases the following features\n",
- "- **Parallel** executions for iterations\n",
- "- **Asynchronous** tracking of progress\n",
- "- **Cancellation** of individual iterations or the entire run\n",
- "- Retrieving models for any iteration or logged metric\n",
- "- Specifying AutoML settings as `**kwargs`"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Setup\n",
- "\n",
- "As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import logging\n",
- "import os\n",
- "import random\n",
- "\n",
- "from matplotlib import pyplot as plt\n",
- "from matplotlib.pyplot import imshow\n",
- "import numpy as np\n",
- "import pandas as pd\n",
- "from sklearn import datasets\n",
- "\n",
- "import azureml.core\n",
- "from azureml.core.experiment import Experiment\n",
- "from azureml.core.workspace import Workspace\n",
- "from azureml.train.automl import AutoMLConfig\n",
- "from azureml.train.automl.run import AutoMLRun"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "ws = Workspace.from_config()\n",
- "\n",
- "# Choose a name for the run history container in the workspace.\n",
- "experiment_name = 'automl-remote-amlcompute'\n",
- "project_folder = './sample_projects/automl-remote-amlcompute'\n",
- "\n",
- "experiment = Experiment(ws, experiment_name)\n",
- "\n",
- "output = {}\n",
- "output['SDK version'] = azureml.core.VERSION\n",
- "output['Subscription ID'] = ws.subscription_id\n",
- "output['Workspace Name'] = ws.name\n",
- "output['Resource Group'] = ws.resource_group\n",
- "output['Location'] = ws.location\n",
- "output['Project Directory'] = project_folder\n",
- "output['Experiment Name'] = experiment.name\n",
- "pd.set_option('display.max_colwidth', -1)\n",
- "pd.DataFrame(data = output, index = ['']).T"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Opt-in diagnostics for better experience, quality, and security of future releases."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.telemetry import set_diagnostics_collection\n",
- "set_diagnostics_collection(send_diagnostics = True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create or Attach existing AmlCompute\n",
- "You will need to create a [compute target](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#compute-target) for your AutoML run. In this tutorial, you create `AmlCompute` as your training compute resource.\n",
- "\n",
- "**Creation of AmlCompute takes approximately 5 minutes.** If the AmlCompute with that name is already in your workspace this code will skip the creation process.\n",
- "\n",
- "As with other Azure services, there are limits on certain resources (e.g. AmlCompute) associated with the Azure Machine Learning service. Please read [this article](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-manage-quotas) on the default limits and how to request more quota."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.compute import AmlCompute\n",
- "from azureml.core.compute import ComputeTarget\n",
- "\n",
- "# Choose a name for your cluster.\n",
- "amlcompute_cluster_name = \"automlcl\"\n",
- "\n",
- "found = False\n",
- "# Check if this compute target already exists in the workspace.\n",
- "cts = ws.compute_targets\n",
- "if amlcompute_cluster_name in cts and cts[amlcompute_cluster_name].type == 'AmlCompute':\n",
- " found = True\n",
- " print('Found existing compute target.')\n",
- " compute_target = cts[amlcompute_cluster_name]\n",
- " \n",
- "if not found:\n",
- " print('Creating a new compute target...')\n",
- " provisioning_config = AmlCompute.provisioning_configuration(vm_size = \"STANDARD_D2_V2\", # for GPU, use \"STANDARD_NC6\"\n",
- " #vm_priority = 'lowpriority', # optional\n",
- " max_nodes = 6)\n",
- "\n",
- " # Create the cluster.\n",
- " compute_target = ComputeTarget.create(ws, amlcompute_cluster_name, provisioning_config)\n",
- " \n",
- " # Can poll for a minimum number of nodes and for a specific timeout.\n",
- " # If no min_node_count is provided, it will use the scale settings for the cluster.\n",
- " compute_target.wait_for_completion(show_output = True, min_node_count = None, timeout_in_minutes = 20)\n",
- " \n",
- " # For a more detailed view of current AmlCompute status, use the 'status' property."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.runconfig import RunConfiguration\n",
- "from azureml.core.conda_dependencies import CondaDependencies\n",
- "\n",
- "# create a new RunConfig object\n",
- "conda_run_config = RunConfiguration(framework=\"python\")\n",
- "\n",
- "# Set compute target to AmlCompute\n",
- "conda_run_config.target = compute_target\n",
- "conda_run_config.environment.docker.enabled = True\n",
- "conda_run_config.environment.docker.base_image = azureml.core.runconfig.DEFAULT_CPU_IMAGE\n",
- "\n",
- "cd = CondaDependencies.create(pip_packages=['azureml-sdk[automl]'], conda_packages=['numpy'])\n",
- "conda_run_config.environment.python.conda_dependencies = cd"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Data\n",
- "For remote executions you should author a `get_data.py` file containing a `get_data()` function. This file should be in the root directory of the project. You can encapsulate code to read data either from a blob storage or local disk in this file.\n",
- "In this example, the `get_data()` function returns data using scikit-learn's [load_digits](http://scikit-learn.org/stable/modules/generated/sklearn.datasets.load_digits.html) method."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "if not os.path.exists(project_folder):\n",
- " os.makedirs(project_folder)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%writefile $project_folder/get_data.py\n",
- "\n",
- "from sklearn import datasets\n",
- "from scipy import sparse\n",
- "import numpy as np\n",
- "\n",
- "def get_data():\n",
- " \n",
- " digits = datasets.load_digits()\n",
- " X_train = digits.data\n",
- " y_train = digits.target\n",
- "\n",
- " return { \"X\" : X_train, \"y\" : y_train }"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Train\n",
- "\n",
- "You can specify `automl_settings` as `**kwargs` as well. Also note that you can use a `get_data()` function for local excutions too.\n",
- "\n",
- "**Note:** When using AmlCompute, you can't pass Numpy arrays directly to the fit method.\n",
- "\n",
- "|Property|Description|\n",
- "|-|-|\n",
- "|**primary_metric**|This is the metric that you want to optimize. Classification supports the following primary metrics:
accuracy
AUC_weighted
average_precision_score_weighted
norm_macro_recall
precision_score_weighted|\n",
- "|**iteration_timeout_minutes**|Time limit in minutes for each iteration.|\n",
- "|**iterations**|Number of iterations. In each iteration AutoML trains a specific pipeline with the data.|\n",
- "|**n_cross_validations**|Number of cross validation splits.|\n",
- "|**max_concurrent_iterations**|Maximum number of iterations that would be executed in parallel. This should be less than the number of cores on the DSVM.|"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "automl_settings = {\n",
- " \"iteration_timeout_minutes\": 2,\n",
- " \"iterations\": 20,\n",
- " \"n_cross_validations\": 5,\n",
- " \"primary_metric\": 'AUC_weighted',\n",
- " \"preprocess\": False,\n",
- " \"max_concurrent_iterations\": 5,\n",
- " \"verbosity\": logging.INFO\n",
- "}\n",
- "\n",
- "automl_config = AutoMLConfig(task = 'classification',\n",
- " debug_log = 'automl_errors.log',\n",
- " path = project_folder,\n",
- " run_configuration=conda_run_config,\n",
- " data_script = project_folder + \"/get_data.py\",\n",
- " **automl_settings\n",
- " )\n"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Call the `submit` method on the experiment object and pass the run configuration. For remote runs the execution is asynchronous, so you will see the iterations get populated as they complete. You can interact with the widgets and models even when the experiment is running to retrieve the best model up to that point. Once you are satisfied with the model, you can cancel a particular iteration or the whole run.\n",
- "In this example, we specify `show_output = False` to suppress console output while the run is in progress."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "remote_run = experiment.submit(automl_config, show_output = False)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "remote_run"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Results\n",
- "\n",
- "#### Loading executed runs\n",
- "In case you need to load a previously executed run, enable the cell below and replace the `run_id` value."
- ]
- },
- {
- "cell_type": "raw",
- "metadata": {},
- "source": [
- "remote_run = AutoMLRun(experiment = experiment, run_id = 'AutoML_5db13491-c92a-4f1d-b622-8ab8d973a058')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Widget for Monitoring Runs\n",
- "\n",
- "The widget will first report a \"loading\" status while running the first iteration. After completing the first iteration, an auto-updating graph and table will be shown. The widget will refresh once per minute, so you should see the graph update as child runs complete.\n",
- "\n",
- "You can click on a pipeline to see run properties and output logs. Logs are also available on the DSVM under `/tmp/azureml_run/{iterationid}/azureml-logs`\n",
- "\n",
- "**Note:** The widget displays a link at the bottom. Use this link to open a web interface to explore the individual run details."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "remote_run"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.widgets import RunDetails\n",
- "RunDetails(remote_run).show() "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Wait until the run finishes.\n",
- "remote_run.wait_for_completion(show_output = True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "\n",
- "#### Retrieve All Child Runs\n",
- "You can also use SDK methods to fetch all the child runs and see individual metrics that we log."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "children = list(remote_run.get_children())\n",
- "metricslist = {}\n",
- "for run in children:\n",
- " properties = run.get_properties()\n",
- " metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)}\n",
- " metricslist[int(properties['iteration'])] = metrics\n",
- "\n",
- "rundata = pd.DataFrame(metricslist).sort_index(1)\n",
- "rundata"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Cancelling Runs\n",
- "\n",
- "You can cancel ongoing remote runs using the `cancel` and `cancel_iteration` functions."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Cancel the ongoing experiment and stop scheduling new iterations.\n",
- "# remote_run.cancel()\n",
- "\n",
- "# Cancel iteration 1 and move onto iteration 2.\n",
- "# remote_run.cancel_iteration(1)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Retrieve the Best Model\n",
- "\n",
- "Below we select the best pipeline from our iterations. The `get_output` method returns the best run and the fitted model. The Model includes the pipeline and any pre-processing. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "best_run, fitted_model = remote_run.get_output()\n",
- "print(best_run)\n",
- "print(fitted_model)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Best Model Based on Any Other Metric\n",
- "Show the run and the model which has the smallest `log_loss` value:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "lookup_metric = \"log_loss\"\n",
- "best_run, fitted_model = remote_run.get_output(metric = lookup_metric)\n",
- "print(best_run)\n",
- "print(fitted_model)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Model from a Specific Iteration\n",
- "Show the run and the model from the third iteration:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "iteration = 3\n",
- "third_run, third_model = remote_run.get_output(iteration=iteration)\n",
- "print(third_run)\n",
- "print(third_model)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Test\n",
- "\n",
- "#### Load Test Data"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "digits = datasets.load_digits()\n",
- "X_test = digits.data[:10, :]\n",
- "y_test = digits.target[:10]\n",
- "images = digits.images[:10]"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Testing Our Best Fitted Model"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Randomly select digits and test.\n",
- "for index in np.random.choice(len(y_test), 2, replace = False):\n",
- " print(index)\n",
- " predicted = fitted_model.predict(X_test[index:index + 1])[0]\n",
- " label = y_test[index]\n",
- " title = \"Label value = %d Predicted value = %d \" % (label, predicted)\n",
- " fig = plt.figure(1, figsize=(3,3))\n",
- " ax1 = fig.add_axes((0,0,.8,.8))\n",
- " ax1.set_title(title)\n",
- " plt.imshow(images[index], cmap = plt.cm.gray_r, interpolation = 'nearest')\n",
- " plt.show()"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "savitam"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Automated Machine Learning\n",
+ "_**Remote Execution using AmlCompute**_\n",
+ "\n",
+ "## Contents\n",
+ "1. [Introduction](#Introduction)\n",
+ "1. [Setup](#Setup)\n",
+ "1. [Data](#Data)\n",
+ "1. [Train](#Train)\n",
+ "1. [Results](#Results)\n",
+ "1. [Test](#Test)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Introduction\n",
+ "In this example we use the scikit-learn's [digit dataset](http://scikit-learn.org/stable/datasets/index.html#optical-recognition-of-handwritten-digits-dataset) to showcase how you can use AutoML for a simple classification problem.\n",
+ "\n",
+ "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
+ "\n",
+ "In this notebook you would see\n",
+ "1. Create an `Experiment` in an existing `Workspace`.\n",
+ "2. Create or Attach existing AmlCompute to a workspace.\n",
+ "3. Configure AutoML using `AutoMLConfig`.\n",
+ "4. Train the model using AmlCompute\n",
+ "5. Explore the results.\n",
+ "6. Test the best fitted model.\n",
+ "\n",
+ "In addition this notebook showcases the following features\n",
+ "- **Parallel** executions for iterations\n",
+ "- **Asynchronous** tracking of progress\n",
+ "- **Cancellation** of individual iterations or the entire run\n",
+ "- Retrieving models for any iteration or logged metric\n",
+ "- Specifying AutoML settings as `**kwargs`"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Setup\n",
+ "\n",
+ "As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import logging\n",
+ "import os\n",
+ "\n",
+ "from matplotlib import pyplot as plt\n",
+ "import numpy as np\n",
+ "import pandas as pd\n",
+ "from sklearn import datasets\n",
+ "\n",
+ "import azureml.core\n",
+ "from azureml.core.experiment import Experiment\n",
+ "from azureml.core.workspace import Workspace\n",
+ "from azureml.train.automl import AutoMLConfig"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "ws = Workspace.from_config()\n",
+ "\n",
+ "# Choose a name for the run history container in the workspace.\n",
+ "experiment_name = 'automl-remote-amlcompute'\n",
+ "project_folder = './sample_projects/automl-remote-amlcompute'\n",
+ "\n",
+ "experiment = Experiment(ws, experiment_name)\n",
+ "\n",
+ "output = {}\n",
+ "output['SDK version'] = azureml.core.VERSION\n",
+ "output['Subscription ID'] = ws.subscription_id\n",
+ "output['Workspace Name'] = ws.name\n",
+ "output['Resource Group'] = ws.resource_group\n",
+ "output['Location'] = ws.location\n",
+ "output['Project Directory'] = project_folder\n",
+ "output['Experiment Name'] = experiment.name\n",
+ "pd.set_option('display.max_colwidth', -1)\n",
+ "outputDf = pd.DataFrame(data = output, index = [''])\n",
+ "outputDf.T"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Opt-in diagnostics for better experience, quality, and security of future releases."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.telemetry import set_diagnostics_collection\n",
+ "set_diagnostics_collection(send_diagnostics = True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create or Attach existing AmlCompute\n",
+ "You will need to create a [compute target](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#compute-target) for your AutoML run. In this tutorial, you create `AmlCompute` as your training compute resource.\n",
+ "\n",
+ "**Creation of AmlCompute takes approximately 5 minutes.** If the AmlCompute with that name is already in your workspace this code will skip the creation process.\n",
+ "\n",
+ "As with other Azure services, there are limits on certain resources (e.g. AmlCompute) associated with the Azure Machine Learning service. Please read [this article](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-manage-quotas) on the default limits and how to request more quota."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.compute import AmlCompute\n",
+ "from azureml.core.compute import ComputeTarget\n",
+ "\n",
+ "# Choose a name for your cluster.\n",
+ "amlcompute_cluster_name = \"automlcl\"\n",
+ "\n",
+ "found = False\n",
+ "# Check if this compute target already exists in the workspace.\n",
+ "cts = ws.compute_targets\n",
+ "if amlcompute_cluster_name in cts and cts[amlcompute_cluster_name].type == 'AmlCompute':\n",
+ " found = True\n",
+ " print('Found existing compute target.')\n",
+ " compute_target = cts[amlcompute_cluster_name]\n",
+ " \n",
+ "if not found:\n",
+ " print('Creating a new compute target...')\n",
+ " provisioning_config = AmlCompute.provisioning_configuration(vm_size = \"STANDARD_D2_V2\", # for GPU, use \"STANDARD_NC6\"\n",
+ " #vm_priority = 'lowpriority', # optional\n",
+ " max_nodes = 6)\n",
+ "\n",
+ " # Create the cluster.\n",
+ " compute_target = ComputeTarget.create(ws, amlcompute_cluster_name, provisioning_config)\n",
+ " \n",
+ " # Can poll for a minimum number of nodes and for a specific timeout.\n",
+ " # If no min_node_count is provided, it will use the scale settings for the cluster.\n",
+ " compute_target.wait_for_completion(show_output = True, min_node_count = None, timeout_in_minutes = 20)\n",
+ " \n",
+ " # For a more detailed view of current AmlCompute status, use get_status()."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.runconfig import RunConfiguration\n",
+ "from azureml.core.conda_dependencies import CondaDependencies\n",
+ "\n",
+ "# create a new RunConfig object\n",
+ "conda_run_config = RunConfiguration(framework=\"python\")\n",
+ "\n",
+ "# Set compute target to AmlCompute\n",
+ "conda_run_config.target = compute_target\n",
+ "conda_run_config.environment.docker.enabled = True\n",
+ "conda_run_config.environment.docker.base_image = azureml.core.runconfig.DEFAULT_CPU_IMAGE\n",
+ "\n",
+ "cd = CondaDependencies.create(pip_packages=['azureml-sdk[automl]'], conda_packages=['numpy'])\n",
+ "conda_run_config.environment.python.conda_dependencies = cd"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Data\n",
+ "For remote executions you should author a `get_data.py` file containing a `get_data()` function. This file should be in the root directory of the project. You can encapsulate code to read data either from a blob storage or local disk in this file.\n",
+ "In this example, the `get_data()` function returns data using scikit-learn's [load_digits](http://scikit-learn.org/stable/modules/generated/sklearn.datasets.load_digits.html) method."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "if not os.path.exists(project_folder):\n",
+ " os.makedirs(project_folder)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%writefile $project_folder/get_data.py\n",
+ "\n",
+ "from sklearn import datasets\n",
+ "from scipy import sparse\n",
+ "import numpy as np\n",
+ "\n",
+ "def get_data():\n",
+ " \n",
+ " digits = datasets.load_digits()\n",
+ " X_train = digits.data\n",
+ " y_train = digits.target\n",
+ "\n",
+ " return { \"X\" : X_train, \"y\" : y_train }"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Train\n",
+ "\n",
+ "You can specify `automl_settings` as `**kwargs` as well. Also note that you can use a `get_data()` function for local excutions too.\n",
+ "\n",
+ "**Note:** When using AmlCompute, you can't pass Numpy arrays directly to the fit method.\n",
+ "\n",
+ "|Property|Description|\n",
+ "|-|-|\n",
+ "|**primary_metric**|This is the metric that you want to optimize. Classification supports the following primary metrics:
accuracy
AUC_weighted
average_precision_score_weighted
norm_macro_recall
precision_score_weighted|\n",
+ "|**iteration_timeout_minutes**|Time limit in minutes for each iteration.|\n",
+ "|**iterations**|Number of iterations. In each iteration AutoML trains a specific pipeline with the data.|\n",
+ "|**n_cross_validations**|Number of cross validation splits.|\n",
+ "|**max_concurrent_iterations**|Maximum number of iterations that would be executed in parallel. This should be less than the number of cores on the DSVM.|"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "automl_settings = {\n",
+ " \"iteration_timeout_minutes\": 2,\n",
+ " \"iterations\": 20,\n",
+ " \"n_cross_validations\": 5,\n",
+ " \"primary_metric\": 'AUC_weighted',\n",
+ " \"preprocess\": False,\n",
+ " \"max_concurrent_iterations\": 5,\n",
+ " \"verbosity\": logging.INFO\n",
+ "}\n",
+ "\n",
+ "automl_config = AutoMLConfig(task = 'classification',\n",
+ " debug_log = 'automl_errors.log',\n",
+ " path = project_folder,\n",
+ " run_configuration=conda_run_config,\n",
+ " data_script = project_folder + \"/get_data.py\",\n",
+ " **automl_settings\n",
+ " )\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Call the `submit` method on the experiment object and pass the run configuration. For remote runs the execution is asynchronous, so you will see the iterations get populated as they complete. You can interact with the widgets and models even when the experiment is running to retrieve the best model up to that point. Once you are satisfied with the model, you can cancel a particular iteration or the whole run.\n",
+ "In this example, we specify `show_output = False` to suppress console output while the run is in progress."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "remote_run = experiment.submit(automl_config, show_output = False)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "remote_run"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Results\n",
+ "\n",
+ "#### Loading executed runs\n",
+ "In case you need to load a previously executed run, enable the cell below and replace the `run_id` value."
+ ]
+ },
+ {
+ "cell_type": "raw",
+ "metadata": {},
+ "source": [
+ "remote_run = AutoMLRun(experiment = experiment, run_id = 'AutoML_5db13491-c92a-4f1d-b622-8ab8d973a058')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Widget for Monitoring Runs\n",
+ "\n",
+ "The widget will first report a \"loading\" status while running the first iteration. After completing the first iteration, an auto-updating graph and table will be shown. The widget will refresh once per minute, so you should see the graph update as child runs complete.\n",
+ "\n",
+ "You can click on a pipeline to see run properties and output logs. Logs are also available on the DSVM under `/tmp/azureml_run/{iterationid}/azureml-logs`\n",
+ "\n",
+ "**Note:** The widget displays a link at the bottom. Use this link to open a web interface to explore the individual run details."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "remote_run"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.widgets import RunDetails\n",
+ "RunDetails(remote_run).show() "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Wait until the run finishes.\n",
+ "remote_run.wait_for_completion(show_output = True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "\n",
+ "#### Retrieve All Child Runs\n",
+ "You can also use SDK methods to fetch all the child runs and see individual metrics that we log."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "children = list(remote_run.get_children())\n",
+ "metricslist = {}\n",
+ "for run in children:\n",
+ " properties = run.get_properties()\n",
+ " metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)}\n",
+ " metricslist[int(properties['iteration'])] = metrics\n",
+ "\n",
+ "rundata = pd.DataFrame(metricslist).sort_index(1)\n",
+ "rundata"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Cancelling Runs\n",
+ "\n",
+ "You can cancel ongoing remote runs using the `cancel` and `cancel_iteration` functions."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Cancel the ongoing experiment and stop scheduling new iterations.\n",
+ "# remote_run.cancel()\n",
+ "\n",
+ "# Cancel iteration 1 and move onto iteration 2.\n",
+ "# remote_run.cancel_iteration(1)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Retrieve the Best Model\n",
+ "\n",
+ "Below we select the best pipeline from our iterations. The `get_output` method returns the best run and the fitted model. The Model includes the pipeline and any pre-processing. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "best_run, fitted_model = remote_run.get_output()\n",
+ "print(best_run)\n",
+ "print(fitted_model)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Best Model Based on Any Other Metric\n",
+ "Show the run and the model which has the smallest `log_loss` value:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "lookup_metric = \"log_loss\"\n",
+ "best_run, fitted_model = remote_run.get_output(metric = lookup_metric)\n",
+ "print(best_run)\n",
+ "print(fitted_model)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Model from a Specific Iteration\n",
+ "Show the run and the model from the third iteration:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "iteration = 3\n",
+ "third_run, third_model = remote_run.get_output(iteration=iteration)\n",
+ "print(third_run)\n",
+ "print(third_model)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Test\n",
+ "\n",
+ "#### Load Test Data"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "digits = datasets.load_digits()\n",
+ "X_test = digits.data[:10, :]\n",
+ "y_test = digits.target[:10]\n",
+ "images = digits.images[:10]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Testing Our Best Fitted Model"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Randomly select digits and test.\n",
+ "for index in np.random.choice(len(y_test), 2, replace = False):\n",
+ " print(index)\n",
+ " predicted = fitted_model.predict(X_test[index:index + 1])[0]\n",
+ " label = y_test[index]\n",
+ " title = \"Label value = %d Predicted value = %d \" % (label, predicted)\n",
+ " fig = plt.figure(1, figsize=(3,3))\n",
+ " ax1 = fig.add_axes((0,0,.8,.8))\n",
+ " ax1.set_title(title)\n",
+ " plt.imshow(images[index], cmap = plt.cm.gray_r, interpolation = 'nearest')\n",
+ " plt.show()"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "savitam"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.6"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/automated-machine-learning/remote-execution-with-datastore/auto-ml-remote-execution-with-datastore.ipynb b/how-to-use-azureml/automated-machine-learning/remote-execution-with-datastore/auto-ml-remote-execution-with-datastore.ipynb
index 52ca5cdb..fe4d9c3e 100644
--- a/how-to-use-azureml/automated-machine-learning/remote-execution-with-datastore/auto-ml-remote-execution-with-datastore.ipynb
+++ b/how-to-use-azureml/automated-machine-learning/remote-execution-with-datastore/auto-ml-remote-execution-with-datastore.ipynb
@@ -1,604 +1,600 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Automated Machine Learning\n",
- "_**Remote Execution with DataStore**_\n",
- "\n",
- "## Contents\n",
- "1. [Introduction](#Introduction)\n",
- "1. [Setup](#Setup)\n",
- "1. [Data](#Data)\n",
- "1. [Train](#Train)\n",
- "1. [Results](#Results)\n",
- "1. [Test](#Test)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Introduction\n",
- "This sample accesses a data file on a remote DSVM through DataStore. Advantages of using data store are:\n",
- "1. DataStore secures the access details.\n",
- "2. DataStore supports read, write to blob and file store\n",
- "3. AutoML natively supports copying data from DataStore to DSVM\n",
- "\n",
- "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
- "\n",
- "In this notebook you would see\n",
- "1. Storing data in DataStore.\n",
- "2. get_data returning data from DataStore."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Setup\n",
- "\n",
- "As part of the setup you have already created a Workspace. For AutoML you would need to create an Experiment. An Experiment is a named object in a Workspace, which is used to run experiments."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import logging\n",
- "import os\n",
- "import random\n",
- "import time\n",
- "\n",
- "from matplotlib import pyplot as plt\n",
- "from matplotlib.pyplot import imshow\n",
- "import numpy as np\n",
- "import pandas as pd\n",
- "from sklearn import datasets\n",
- "\n",
- "import azureml.core\n",
- "from azureml.core.compute import DsvmCompute\n",
- "from azureml.core.experiment import Experiment\n",
- "from azureml.core.workspace import Workspace\n",
- "from azureml.train.automl import AutoMLConfig\n",
- "from azureml.train.automl.run import AutoMLRun"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "ws = Workspace.from_config()\n",
- "\n",
- "# choose a name for experiment\n",
- "experiment_name = 'automl-remote-datastore-file'\n",
- "# project folder\n",
- "project_folder = './sample_projects/automl-remote-datastore-file'\n",
- "\n",
- "experiment=Experiment(ws, experiment_name)\n",
- "\n",
- "output = {}\n",
- "output['SDK version'] = azureml.core.VERSION\n",
- "output['Subscription ID'] = ws.subscription_id\n",
- "output['Workspace'] = ws.name\n",
- "output['Resource Group'] = ws.resource_group\n",
- "output['Location'] = ws.location\n",
- "output['Project Directory'] = project_folder\n",
- "output['Experiment Name'] = experiment.name\n",
- "pd.set_option('display.max_colwidth', -1)\n",
- "pd.DataFrame(data=output, index=['']).T"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Opt-in diagnostics for better experience, quality, and security of future releases"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.telemetry import set_diagnostics_collection\n",
- "set_diagnostics_collection(send_diagnostics=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create a Remote Linux DSVM\n",
- "Note: If creation fails with a message about Marketplace purchase eligibilty, go to portal.azure.com, start creating DSVM there, and select \"Want to create programmatically\" to enable programmatic creation. Once you've enabled it, you can exit without actually creating VM.\n",
- "\n",
- "**Note**: By default SSH runs on port 22 and you don't need to specify it. But if for security reasons you can switch to a different port (such as 5022), you can append the port number to the address. [Read more](https://render.githubusercontent.com/documentation/sdk/ssh-issue.md) on this."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "compute_target_name = 'mydsvmc'\n",
- "\n",
- "try:\n",
- " while ws.compute_targets[compute_target_name].provisioning_state == 'Creating':\n",
- " time.sleep(1)\n",
- " \n",
- " dsvm_compute = DsvmCompute(workspace=ws, name=compute_target_name)\n",
- " print('found existing:', dsvm_compute.name)\n",
- "except:\n",
- " dsvm_config = DsvmCompute.provisioning_configuration(vm_size=\"Standard_D2_v2\")\n",
- " dsvm_compute = DsvmCompute.create(ws, name=compute_target_name, provisioning_configuration=dsvm_config)\n",
- " dsvm_compute.wait_for_completion(show_output=True)\n",
- " print(\"Waiting one minute for ssh to be accessible\")\n",
- " time.sleep(60) # Wait for ssh to be accessible"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Data\n",
- "\n",
- "### Copy data file to local\n",
- "\n",
- "Download the data file.\n"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "mkdir data"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from sklearn.datasets import fetch_20newsgroups\n",
- "import csv\n",
- "\n",
- "remove = ('headers', 'footers', 'quotes')\n",
- "categories = [\n",
- " 'alt.atheism',\n",
- " 'talk.religion.misc',\n",
- " 'comp.graphics',\n",
- " 'sci.space',\n",
- " ]\n",
- "data_train = fetch_20newsgroups(subset = 'train', categories = categories,\n",
- " shuffle = True, random_state = 42,\n",
- " remove = remove)\n",
- " \n",
- "pd.DataFrame(data_train.data).to_csv(\"data/X_train.tsv\", index=False, header=False, quoting=csv.QUOTE_ALL, sep=\"\\t\")\n",
- "pd.DataFrame(data_train.target).to_csv(\"data/y_train.tsv\", index=False, header=False, sep=\"\\t\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Upload data to the cloud"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Now make the data accessible remotely by uploading that data from your local machine into Azure so it can be accessed for remote training. The datastore is a convenient construct associated with your workspace for you to upload/download data, and interact with it from your remote compute targets. It is backed by Azure blob storage account.\n",
- "\n",
- "The data.tsv files are uploaded into a directory named data at the root of the datastore."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core import Workspace, Datastore\n",
- "#blob_datastore = Datastore(ws, blob_datastore_name)\n",
- "ds = ws.get_default_datastore()\n",
- "print(ds.datastore_type, ds.account_name, ds.container_name)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# ds.upload_files(\"data.tsv\")\n",
- "ds.upload(src_dir='./data', target_path='data', overwrite=True, show_progress=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Configure & Run\n",
- "\n",
- "First let's create a DataReferenceConfigruation object to inform the system what data folder to download to the compute target.\n",
- "The path_on_compute should be an absolute path to ensure that the data files are downloaded only once. The get_data method should use this same path to access the data files."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.runconfig import DataReferenceConfiguration\n",
- "dr = DataReferenceConfiguration(datastore_name=ds.name, \n",
- " path_on_datastore='data', \n",
- " path_on_compute='/tmp/azureml_runs',\n",
- " mode='download', # download files from datastore to compute target\n",
- " overwrite=False)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.runconfig import RunConfiguration\n",
- "from azureml.core.conda_dependencies import CondaDependencies\n",
- "\n",
- "# create a new RunConfig object\n",
- "conda_run_config = RunConfiguration(framework=\"python\")\n",
- "\n",
- "# Set compute target to the Linux DSVM\n",
- "conda_run_config.target = dsvm_compute\n",
- "# set the data reference of the run coonfiguration\n",
- "conda_run_config.data_references = {ds.name: dr}\n",
- "\n",
- "cd = CondaDependencies.create(pip_packages=['azureml-sdk[automl]'], conda_packages=['numpy'])\n",
- "conda_run_config.environment.python.conda_dependencies = cd"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create Get Data File\n",
- "For remote executions you should author a get_data.py file containing a get_data() function. This file should be in the root directory of the project. You can encapsulate code to read data either from a blob storage or local disk in this file.\n",
- "\n",
- "The *get_data()* function returns a [dictionary](README.md#getdata).\n",
- "\n",
- "The read_csv uses the path_on_compute value specified in the DataReferenceConfiguration call plus the path_on_datastore folder and then the actual file name."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "if not os.path.exists(project_folder):\n",
- " os.makedirs(project_folder)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%writefile $project_folder/get_data.py\n",
- "\n",
- "import pandas as pd\n",
- "\n",
- "def get_data():\n",
- " X_train = pd.read_csv(\"/tmp/azureml_runs/data/X_train.tsv\", delimiter=\"\\t\", header=None, quotechar='\"')\n",
- " y_train = pd.read_csv(\"/tmp/azureml_runs/data/y_train.tsv\", delimiter=\"\\t\", header=None, quotechar='\"')\n",
- "\n",
- " return { \"X\" : X_train.values, \"y\" : y_train[0].values }"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Train\n",
- "\n",
- "You can specify automl_settings as **kwargs** as well. Also note that you can use the get_data() symantic for local excutions too. \n",
- "\n",
- "Note: For Remote DSVM and Batch AI you cannot pass Numpy arrays directly to AutoMLConfig.\n",
- "\n",
- "|Property|Description|\n",
- "|-|-|\n",
- "|**primary_metric**|This is the metric that you want to optimize. Classification supports the following primary metrics:
accuracy
AUC_weighted
average_precision_score_weighted
norm_macro_recall
precision_score_weighted|\n",
- "|**iteration_timeout_minutes**|Time limit in minutes for each iteration|\n",
- "|**iterations**|Number of iterations. In each iteration Auto ML trains a specific pipeline with the data|\n",
- "|**n_cross_validations**|Number of cross validation splits|\n",
- "|**max_concurrent_iterations**|Max number of iterations that would be executed in parallel. This should be less than the number of cores on the DSVM\n",
- "|**preprocess**| *True/False*
Setting this to *True* enables Auto ML to perform preprocessing
on the input to handle *missing data*, and perform some common *feature extraction*|\n",
- "|**enable_cache**|Setting this to *True* enables preprocess done once and reuse the same preprocessed data for all the iterations. Default value is True.|\n",
- "|**max_cores_per_iteration**| Indicates how many cores on the compute target would be used to train a single pipeline.
Default is *1*, you can set it to *-1* to use all cores|"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "automl_settings = {\n",
- " \"iteration_timeout_minutes\": 60,\n",
- " \"iterations\": 4,\n",
- " \"n_cross_validations\": 5,\n",
- " \"primary_metric\": 'AUC_weighted',\n",
- " \"preprocess\": True,\n",
- " \"max_cores_per_iteration\": 1,\n",
- " \"verbosity\": logging.INFO\n",
- "}\n",
- "automl_config = AutoMLConfig(task = 'classification',\n",
- " debug_log = 'automl_errors.log',\n",
- " path=project_folder,\n",
- " run_configuration=conda_run_config,\n",
- " #compute_target = dsvm_compute,\n",
- " data_script = project_folder + \"/get_data.py\",\n",
- " **automl_settings\n",
- " )"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "For remote runs the execution is asynchronous, so you will see the iterations get populated as they complete. You can interact with the widgets/models even when the experiment is running to retreive the best model up to that point. Once you are satisfied with the model you can cancel a particular iteration or the whole run."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "remote_run = experiment.submit(automl_config, show_output=False)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "remote_run"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Results\n",
- "#### Widget for monitoring runs\n",
- "\n",
- "The widget will sit on \"loading\" until the first iteration completed, then you will see an auto-updating graph and table show up. It refreshed once per minute, so you should see the graph update as child runs complete.\n",
- "\n",
- "You can click on a pipeline to see run properties and output logs. Logs are also available on the DSVM under /tmp/azureml_run/{iterationid}/azureml-logs\n",
- "\n",
- "NOTE: The widget displays a link at the bottom. This links to a web-ui to explore the individual run details."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.widgets import RunDetails\n",
- "RunDetails(remote_run).show() "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Wait until the run finishes.\n",
- "remote_run.wait_for_completion(show_output = True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "\n",
- "#### Retrieve All Child Runs\n",
- "You can also use sdk methods to fetch all the child runs and see individual metrics that we log. "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "children = list(remote_run.get_children())\n",
- "metricslist = {}\n",
- "for run in children:\n",
- " properties = run.get_properties()\n",
- " metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)} \n",
- " metricslist[int(properties['iteration'])] = metrics\n",
- "\n",
- "rundata = pd.DataFrame(metricslist).sort_index(1)\n",
- "rundata"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Canceling Runs\n",
- "You can cancel ongoing remote runs using the *cancel()* and *cancel_iteration()* functions"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Cancel the ongoing experiment and stop scheduling new iterations\n",
- "# remote_run.cancel()\n",
- "\n",
- "# Cancel iteration 1 and move onto iteration 2\n",
- "# remote_run.cancel_iteration(1)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Pre-process cache cleanup\n",
- "The preprocess data gets cache at user default file store. When the run is completed the cache can be cleaned by running below cell"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "remote_run.clean_preprocessor_cache()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Retrieve the Best Model\n",
- "\n",
- "Below we select the best pipeline from our iterations. The *get_output* method returns the best run and the fitted model. There are overloads on *get_output* that allow you to retrieve the best run and fitted model for *any* logged metric or a particular *iteration*."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "best_run, fitted_model = remote_run.get_output()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Best Model based on any other metric"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# lookup_metric = \"accuracy\"\n",
- "# best_run, fitted_model = remote_run.get_output(metric=lookup_metric)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Model from a specific iteration"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# iteration = 1\n",
- "# best_run, fitted_model = remote_run.get_output(iteration=iteration)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Test\n"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Load test data.\n",
- "from pandas_ml import ConfusionMatrix\n",
- "\n",
- "data_test = fetch_20newsgroups(subset = 'test', categories = categories,\n",
- " shuffle = True, random_state = 42,\n",
- " remove = remove)\n",
- "\n",
- "X_test = np.array(data_test.data).reshape((len(data_test.data),1))\n",
- "y_test = data_test.target\n",
- "\n",
- "# Test our best pipeline.\n",
- "\n",
- "y_pred = fitted_model.predict(X_test)\n",
- "y_pred_strings = [data_test.target_names[i] for i in y_pred]\n",
- "y_test_strings = [data_test.target_names[i] for i in y_test]\n",
- "\n",
- "cm = ConfusionMatrix(y_test_strings, y_pred_strings)\n",
- "print(cm)\n",
- "cm.plot()"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "savitam"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Automated Machine Learning\n",
+ "_**Remote Execution with DataStore**_\n",
+ "\n",
+ "## Contents\n",
+ "1. [Introduction](#Introduction)\n",
+ "1. [Setup](#Setup)\n",
+ "1. [Data](#Data)\n",
+ "1. [Train](#Train)\n",
+ "1. [Results](#Results)\n",
+ "1. [Test](#Test)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Introduction\n",
+ "This sample accesses a data file on a remote DSVM through DataStore. Advantages of using data store are:\n",
+ "1. DataStore secures the access details.\n",
+ "2. DataStore supports read, write to blob and file store\n",
+ "3. AutoML natively supports copying data from DataStore to DSVM\n",
+ "\n",
+ "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
+ "\n",
+ "In this notebook you would see\n",
+ "1. Storing data in DataStore.\n",
+ "2. get_data returning data from DataStore."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Setup\n",
+ "\n",
+ "As part of the setup you have already created a Workspace. For AutoML you would need to create an Experiment. An Experiment is a named object in a Workspace, which is used to run experiments."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import logging\n",
+ "import os\n",
+ "import time\n",
+ "\n",
+ "import numpy as np\n",
+ "import pandas as pd\n",
+ "\n",
+ "import azureml.core\n",
+ "from azureml.core.compute import DsvmCompute\n",
+ "from azureml.core.experiment import Experiment\n",
+ "from azureml.core.workspace import Workspace\n",
+ "from azureml.train.automl import AutoMLConfig"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "ws = Workspace.from_config()\n",
+ "\n",
+ "# choose a name for experiment\n",
+ "experiment_name = 'automl-remote-datastore-file'\n",
+ "# project folder\n",
+ "project_folder = './sample_projects/automl-remote-datastore-file'\n",
+ "\n",
+ "experiment=Experiment(ws, experiment_name)\n",
+ "\n",
+ "output = {}\n",
+ "output['SDK version'] = azureml.core.VERSION\n",
+ "output['Subscription ID'] = ws.subscription_id\n",
+ "output['Workspace'] = ws.name\n",
+ "output['Resource Group'] = ws.resource_group\n",
+ "output['Location'] = ws.location\n",
+ "output['Project Directory'] = project_folder\n",
+ "output['Experiment Name'] = experiment.name\n",
+ "pd.set_option('display.max_colwidth', -1)\n",
+ "outputDf = pd.DataFrame(data = output, index = [''])\n",
+ "outputDf.T"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Opt-in diagnostics for better experience, quality, and security of future releases"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.telemetry import set_diagnostics_collection\n",
+ "set_diagnostics_collection(send_diagnostics=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create a Remote Linux DSVM\n",
+ "Note: If creation fails with a message about Marketplace purchase eligibilty, go to portal.azure.com, start creating DSVM there, and select \"Want to create programmatically\" to enable programmatic creation. Once you've enabled it, you can exit without actually creating VM.\n",
+ "\n",
+ "**Note**: By default SSH runs on port 22 and you don't need to specify it. But if for security reasons you can switch to a different port (such as 5022), you can append the port number to the address. [Read more](https://render.githubusercontent.com/documentation/sdk/ssh-issue.md) on this."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "compute_target_name = 'mydsvmc'\n",
+ "\n",
+ "try:\n",
+ " while ws.compute_targets[compute_target_name].provisioning_state == 'Creating':\n",
+ " time.sleep(1)\n",
+ " \n",
+ " dsvm_compute = DsvmCompute(workspace=ws, name=compute_target_name)\n",
+ " print('found existing:', dsvm_compute.name)\n",
+ "except:\n",
+ " dsvm_config = DsvmCompute.provisioning_configuration(vm_size=\"Standard_D2_v2\")\n",
+ " dsvm_compute = DsvmCompute.create(ws, name=compute_target_name, provisioning_configuration=dsvm_config)\n",
+ " dsvm_compute.wait_for_completion(show_output=True)\n",
+ " print(\"Waiting one minute for ssh to be accessible\")\n",
+ " time.sleep(60) # Wait for ssh to be accessible"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Data\n",
+ "\n",
+ "### Copy data file to local\n",
+ "\n",
+ "Download the data file.\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "if not os.path.isdir('data'):\n",
+ " os.mkdir('data') "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from sklearn.datasets import fetch_20newsgroups\n",
+ "import csv\n",
+ "\n",
+ "remove = ('headers', 'footers', 'quotes')\n",
+ "categories = [\n",
+ " 'alt.atheism',\n",
+ " 'talk.religion.misc',\n",
+ " 'comp.graphics',\n",
+ " 'sci.space',\n",
+ " ]\n",
+ "data_train = fetch_20newsgroups(subset = 'train', categories = categories,\n",
+ " shuffle = True, random_state = 42,\n",
+ " remove = remove)\n",
+ " \n",
+ "pd.DataFrame(data_train.data).to_csv(\"data/X_train.tsv\", index=False, header=False, quoting=csv.QUOTE_ALL, sep=\"\\t\")\n",
+ "pd.DataFrame(data_train.target).to_csv(\"data/y_train.tsv\", index=False, header=False, sep=\"\\t\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Upload data to the cloud"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Now make the data accessible remotely by uploading that data from your local machine into Azure so it can be accessed for remote training. The datastore is a convenient construct associated with your workspace for you to upload/download data, and interact with it from your remote compute targets. It is backed by Azure blob storage account.\n",
+ "\n",
+ "The data.tsv files are uploaded into a directory named data at the root of the datastore."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "#blob_datastore = Datastore(ws, blob_datastore_name)\n",
+ "ds = ws.get_default_datastore()\n",
+ "print(ds.datastore_type, ds.account_name, ds.container_name)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# ds.upload_files(\"data.tsv\")\n",
+ "ds.upload(src_dir='./data', target_path='data', overwrite=True, show_progress=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Configure & Run\n",
+ "\n",
+ "First let's create a DataReferenceConfigruation object to inform the system what data folder to download to the compute target.\n",
+ "The path_on_compute should be an absolute path to ensure that the data files are downloaded only once. The get_data method should use this same path to access the data files."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.runconfig import DataReferenceConfiguration\n",
+ "dr = DataReferenceConfiguration(datastore_name=ds.name, \n",
+ " path_on_datastore='data', \n",
+ " path_on_compute='/tmp/azureml_runs',\n",
+ " mode='download', # download files from datastore to compute target\n",
+ " overwrite=False)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.runconfig import RunConfiguration\n",
+ "from azureml.core.conda_dependencies import CondaDependencies\n",
+ "\n",
+ "# create a new RunConfig object\n",
+ "conda_run_config = RunConfiguration(framework=\"python\")\n",
+ "\n",
+ "# Set compute target to the Linux DSVM\n",
+ "conda_run_config.target = dsvm_compute\n",
+ "# set the data reference of the run coonfiguration\n",
+ "conda_run_config.data_references = {ds.name: dr}\n",
+ "\n",
+ "cd = CondaDependencies.create(pip_packages=['azureml-sdk[automl]'], conda_packages=['numpy'])\n",
+ "conda_run_config.environment.python.conda_dependencies = cd"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create Get Data File\n",
+ "For remote executions you should author a get_data.py file containing a get_data() function. This file should be in the root directory of the project. You can encapsulate code to read data either from a blob storage or local disk in this file.\n",
+ "\n",
+ "The *get_data()* function returns a [dictionary](README.md#getdata).\n",
+ "\n",
+ "The read_csv uses the path_on_compute value specified in the DataReferenceConfiguration call plus the path_on_datastore folder and then the actual file name."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "if not os.path.exists(project_folder):\n",
+ " os.makedirs(project_folder)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%writefile $project_folder/get_data.py\n",
+ "\n",
+ "import pandas as pd\n",
+ "\n",
+ "def get_data():\n",
+ " X_train = pd.read_csv(\"/tmp/azureml_runs/data/X_train.tsv\", delimiter=\"\\t\", header=None, quotechar='\"')\n",
+ " y_train = pd.read_csv(\"/tmp/azureml_runs/data/y_train.tsv\", delimiter=\"\\t\", header=None, quotechar='\"')\n",
+ "\n",
+ " return { \"X\" : X_train.values, \"y\" : y_train[0].values }"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Train\n",
+ "\n",
+ "You can specify automl_settings as **kwargs** as well. Also note that you can use the get_data() symantic for local excutions too. \n",
+ "\n",
+ "Note: For Remote DSVM and Batch AI you cannot pass Numpy arrays directly to AutoMLConfig.\n",
+ "\n",
+ "|Property|Description|\n",
+ "|-|-|\n",
+ "|**primary_metric**|This is the metric that you want to optimize. Classification supports the following primary metrics:
accuracy
AUC_weighted
average_precision_score_weighted
norm_macro_recall
precision_score_weighted|\n",
+ "|**iteration_timeout_minutes**|Time limit in minutes for each iteration|\n",
+ "|**iterations**|Number of iterations. In each iteration Auto ML trains a specific pipeline with the data|\n",
+ "|**n_cross_validations**|Number of cross validation splits|\n",
+ "|**max_concurrent_iterations**|Max number of iterations that would be executed in parallel. This should be less than the number of cores on the DSVM\n",
+ "|**preprocess**| *True/False*
Setting this to *True* enables Auto ML to perform preprocessing
on the input to handle *missing data*, and perform some common *feature extraction*|\n",
+ "|**enable_cache**|Setting this to *True* enables preprocess done once and reuse the same preprocessed data for all the iterations. Default value is True.|\n",
+ "|**max_cores_per_iteration**| Indicates how many cores on the compute target would be used to train a single pipeline.
Default is *1*, you can set it to *-1* to use all cores|"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "automl_settings = {\n",
+ " \"iteration_timeout_minutes\": 60,\n",
+ " \"iterations\": 4,\n",
+ " \"n_cross_validations\": 5,\n",
+ " \"primary_metric\": 'AUC_weighted',\n",
+ " \"preprocess\": True,\n",
+ " \"max_cores_per_iteration\": 1,\n",
+ " \"verbosity\": logging.INFO\n",
+ "}\n",
+ "automl_config = AutoMLConfig(task = 'classification',\n",
+ " debug_log = 'automl_errors.log',\n",
+ " path=project_folder,\n",
+ " run_configuration=conda_run_config,\n",
+ " #compute_target = dsvm_compute,\n",
+ " data_script = project_folder + \"/get_data.py\",\n",
+ " **automl_settings\n",
+ " )"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "For remote runs the execution is asynchronous, so you will see the iterations get populated as they complete. You can interact with the widgets/models even when the experiment is running to retreive the best model up to that point. Once you are satisfied with the model you can cancel a particular iteration or the whole run."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "remote_run = experiment.submit(automl_config, show_output=False)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "remote_run"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Results\n",
+ "#### Widget for monitoring runs\n",
+ "\n",
+ "The widget will sit on \"loading\" until the first iteration completed, then you will see an auto-updating graph and table show up. It refreshed once per minute, so you should see the graph update as child runs complete.\n",
+ "\n",
+ "You can click on a pipeline to see run properties and output logs. Logs are also available on the DSVM under /tmp/azureml_run/{iterationid}/azureml-logs\n",
+ "\n",
+ "NOTE: The widget displays a link at the bottom. This links to a web-ui to explore the individual run details."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.widgets import RunDetails\n",
+ "RunDetails(remote_run).show() "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Wait until the run finishes.\n",
+ "remote_run.wait_for_completion(show_output = True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "\n",
+ "#### Retrieve All Child Runs\n",
+ "You can also use sdk methods to fetch all the child runs and see individual metrics that we log. "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "children = list(remote_run.get_children())\n",
+ "metricslist = {}\n",
+ "for run in children:\n",
+ " properties = run.get_properties()\n",
+ " metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)} \n",
+ " metricslist[int(properties['iteration'])] = metrics\n",
+ "\n",
+ "rundata = pd.DataFrame(metricslist).sort_index(1)\n",
+ "rundata"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Canceling Runs\n",
+ "You can cancel ongoing remote runs using the *cancel()* and *cancel_iteration()* functions"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Cancel the ongoing experiment and stop scheduling new iterations\n",
+ "# remote_run.cancel()\n",
+ "\n",
+ "# Cancel iteration 1 and move onto iteration 2\n",
+ "# remote_run.cancel_iteration(1)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Pre-process cache cleanup\n",
+ "The preprocess data gets cache at user default file store. When the run is completed the cache can be cleaned by running below cell"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "remote_run.clean_preprocessor_cache()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Retrieve the Best Model\n",
+ "\n",
+ "Below we select the best pipeline from our iterations. The *get_output* method returns the best run and the fitted model. There are overloads on *get_output* that allow you to retrieve the best run and fitted model for *any* logged metric or a particular *iteration*."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "best_run, fitted_model = remote_run.get_output()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Best Model based on any other metric"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# lookup_metric = \"accuracy\"\n",
+ "# best_run, fitted_model = remote_run.get_output(metric=lookup_metric)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Model from a specific iteration"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# iteration = 1\n",
+ "# best_run, fitted_model = remote_run.get_output(iteration=iteration)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Test\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Load test data.\n",
+ "from pandas_ml import ConfusionMatrix\n",
+ "\n",
+ "data_test = fetch_20newsgroups(subset = 'test', categories = categories,\n",
+ " shuffle = True, random_state = 42,\n",
+ " remove = remove)\n",
+ "\n",
+ "X_test = np.array(data_test.data).reshape((len(data_test.data),1))\n",
+ "y_test = data_test.target\n",
+ "\n",
+ "# Test our best pipeline.\n",
+ "\n",
+ "y_pred = fitted_model.predict(X_test)\n",
+ "y_pred_strings = [data_test.target_names[i] for i in y_pred]\n",
+ "y_test_strings = [data_test.target_names[i] for i in y_test]\n",
+ "\n",
+ "cm = ConfusionMatrix(y_test_strings, y_pred_strings)\n",
+ "print(cm)\n",
+ "cm.plot()"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "savitam"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.6"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/automated-machine-learning/remote-execution/auto-ml-remote-execution.ipynb b/how-to-use-azureml/automated-machine-learning/remote-execution/auto-ml-remote-execution.ipynb
index 927d8ccc..b6f4ceb5 100644
--- a/how-to-use-azureml/automated-machine-learning/remote-execution/auto-ml-remote-execution.ipynb
+++ b/how-to-use-azureml/automated-machine-learning/remote-execution/auto-ml-remote-execution.ipynb
@@ -1,527 +1,525 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Automated Machine Learning\n",
- "_**Remote Execution using DSVM (Ubuntu)**_\n",
- "\n",
- "## Contents\n",
- "1. [Introduction](#Introduction)\n",
- "1. [Setup](#Setup)\n",
- "1. [Data](#Data)\n",
- "1. [Train](#Train)\n",
- "1. [Results](#Results)\n",
- "1. [Test](#Test)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Introduction\n",
- "In this example we use the scikit-learn's [digit dataset](http://scikit-learn.org/stable/datasets/index.html#optical-recognition-of-handwritten-digits-dataset) to showcase how you can use AutoML for a simple classification problem.\n",
- "\n",
- "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
- "\n",
- "In this notebook you wiil learn how to:\n",
- "1. Create an `Experiment` in an existing `Workspace`.\n",
- "2. Attach an existing DSVM to a workspace.\n",
- "3. Configure AutoML using `AutoMLConfig`.\n",
- "4. Train the model using the DSVM.\n",
- "5. Explore the results.\n",
- "6. Test the best fitted model.\n",
- "\n",
- "In addition, this notebook showcases the following features:\n",
- "- **Parallel** executions for iterations\n",
- "- **Asynchronous** tracking of progress\n",
- "- **Cancellation** of individual iterations or the entire run\n",
- "- Retrieving models for any iteration or logged metric\n",
- "- Specifying AutoML settings as `**kwargs`"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Setup\n",
- "\n",
- "As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import logging\n",
- "import os\n",
- "import random\n",
- "import time\n",
- "\n",
- "from matplotlib import pyplot as plt\n",
- "from matplotlib.pyplot import imshow\n",
- "import numpy as np\n",
- "import pandas as pd\n",
- "from sklearn import datasets\n",
- "\n",
- "import azureml.core\n",
- "from azureml.core.experiment import Experiment\n",
- "from azureml.core.workspace import Workspace\n",
- "from azureml.train.automl import AutoMLConfig\n",
- "from azureml.train.automl.run import AutoMLRun"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "ws = Workspace.from_config()\n",
- "\n",
- "# Choose a name for the run history container in the workspace.\n",
- "experiment_name = 'automl-remote-dsvm'\n",
- "project_folder = './sample_projects/automl-remote-dsvm'\n",
- "\n",
- "experiment = Experiment(ws, experiment_name)\n",
- "\n",
- "output = {}\n",
- "output['SDK version'] = azureml.core.VERSION\n",
- "output['Subscription ID'] = ws.subscription_id\n",
- "output['Workspace Name'] = ws.name\n",
- "output['Resource Group'] = ws.resource_group\n",
- "output['Location'] = ws.location\n",
- "output['Project Directory'] = project_folder\n",
- "output['Experiment Name'] = experiment.name\n",
- "pd.set_option('display.max_colwidth', -1)\n",
- "pd.DataFrame(data = output, index = ['']).T"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Opt-in diagnostics for better experience, quality, and security of future releases."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.telemetry import set_diagnostics_collection\n",
- "set_diagnostics_collection(send_diagnostics = True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create a Remote Linux DSVM\n",
- "**Note:** If creation fails with a message about Marketplace purchase eligibilty, start creation of a DSVM through the [Azure portal](https://portal.azure.com), and select \"Want to create programmatically\" to enable programmatic creation. Once you've enabled this setting, you can exit the portal without actually creating the DSVM, and creation of the DSVM through the notebook should work.\n"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.compute import DsvmCompute\n",
- "\n",
- "dsvm_name = 'mydsvma'\n",
- "try:\n",
- " dsvm_compute = DsvmCompute(ws, dsvm_name)\n",
- " print('Found an existing DSVM.')\n",
- "except:\n",
- " print('Creating a new DSVM.')\n",
- " dsvm_config = DsvmCompute.provisioning_configuration(vm_size = \"Standard_D2s_v3\")\n",
- " dsvm_compute = DsvmCompute.create(ws, name = dsvm_name, provisioning_configuration = dsvm_config)\n",
- " dsvm_compute.wait_for_completion(show_output = True)\n",
- " print(\"Waiting one minute for ssh to be accessible\")\n",
- " time.sleep(60) # Wait for ssh to be accessible"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.runconfig import RunConfiguration\n",
- "from azureml.core.conda_dependencies import CondaDependencies\n",
- "\n",
- "# create a new RunConfig object\n",
- "conda_run_config = RunConfiguration(framework=\"python\")\n",
- "\n",
- "# Set compute target to the Linux DSVM\n",
- "conda_run_config.target = dsvm_compute\n",
- "\n",
- "cd = CondaDependencies.create(pip_packages=['azureml-sdk[automl]'], conda_packages=['numpy'])\n",
- "conda_run_config.environment.python.conda_dependencies = cd"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Data\n",
- "For remote executions you should author a `get_data.py` file containing a `get_data()` function. This file should be in the root directory of the project. You can encapsulate code to read data either from a blob storage or local disk in this file.\n",
- "In this example, the `get_data()` function returns data using scikit-learn's [load_digits](http://scikit-learn.org/stable/modules/generated/sklearn.datasets.load_digits.html) method."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "if not os.path.exists(project_folder):\n",
- " os.makedirs(project_folder)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%writefile $project_folder/get_data.py\n",
- "\n",
- "from sklearn import datasets\n",
- "from scipy import sparse\n",
- "import numpy as np\n",
- "\n",
- "def get_data():\n",
- " \n",
- " digits = datasets.load_digits()\n",
- " X_train = digits.data[100:,:]\n",
- " y_train = digits.target[100:]\n",
- "\n",
- " return { \"X\" : X_train, \"y\" : y_train }"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Train\n",
- "\n",
- "You can specify `automl_settings` as `**kwargs` as well. Also note that you can use a `get_data()` function for local excutions too.\n",
- "\n",
- "**Note:** When using Remote DSVM, you can't pass Numpy arrays directly to the fit method.\n",
- "\n",
- "|Property|Description|\n",
- "|-|-|\n",
- "|**primary_metric**|This is the metric that you want to optimize. Classification supports the following primary metrics:
accuracy
AUC_weighted
average_precision_score_weighted
norm_macro_recall
precision_score_weighted|\n",
- "|**iteration_timeout_minutes**|Time limit in minutes for each iteration.|\n",
- "|**iterations**|Number of iterations. In each iteration AutoML trains a specific pipeline with the data.|\n",
- "|**n_cross_validations**|Number of cross validation splits.|\n",
- "|**max_concurrent_iterations**|Maximum number of iterations to execute in parallel. This should be less than the number of cores on the DSVM.|"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "automl_settings = {\n",
- " \"iteration_timeout_minutes\": 10,\n",
- " \"iterations\": 20,\n",
- " \"n_cross_validations\": 5,\n",
- " \"primary_metric\": 'AUC_weighted',\n",
- " \"preprocess\": False,\n",
- " \"max_concurrent_iterations\": 2,\n",
- " \"verbosity\": logging.INFO\n",
- "}\n",
- "\n",
- "automl_config = AutoMLConfig(task = 'classification',\n",
- " debug_log = 'automl_errors.log',\n",
- " path = project_folder, \n",
- " run_configuration=conda_run_config,\n",
- " data_script = project_folder + \"/get_data.py\",\n",
- " **automl_settings\n",
- " )\n"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "**Note:** The first run on a new DSVM may take several minutes to prepare the environment."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Call the `submit` method on the experiment object and pass the run configuration. For remote runs the execution is asynchronous, so you will see the iterations get populated as they complete. You can interact with the widgets and models even when the experiment is running to retrieve the best model up to that point. Once you are satisfied with the model, you can cancel a particular iteration or the whole run.\n",
- "\n",
- "In this example, we specify `show_output = False` to suppress console output while the run is in progress."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "remote_run = experiment.submit(automl_config, show_output = False)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "remote_run"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Results\n",
- "\n",
- "#### Loading Executed Runs\n",
- "In case you need to load a previously executed run, enable the cell below and replace the `run_id` value."
- ]
- },
- {
- "cell_type": "raw",
- "metadata": {},
- "source": [
- "remote_run = AutoMLRun(experiment=experiment, run_id = 'AutoML_480d3ed6-fc94-44aa-8f4e-0b945db9d3ef')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Widget for Monitoring Runs\n",
- "\n",
- "The widget will first report a \"loading\" status while running the first iteration. After completing the first iteration, an auto-updating graph and table will be shown. The widget will refresh once per minute, so you should see the graph update as child runs complete.\n",
- "\n",
- "You can click on a pipeline to see run properties and output logs. Logs are also available on the DSVM under `/tmp/azureml_run/{iterationid}/azureml-logs`\n",
- "\n",
- "**Note:** The widget displays a link at the bottom. Use this link to open a web interface to explore the individual run details."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.widgets import RunDetails\n",
- "RunDetails(remote_run).show() "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Wait until the run finishes.\n",
- "remote_run.wait_for_completion(show_output = True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "\n",
- "#### Retrieve All Child Runs\n",
- "You can also use SDK methods to fetch all the child runs and see individual metrics that we log."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "children = list(remote_run.get_children())\n",
- "metricslist = {}\n",
- "for run in children:\n",
- " properties = run.get_properties()\n",
- " metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)} \n",
- " metricslist[int(properties['iteration'])] = metrics\n",
- "\n",
- "rundata = pd.DataFrame(metricslist).sort_index(1)\n",
- "rundata"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Cancelling Runs\n",
- "\n",
- "You can cancel ongoing remote runs using the `cancel` and `cancel_iteration` functions."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Cancel the ongoing experiment and stop scheduling new iterations.\n",
- "# remote_run.cancel()\n",
- "\n",
- "# Cancel iteration 1 and move onto iteration 2.\n",
- "# remote_run.cancel_iteration(1)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Retrieve the Best Model\n",
- "\n",
- "Below we select the best pipeline from our iterations. The `get_output` method returns the best run and the fitted model. The Model includes the pipeline and any pre-processing. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "best_run, fitted_model = remote_run.get_output()\n",
- "print(best_run)\n",
- "print(fitted_model)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Best Model Based on Any Other Metric\n",
- "Show the run and the model which has the smallest `log_loss` value:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "lookup_metric = \"log_loss\"\n",
- "best_run, fitted_model = remote_run.get_output(metric = lookup_metric)\n",
- "print(best_run)\n",
- "print(fitted_model)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Model from a Specific Iteration\n",
- "Show the run and the model from the third iteration:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "iteration = 3\n",
- "third_run, third_model = remote_run.get_output(iteration = iteration)\n",
- "print(third_run)\n",
- "print(third_model)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Test\n",
- "\n",
- "#### Load Test Data"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "digits = datasets.load_digits()\n",
- "X_test = digits.data[:10, :]\n",
- "y_test = digits.target[:10]\n",
- "images = digits.images[:10]"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Test Our Best Fitted Model"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Randomly select digits and test.\n",
- "for index in np.random.choice(len(y_test), 2, replace = False):\n",
- " print(index)\n",
- " predicted = fitted_model.predict(X_test[index:index + 1])[0]\n",
- " label = y_test[index]\n",
- " title = \"Label value = %d Predicted value = %d \" % (label, predicted)\n",
- " fig = plt.figure(1, figsize=(3,3))\n",
- " ax1 = fig.add_axes((0,0,.8,.8))\n",
- " ax1.set_title(title)\n",
- " plt.imshow(images[index], cmap = plt.cm.gray_r, interpolation = 'nearest')\n",
- " plt.show()"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "savitam"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Automated Machine Learning\n",
+ "_**Remote Execution using DSVM (Ubuntu)**_\n",
+ "\n",
+ "## Contents\n",
+ "1. [Introduction](#Introduction)\n",
+ "1. [Setup](#Setup)\n",
+ "1. [Data](#Data)\n",
+ "1. [Train](#Train)\n",
+ "1. [Results](#Results)\n",
+ "1. [Test](#Test)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Introduction\n",
+ "In this example we use the scikit-learn's [digit dataset](http://scikit-learn.org/stable/datasets/index.html#optical-recognition-of-handwritten-digits-dataset) to showcase how you can use AutoML for a simple classification problem.\n",
+ "\n",
+ "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
+ "\n",
+ "In this notebook you wiil learn how to:\n",
+ "1. Create an `Experiment` in an existing `Workspace`.\n",
+ "2. Attach an existing DSVM to a workspace.\n",
+ "3. Configure AutoML using `AutoMLConfig`.\n",
+ "4. Train the model using the DSVM.\n",
+ "5. Explore the results.\n",
+ "6. Test the best fitted model.\n",
+ "\n",
+ "In addition, this notebook showcases the following features:\n",
+ "- **Parallel** executions for iterations\n",
+ "- **Asynchronous** tracking of progress\n",
+ "- **Cancellation** of individual iterations or the entire run\n",
+ "- Retrieving models for any iteration or logged metric\n",
+ "- Specifying AutoML settings as `**kwargs`"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Setup\n",
+ "\n",
+ "As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import logging\n",
+ "import os\n",
+ "import time\n",
+ "\n",
+ "from matplotlib import pyplot as plt\n",
+ "import numpy as np\n",
+ "import pandas as pd\n",
+ "from sklearn import datasets\n",
+ "\n",
+ "import azureml.core\n",
+ "from azureml.core.experiment import Experiment\n",
+ "from azureml.core.workspace import Workspace\n",
+ "from azureml.train.automl import AutoMLConfig"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "ws = Workspace.from_config()\n",
+ "\n",
+ "# Choose a name for the run history container in the workspace.\n",
+ "experiment_name = 'automl-remote-dsvm'\n",
+ "project_folder = './sample_projects/automl-remote-dsvm'\n",
+ "\n",
+ "experiment = Experiment(ws, experiment_name)\n",
+ "\n",
+ "output = {}\n",
+ "output['SDK version'] = azureml.core.VERSION\n",
+ "output['Subscription ID'] = ws.subscription_id\n",
+ "output['Workspace Name'] = ws.name\n",
+ "output['Resource Group'] = ws.resource_group\n",
+ "output['Location'] = ws.location\n",
+ "output['Project Directory'] = project_folder\n",
+ "output['Experiment Name'] = experiment.name\n",
+ "pd.set_option('display.max_colwidth', -1)\n",
+ "outputDf = pd.DataFrame(data = output, index = [''])\n",
+ "outputDf.T"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Opt-in diagnostics for better experience, quality, and security of future releases."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.telemetry import set_diagnostics_collection\n",
+ "set_diagnostics_collection(send_diagnostics = True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create a Remote Linux DSVM\n",
+ "**Note:** If creation fails with a message about Marketplace purchase eligibilty, start creation of a DSVM through the [Azure portal](https://portal.azure.com), and select \"Want to create programmatically\" to enable programmatic creation. Once you've enabled this setting, you can exit the portal without actually creating the DSVM, and creation of the DSVM through the notebook should work.\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.compute import DsvmCompute\n",
+ "\n",
+ "dsvm_name = 'mydsvma'\n",
+ "try:\n",
+ " dsvm_compute = DsvmCompute(ws, dsvm_name)\n",
+ " print('Found an existing DSVM.')\n",
+ "except:\n",
+ " print('Creating a new DSVM.')\n",
+ " dsvm_config = DsvmCompute.provisioning_configuration(vm_size = \"Standard_D2s_v3\")\n",
+ " dsvm_compute = DsvmCompute.create(ws, name = dsvm_name, provisioning_configuration = dsvm_config)\n",
+ " dsvm_compute.wait_for_completion(show_output = True)\n",
+ " print(\"Waiting one minute for ssh to be accessible\")\n",
+ " time.sleep(60) # Wait for ssh to be accessible"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.runconfig import RunConfiguration\n",
+ "from azureml.core.conda_dependencies import CondaDependencies\n",
+ "\n",
+ "# create a new RunConfig object\n",
+ "conda_run_config = RunConfiguration(framework=\"python\")\n",
+ "\n",
+ "# Set compute target to the Linux DSVM\n",
+ "conda_run_config.target = dsvm_compute\n",
+ "\n",
+ "cd = CondaDependencies.create(pip_packages=['azureml-sdk[automl]'], conda_packages=['numpy'])\n",
+ "conda_run_config.environment.python.conda_dependencies = cd"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Data\n",
+ "For remote executions you should author a `get_data.py` file containing a `get_data()` function. This file should be in the root directory of the project. You can encapsulate code to read data either from a blob storage or local disk in this file.\n",
+ "In this example, the `get_data()` function returns data using scikit-learn's [load_digits](http://scikit-learn.org/stable/modules/generated/sklearn.datasets.load_digits.html) method."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "if not os.path.exists(project_folder):\n",
+ " os.makedirs(project_folder)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%writefile $project_folder/get_data.py\n",
+ "\n",
+ "from sklearn import datasets\n",
+ "from scipy import sparse\n",
+ "import numpy as np\n",
+ "\n",
+ "def get_data():\n",
+ " \n",
+ " digits = datasets.load_digits()\n",
+ " X_train = digits.data[100:,:]\n",
+ " y_train = digits.target[100:]\n",
+ "\n",
+ " return { \"X\" : X_train, \"y\" : y_train }"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Train\n",
+ "\n",
+ "You can specify `automl_settings` as `**kwargs` as well. Also note that you can use a `get_data()` function for local excutions too.\n",
+ "\n",
+ "**Note:** When using Remote DSVM, you can't pass Numpy arrays directly to the fit method.\n",
+ "\n",
+ "|Property|Description|\n",
+ "|-|-|\n",
+ "|**primary_metric**|This is the metric that you want to optimize. Classification supports the following primary metrics:
accuracy
AUC_weighted
average_precision_score_weighted
norm_macro_recall
precision_score_weighted|\n",
+ "|**iteration_timeout_minutes**|Time limit in minutes for each iteration.|\n",
+ "|**iterations**|Number of iterations. In each iteration AutoML trains a specific pipeline with the data.|\n",
+ "|**n_cross_validations**|Number of cross validation splits.|\n",
+ "|**max_concurrent_iterations**|Maximum number of iterations to execute in parallel. This should be less than the number of cores on the DSVM.|"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "automl_settings = {\n",
+ " \"iteration_timeout_minutes\": 10,\n",
+ " \"iterations\": 20,\n",
+ " \"n_cross_validations\": 5,\n",
+ " \"primary_metric\": 'AUC_weighted',\n",
+ " \"preprocess\": False,\n",
+ " \"max_concurrent_iterations\": 2,\n",
+ " \"verbosity\": logging.INFO\n",
+ "}\n",
+ "\n",
+ "automl_config = AutoMLConfig(task = 'classification',\n",
+ " debug_log = 'automl_errors.log',\n",
+ " path = project_folder, \n",
+ " run_configuration=conda_run_config,\n",
+ " data_script = project_folder + \"/get_data.py\",\n",
+ " **automl_settings\n",
+ " )\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "**Note:** The first run on a new DSVM may take several minutes to prepare the environment."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Call the `submit` method on the experiment object and pass the run configuration. For remote runs the execution is asynchronous, so you will see the iterations get populated as they complete. You can interact with the widgets and models even when the experiment is running to retrieve the best model up to that point. Once you are satisfied with the model, you can cancel a particular iteration or the whole run.\n",
+ "\n",
+ "In this example, we specify `show_output = False` to suppress console output while the run is in progress."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "remote_run = experiment.submit(automl_config, show_output = False)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "remote_run"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Results\n",
+ "\n",
+ "#### Loading Executed Runs\n",
+ "In case you need to load a previously executed run, enable the cell below and replace the `run_id` value."
+ ]
+ },
+ {
+ "cell_type": "raw",
+ "metadata": {},
+ "source": [
+ "remote_run = AutoMLRun(experiment=experiment, run_id = 'AutoML_480d3ed6-fc94-44aa-8f4e-0b945db9d3ef')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Widget for Monitoring Runs\n",
+ "\n",
+ "The widget will first report a \"loading\" status while running the first iteration. After completing the first iteration, an auto-updating graph and table will be shown. The widget will refresh once per minute, so you should see the graph update as child runs complete.\n",
+ "\n",
+ "You can click on a pipeline to see run properties and output logs. Logs are also available on the DSVM under `/tmp/azureml_run/{iterationid}/azureml-logs`\n",
+ "\n",
+ "**Note:** The widget displays a link at the bottom. Use this link to open a web interface to explore the individual run details."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.widgets import RunDetails\n",
+ "RunDetails(remote_run).show() "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Wait until the run finishes.\n",
+ "remote_run.wait_for_completion(show_output = True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "\n",
+ "#### Retrieve All Child Runs\n",
+ "You can also use SDK methods to fetch all the child runs and see individual metrics that we log."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "children = list(remote_run.get_children())\n",
+ "metricslist = {}\n",
+ "for run in children:\n",
+ " properties = run.get_properties()\n",
+ " metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)} \n",
+ " metricslist[int(properties['iteration'])] = metrics\n",
+ "\n",
+ "rundata = pd.DataFrame(metricslist).sort_index(1)\n",
+ "rundata"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Cancelling Runs\n",
+ "\n",
+ "You can cancel ongoing remote runs using the `cancel` and `cancel_iteration` functions."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Cancel the ongoing experiment and stop scheduling new iterations.\n",
+ "# remote_run.cancel()\n",
+ "\n",
+ "# Cancel iteration 1 and move onto iteration 2.\n",
+ "# remote_run.cancel_iteration(1)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Retrieve the Best Model\n",
+ "\n",
+ "Below we select the best pipeline from our iterations. The `get_output` method returns the best run and the fitted model. The Model includes the pipeline and any pre-processing. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "best_run, fitted_model = remote_run.get_output()\n",
+ "print(best_run)\n",
+ "print(fitted_model)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Best Model Based on Any Other Metric\n",
+ "Show the run and the model which has the smallest `log_loss` value:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "lookup_metric = \"log_loss\"\n",
+ "best_run, fitted_model = remote_run.get_output(metric = lookup_metric)\n",
+ "print(best_run)\n",
+ "print(fitted_model)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Model from a Specific Iteration\n",
+ "Show the run and the model from the third iteration:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "iteration = 3\n",
+ "third_run, third_model = remote_run.get_output(iteration = iteration)\n",
+ "print(third_run)\n",
+ "print(third_model)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Test\n",
+ "\n",
+ "#### Load Test Data"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "digits = datasets.load_digits()\n",
+ "X_test = digits.data[:10, :]\n",
+ "y_test = digits.target[:10]\n",
+ "images = digits.images[:10]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Test Our Best Fitted Model"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Randomly select digits and test.\n",
+ "for index in np.random.choice(len(y_test), 2, replace = False):\n",
+ " print(index)\n",
+ " predicted = fitted_model.predict(X_test[index:index + 1])[0]\n",
+ " label = y_test[index]\n",
+ " title = \"Label value = %d Predicted value = %d \" % (label, predicted)\n",
+ " fig = plt.figure(1, figsize=(3,3))\n",
+ " ax1 = fig.add_axes((0,0,.8,.8))\n",
+ " ax1.set_title(title)\n",
+ " plt.imshow(images[index], cmap = plt.cm.gray_r, interpolation = 'nearest')\n",
+ " plt.show()"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "savitam"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.6"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/automated-machine-learning/sample-weight/auto-ml-sample-weight.ipynb b/how-to-use-azureml/automated-machine-learning/sample-weight/auto-ml-sample-weight.ipynb
index 65ad698d..f64c282d 100644
--- a/how-to-use-azureml/automated-machine-learning/sample-weight/auto-ml-sample-weight.ipynb
+++ b/how-to-use-azureml/automated-machine-learning/sample-weight/auto-ml-sample-weight.ipynb
@@ -1,260 +1,257 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Automated Machine Learning\n",
- "_**Sample Weight**_\n",
- "\n",
- "## Contents\n",
- "1. [Introduction](#Introduction)\n",
- "1. [Setup](#Setup)\n",
- "1. [Train](#Train)\n",
- "1. [Test](#Test)\n"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Introduction\n",
- "In this example we use the scikit-learn's [digit dataset](http://scikit-learn.org/stable/datasets/index.html#optical-recognition-of-handwritten-digits-dataset) to showcase how you can use sample weight with AutoML. Sample weight is used where some sample values are more important than others.\n",
- "\n",
- "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
- "\n",
- "In this notebook you will learn how to configure AutoML to use `sample_weight` and you will see the difference sample weight makes to the test results."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Setup\n",
- "\n",
- "As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import logging\n",
- "import os\n",
- "import random\n",
- "\n",
- "from matplotlib import pyplot as plt\n",
- "from matplotlib.pyplot import imshow\n",
- "import numpy as np\n",
- "import pandas as pd\n",
- "from sklearn import datasets\n",
- "\n",
- "import azureml.core\n",
- "from azureml.core.experiment import Experiment\n",
- "from azureml.core.workspace import Workspace\n",
- "from azureml.train.automl import AutoMLConfig\n",
- "from azureml.train.automl.run import AutoMLRun"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "ws = Workspace.from_config()\n",
- "\n",
- "# Choose names for the regular and the sample weight experiments.\n",
- "experiment_name = 'non_sample_weight_experiment'\n",
- "sample_weight_experiment_name = 'sample_weight_experiment'\n",
- "\n",
- "project_folder = './sample_projects/automl-local-classification'\n",
- "\n",
- "experiment = Experiment(ws, experiment_name)\n",
- "sample_weight_experiment=Experiment(ws, sample_weight_experiment_name)\n",
- "\n",
- "output = {}\n",
- "output['SDK version'] = azureml.core.VERSION\n",
- "output['Subscription ID'] = ws.subscription_id\n",
- "output['Workspace Name'] = ws.name\n",
- "output['Resource Group'] = ws.resource_group\n",
- "output['Location'] = ws.location\n",
- "output['Project Directory'] = project_folder\n",
- "output['Experiment Name'] = experiment.name\n",
- "pd.set_option('display.max_colwidth', -1)\n",
- "pd.DataFrame(data = output, index = ['']).T"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Opt-in diagnostics for better experience, quality, and security of future releases."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.telemetry import set_diagnostics_collection\n",
- "set_diagnostics_collection(send_diagnostics = True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Train\n",
- "\n",
- "Instantiate two `AutoMLConfig` objects. One will be used with `sample_weight` and one without."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "digits = datasets.load_digits()\n",
- "X_train = digits.data[100:,:]\n",
- "y_train = digits.target[100:]\n",
- "\n",
- "# The example makes the sample weight 0.9 for the digit 4 and 0.1 for all other digits.\n",
- "# This makes the model more likely to classify as 4 if the image it not clear.\n",
- "sample_weight = np.array([(0.9 if x == 4 else 0.01) for x in y_train])\n",
- "\n",
- "automl_classifier = AutoMLConfig(task = 'classification',\n",
- " debug_log = 'automl_errors.log',\n",
- " primary_metric = 'AUC_weighted',\n",
- " iteration_timeout_minutes = 60,\n",
- " iterations = 10,\n",
- " n_cross_validations = 2,\n",
- " verbosity = logging.INFO,\n",
- " X = X_train, \n",
- " y = y_train,\n",
- " path = project_folder)\n",
- "\n",
- "automl_sample_weight = AutoMLConfig(task = 'classification',\n",
- " debug_log = 'automl_errors.log',\n",
- " primary_metric = 'AUC_weighted',\n",
- " iteration_timeout_minutes = 60,\n",
- " iterations = 10,\n",
- " n_cross_validations = 2,\n",
- " verbosity = logging.INFO,\n",
- " X = X_train, \n",
- " y = y_train,\n",
- " sample_weight = sample_weight,\n",
- " path = project_folder)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Call the `submit` method on the experiment objects and pass the run configuration. Execution of local runs is synchronous. Depending on the data and the number of iterations this can run for a while.\n",
- "In this example, we specify `show_output = True` to print currently running iterations to the console."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "local_run = experiment.submit(automl_classifier, show_output = True)\n",
- "sample_weight_run = sample_weight_experiment.submit(automl_sample_weight, show_output = True)\n",
- "\n",
- "best_run, fitted_model = local_run.get_output()\n",
- "best_run_sample_weight, fitted_model_sample_weight = sample_weight_run.get_output()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Test\n",
- "\n",
- "#### Load Test Data"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "digits = datasets.load_digits()\n",
- "X_test = digits.data[:100, :]\n",
- "y_test = digits.target[:100]\n",
- "images = digits.images[:100]"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Compare the Models\n",
- "The prediction from the sample weight model is more likely to correctly predict 4's. However, it is also more likely to predict 4 for some images that are not labelled as 4."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Randomly select digits and test.\n",
- "for index in range(0,len(y_test)):\n",
- " predicted = fitted_model.predict(X_test[index:index + 1])[0]\n",
- " predicted_sample_weight = fitted_model_sample_weight.predict(X_test[index:index + 1])[0]\n",
- " label = y_test[index]\n",
- " if predicted == 4 or predicted_sample_weight == 4 or label == 4:\n",
- " title = \"Label value = %d Predicted value = %d Prediced with sample weight = %d\" % (label, predicted, predicted_sample_weight)\n",
- " fig = plt.figure(1, figsize=(3,3))\n",
- " ax1 = fig.add_axes((0,0,.8,.8))\n",
- " ax1.set_title(title)\n",
- " plt.imshow(images[index], cmap = plt.cm.gray_r, interpolation = 'nearest')\n",
- " plt.show()"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "savitam"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Automated Machine Learning\n",
+ "_**Sample Weight**_\n",
+ "\n",
+ "## Contents\n",
+ "1. [Introduction](#Introduction)\n",
+ "1. [Setup](#Setup)\n",
+ "1. [Train](#Train)\n",
+ "1. [Test](#Test)\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Introduction\n",
+ "In this example we use the scikit-learn's [digit dataset](http://scikit-learn.org/stable/datasets/index.html#optical-recognition-of-handwritten-digits-dataset) to showcase how you can use sample weight with AutoML. Sample weight is used where some sample values are more important than others.\n",
+ "\n",
+ "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
+ "\n",
+ "In this notebook you will learn how to configure AutoML to use `sample_weight` and you will see the difference sample weight makes to the test results."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Setup\n",
+ "\n",
+ "As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import logging\n",
+ "\n",
+ "from matplotlib import pyplot as plt\n",
+ "import numpy as np\n",
+ "import pandas as pd\n",
+ "from sklearn import datasets\n",
+ "\n",
+ "import azureml.core\n",
+ "from azureml.core.experiment import Experiment\n",
+ "from azureml.core.workspace import Workspace\n",
+ "from azureml.train.automl import AutoMLConfig"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "ws = Workspace.from_config()\n",
+ "\n",
+ "# Choose names for the regular and the sample weight experiments.\n",
+ "experiment_name = 'non_sample_weight_experiment'\n",
+ "sample_weight_experiment_name = 'sample_weight_experiment'\n",
+ "\n",
+ "project_folder = './sample_projects/automl-local-classification'\n",
+ "\n",
+ "experiment = Experiment(ws, experiment_name)\n",
+ "sample_weight_experiment=Experiment(ws, sample_weight_experiment_name)\n",
+ "\n",
+ "output = {}\n",
+ "output['SDK version'] = azureml.core.VERSION\n",
+ "output['Subscription ID'] = ws.subscription_id\n",
+ "output['Workspace Name'] = ws.name\n",
+ "output['Resource Group'] = ws.resource_group\n",
+ "output['Location'] = ws.location\n",
+ "output['Project Directory'] = project_folder\n",
+ "output['Experiment Name'] = experiment.name\n",
+ "pd.set_option('display.max_colwidth', -1)\n",
+ "outputDf = pd.DataFrame(data = output, index = [''])\n",
+ "outputDf.T"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Opt-in diagnostics for better experience, quality, and security of future releases."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.telemetry import set_diagnostics_collection\n",
+ "set_diagnostics_collection(send_diagnostics = True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Train\n",
+ "\n",
+ "Instantiate two `AutoMLConfig` objects. One will be used with `sample_weight` and one without."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "digits = datasets.load_digits()\n",
+ "X_train = digits.data[100:,:]\n",
+ "y_train = digits.target[100:]\n",
+ "\n",
+ "# The example makes the sample weight 0.9 for the digit 4 and 0.1 for all other digits.\n",
+ "# This makes the model more likely to classify as 4 if the image it not clear.\n",
+ "sample_weight = np.array([(0.9 if x == 4 else 0.01) for x in y_train])\n",
+ "\n",
+ "automl_classifier = AutoMLConfig(task = 'classification',\n",
+ " debug_log = 'automl_errors.log',\n",
+ " primary_metric = 'AUC_weighted',\n",
+ " iteration_timeout_minutes = 60,\n",
+ " iterations = 10,\n",
+ " n_cross_validations = 2,\n",
+ " verbosity = logging.INFO,\n",
+ " X = X_train, \n",
+ " y = y_train,\n",
+ " path = project_folder)\n",
+ "\n",
+ "automl_sample_weight = AutoMLConfig(task = 'classification',\n",
+ " debug_log = 'automl_errors.log',\n",
+ " primary_metric = 'AUC_weighted',\n",
+ " iteration_timeout_minutes = 60,\n",
+ " iterations = 10,\n",
+ " n_cross_validations = 2,\n",
+ " verbosity = logging.INFO,\n",
+ " X = X_train, \n",
+ " y = y_train,\n",
+ " sample_weight = sample_weight,\n",
+ " path = project_folder)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Call the `submit` method on the experiment objects and pass the run configuration. Execution of local runs is synchronous. Depending on the data and the number of iterations this can run for a while.\n",
+ "In this example, we specify `show_output = True` to print currently running iterations to the console."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "local_run = experiment.submit(automl_classifier, show_output = True)\n",
+ "sample_weight_run = sample_weight_experiment.submit(automl_sample_weight, show_output = True)\n",
+ "\n",
+ "best_run, fitted_model = local_run.get_output()\n",
+ "best_run_sample_weight, fitted_model_sample_weight = sample_weight_run.get_output()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Test\n",
+ "\n",
+ "#### Load Test Data"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "digits = datasets.load_digits()\n",
+ "X_test = digits.data[:100, :]\n",
+ "y_test = digits.target[:100]\n",
+ "images = digits.images[:100]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Compare the Models\n",
+ "The prediction from the sample weight model is more likely to correctly predict 4's. However, it is also more likely to predict 4 for some images that are not labelled as 4."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Randomly select digits and test.\n",
+ "for index in range(0,len(y_test)):\n",
+ " predicted = fitted_model.predict(X_test[index:index + 1])[0]\n",
+ " predicted_sample_weight = fitted_model_sample_weight.predict(X_test[index:index + 1])[0]\n",
+ " label = y_test[index]\n",
+ " if predicted == 4 or predicted_sample_weight == 4 or label == 4:\n",
+ " title = \"Label value = %d Predicted value = %d Prediced with sample weight = %d\" % (label, predicted, predicted_sample_weight)\n",
+ " fig = plt.figure(1, figsize=(3,3))\n",
+ " ax1 = fig.add_axes((0,0,.8,.8))\n",
+ " ax1.set_title(title)\n",
+ " plt.imshow(images[index], cmap = plt.cm.gray_r, interpolation = 'nearest')\n",
+ " plt.show()"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "savitam"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.5"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.5"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/automated-machine-learning/sparse-data-train-test-split/auto-ml-sparse-data-train-test-split.ipynb b/how-to-use-azureml/automated-machine-learning/sparse-data-train-test-split/auto-ml-sparse-data-train-test-split.ipynb
index 0a08890f..24a1989f 100644
--- a/how-to-use-azureml/automated-machine-learning/sparse-data-train-test-split/auto-ml-sparse-data-train-test-split.ipynb
+++ b/how-to-use-azureml/automated-machine-learning/sparse-data-train-test-split/auto-ml-sparse-data-train-test-split.ipynb
@@ -1,403 +1,397 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Automated Machine Learning\n",
- "_**Train Test Split and Handling Sparse Data**_\n",
- "\n",
- "## Contents\n",
- "1. [Introduction](#Introduction)\n",
- "1. [Setup](#Setup)\n",
- "1. [Data](#Data)\n",
- "1. [Train](#Train)\n",
- "1. [Results](#Results)\n",
- "1. [Test](#Test)\n"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Introduction\n",
- "In this example we use the scikit-learn's [20newsgroup](http://scikit-learn.org/stable/modules/generated/sklearn.datasets.fetch_20newsgroups.html) to showcase how you can use AutoML for handling sparse data and how to specify custom cross validations splits.\n",
- "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
- "\n",
- "In this notebook you will learn how to:\n",
- "1. Create an `Experiment` in an existing `Workspace`.\n",
- "2. Configure AutoML using `AutoMLConfig`.\n",
- "4. Train the model.\n",
- "5. Explore the results.\n",
- "6. Test the best fitted model.\n",
- "\n",
- "In addition this notebook showcases the following features\n",
- "- Explicit train test splits \n",
- "- Handling **sparse data** in the input"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Setup\n",
- "\n",
- "As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import logging\n",
- "import os\n",
- "import random\n",
- "\n",
- "from matplotlib import pyplot as plt\n",
- "from matplotlib.pyplot import imshow\n",
- "import numpy as np\n",
- "import pandas as pd\n",
- "from sklearn import datasets\n",
- "\n",
- "import azureml.core\n",
- "from azureml.core.experiment import Experiment\n",
- "from azureml.core.workspace import Workspace\n",
- "from azureml.train.automl import AutoMLConfig\n",
- "from azureml.train.automl.run import AutoMLRun"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "ws = Workspace.from_config()\n",
- "\n",
- "# choose a name for the experiment\n",
- "experiment_name = 'automl-local-missing-data'\n",
- "# project folder\n",
- "project_folder = './sample_projects/automl-local-missing-data'\n",
- "\n",
- "experiment = Experiment(ws, experiment_name)\n",
- "\n",
- "output = {}\n",
- "output['SDK version'] = azureml.core.VERSION\n",
- "output['Subscription ID'] = ws.subscription_id\n",
- "output['Workspace'] = ws.name\n",
- "output['Resource Group'] = ws.resource_group\n",
- "output['Location'] = ws.location\n",
- "output['Project Directory'] = project_folder\n",
- "output['Experiment Name'] = experiment.name\n",
- "pd.set_option('display.max_colwidth', -1)\n",
- "pd.DataFrame(data=output, index=['']).T"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Opt-in diagnostics for better experience, quality, and security of future releases."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.telemetry import set_diagnostics_collection\n",
- "set_diagnostics_collection(send_diagnostics = True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Data"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from sklearn.datasets import fetch_20newsgroups\n",
- "from sklearn.feature_extraction.text import HashingVectorizer\n",
- "from sklearn.model_selection import train_test_split\n",
- "\n",
- "remove = ('headers', 'footers', 'quotes')\n",
- "categories = [\n",
- " 'alt.atheism',\n",
- " 'talk.religion.misc',\n",
- " 'comp.graphics',\n",
- " 'sci.space',\n",
- "]\n",
- "data_train = fetch_20newsgroups(subset = 'train', categories = categories,\n",
- " shuffle = True, random_state = 42,\n",
- " remove = remove)\n",
- "\n",
- "X_train, X_valid, y_train, y_valid = train_test_split(data_train.data, data_train.target, test_size = 0.33, random_state = 42)\n",
- "\n",
- "\n",
- "vectorizer = HashingVectorizer(stop_words = 'english', alternate_sign = False,\n",
- " n_features = 2**16)\n",
- "X_train = vectorizer.transform(X_train)\n",
- "X_valid = vectorizer.transform(X_valid)\n",
- "\n",
- "summary_df = pd.DataFrame(index = ['No of Samples', 'No of Features'])\n",
- "summary_df['Train Set'] = [X_train.shape[0], X_train.shape[1]]\n",
- "summary_df['Validation Set'] = [X_valid.shape[0], X_valid.shape[1]]\n",
- "summary_df"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Train\n",
- "\n",
- "Instantiate an `AutoMLConfig` object to specify the settings and data used to run the experiment.\n",
- "\n",
- "|Property|Description|\n",
- "|-|-|\n",
- "|**task**|classification or regression|\n",
- "|**primary_metric**|This is the metric that you want to optimize. Classification supports the following primary metrics:
accuracy
AUC_weighted
average_precision_score_weighted
norm_macro_recall
precision_score_weighted|\n",
- "|**iteration_timeout_minutes**|Time limit in minutes for each iteration.|\n",
- "|**iterations**|Number of iterations. In each iteration AutoML trains a specific pipeline with the data.|\n",
- "|**preprocess**|Setting this to *True* enables AutoML to perform preprocessing on the input to handle *missing data*, and to perform some common *feature extraction*.
**Note:** If input data is sparse, you cannot use *True*.|\n",
- "|**X**|(sparse) array-like, shape = [n_samples, n_features]|\n",
- "|**y**|(sparse) array-like, shape = [n_samples, ], [n_samples, n_classes]
Multi-class targets. An indicator matrix turns on multilabel classification. This should be an array of integers.|\n",
- "|**X_valid**|(sparse) array-like, shape = [n_samples, n_features] for the custom validation set.|\n",
- "|**y_valid**|(sparse) array-like, shape = [n_samples, ], [n_samples, n_classes]
Multi-class targets. An indicator matrix turns on multilabel classification for the custom validation set.|\n",
- "|**path**|Relative path to the project folder. AutoML stores configuration files for the experiment under this folder. You can specify a new empty folder.|"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "automl_config = AutoMLConfig(task = 'classification',\n",
- " debug_log = 'automl_errors.log',\n",
- " primary_metric = 'AUC_weighted',\n",
- " iteration_timeout_minutes = 60,\n",
- " iterations = 5,\n",
- " preprocess = False,\n",
- " verbosity = logging.INFO,\n",
- " X = X_train, \n",
- " y = y_train,\n",
- " X_valid = X_valid, \n",
- " y_valid = y_valid, \n",
- " path = project_folder)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Call the `submit` method on the experiment object and pass the run configuration. Execution of local runs is synchronous. Depending on the data and the number of iterations this can run for a while.\n",
- "In this example, we specify `show_output = True` to print currently running iterations to the console."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "local_run = experiment.submit(automl_config, show_output=True)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "local_run"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Results"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Widget for Monitoring Runs\n",
- "\n",
- "The widget will first report a \"loading\" status while running the first iteration. After completing the first iteration, an auto-updating graph and table will be shown. The widget will refresh once per minute, so you should see the graph update as child runs complete.\n",
- "\n",
- "**Note:** The widget displays a link at the bottom. Use this link to open a web interface to explore the individual run details."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.widgets import RunDetails\n",
- "RunDetails(local_run).show() "
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "\n",
- "#### Retrieve All Child Runs\n",
- "You can also use SDK methods to fetch all the child runs and see individual metrics that we log."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "children = list(local_run.get_children())\n",
- "metricslist = {}\n",
- "for run in children:\n",
- " properties = run.get_properties()\n",
- " metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)}\n",
- " metricslist[int(properties['iteration'])] = metrics\n",
- " \n",
- "rundata = pd.DataFrame(metricslist).sort_index(1)\n",
- "rundata"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Retrieve the Best Model\n",
- "\n",
- "Below we select the best pipeline from our iterations. The `get_output` method returns the best run and the fitted model. The Model includes the pipeline and any pre-processing. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "best_run, fitted_model = local_run.get_output()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Best Model Based on Any Other Metric\n",
- "Show the run and the model which has the smallest `accuracy` value:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# lookup_metric = \"accuracy\"\n",
- "# best_run, fitted_model = local_run.get_output(metric = lookup_metric)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Model from a Specific Iteration\n",
- "Show the run and the model from the third iteration:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# iteration = 3\n",
- "# best_run, fitted_model = local_run.get_output(iteration = iteration)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Test"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Load test data.\n",
- "from pandas_ml import ConfusionMatrix\n",
- "\n",
- "data_test = fetch_20newsgroups(subset = 'test', categories = categories,\n",
- " shuffle = True, random_state = 42,\n",
- " remove = remove)\n",
- "\n",
- "X_test = vectorizer.transform(data_test.data)\n",
- "y_test = data_test.target\n",
- "\n",
- "# Test our best pipeline.\n",
- "\n",
- "y_pred = fitted_model.predict(X_test)\n",
- "y_pred_strings = [data_test.target_names[i] for i in y_pred]\n",
- "y_test_strings = [data_test.target_names[i] for i in y_test]\n",
- "\n",
- "cm = ConfusionMatrix(y_test_strings, y_pred_strings)\n",
- "print(cm)\n",
- "cm.plot()"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "savitam"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Automated Machine Learning\n",
+ "_**Train Test Split and Handling Sparse Data**_\n",
+ "\n",
+ "## Contents\n",
+ "1. [Introduction](#Introduction)\n",
+ "1. [Setup](#Setup)\n",
+ "1. [Data](#Data)\n",
+ "1. [Train](#Train)\n",
+ "1. [Results](#Results)\n",
+ "1. [Test](#Test)\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Introduction\n",
+ "In this example we use the scikit-learn's [20newsgroup](http://scikit-learn.org/stable/modules/generated/sklearn.datasets.fetch_20newsgroups.html) to showcase how you can use AutoML for handling sparse data and how to specify custom cross validations splits.\n",
+ "Make sure you have executed the [configuration](../../../configuration.ipynb) before running this notebook.\n",
+ "\n",
+ "In this notebook you will learn how to:\n",
+ "1. Create an `Experiment` in an existing `Workspace`.\n",
+ "2. Configure AutoML using `AutoMLConfig`.\n",
+ "4. Train the model.\n",
+ "5. Explore the results.\n",
+ "6. Test the best fitted model.\n",
+ "\n",
+ "In addition this notebook showcases the following features\n",
+ "- Explicit train test splits \n",
+ "- Handling **sparse data** in the input"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Setup\n",
+ "\n",
+ "As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import logging\n",
+ "\n",
+ "import pandas as pd\n",
+ "\n",
+ "import azureml.core\n",
+ "from azureml.core.experiment import Experiment\n",
+ "from azureml.core.workspace import Workspace\n",
+ "from azureml.train.automl import AutoMLConfig"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "ws = Workspace.from_config()\n",
+ "\n",
+ "# choose a name for the experiment\n",
+ "experiment_name = 'automl-local-missing-data'\n",
+ "# project folder\n",
+ "project_folder = './sample_projects/automl-local-missing-data'\n",
+ "\n",
+ "experiment = Experiment(ws, experiment_name)\n",
+ "\n",
+ "output = {}\n",
+ "output['SDK version'] = azureml.core.VERSION\n",
+ "output['Subscription ID'] = ws.subscription_id\n",
+ "output['Workspace'] = ws.name\n",
+ "output['Resource Group'] = ws.resource_group\n",
+ "output['Location'] = ws.location\n",
+ "output['Project Directory'] = project_folder\n",
+ "output['Experiment Name'] = experiment.name\n",
+ "pd.set_option('display.max_colwidth', -1)\n",
+ "outputDf = pd.DataFrame(data = output, index = [''])\n",
+ "outputDf.T"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Opt-in diagnostics for better experience, quality, and security of future releases."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.telemetry import set_diagnostics_collection\n",
+ "set_diagnostics_collection(send_diagnostics = True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Data"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from sklearn.datasets import fetch_20newsgroups\n",
+ "from sklearn.feature_extraction.text import HashingVectorizer\n",
+ "from sklearn.model_selection import train_test_split\n",
+ "\n",
+ "remove = ('headers', 'footers', 'quotes')\n",
+ "categories = [\n",
+ " 'alt.atheism',\n",
+ " 'talk.religion.misc',\n",
+ " 'comp.graphics',\n",
+ " 'sci.space',\n",
+ "]\n",
+ "data_train = fetch_20newsgroups(subset = 'train', categories = categories,\n",
+ " shuffle = True, random_state = 42,\n",
+ " remove = remove)\n",
+ "\n",
+ "X_train, X_valid, y_train, y_valid = train_test_split(data_train.data, data_train.target, test_size = 0.33, random_state = 42)\n",
+ "\n",
+ "\n",
+ "vectorizer = HashingVectorizer(stop_words = 'english', alternate_sign = False,\n",
+ " n_features = 2**16)\n",
+ "X_train = vectorizer.transform(X_train)\n",
+ "X_valid = vectorizer.transform(X_valid)\n",
+ "\n",
+ "summary_df = pd.DataFrame(index = ['No of Samples', 'No of Features'])\n",
+ "summary_df['Train Set'] = [X_train.shape[0], X_train.shape[1]]\n",
+ "summary_df['Validation Set'] = [X_valid.shape[0], X_valid.shape[1]]\n",
+ "summary_df"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Train\n",
+ "\n",
+ "Instantiate an `AutoMLConfig` object to specify the settings and data used to run the experiment.\n",
+ "\n",
+ "|Property|Description|\n",
+ "|-|-|\n",
+ "|**task**|classification or regression|\n",
+ "|**primary_metric**|This is the metric that you want to optimize. Classification supports the following primary metrics:
accuracy
AUC_weighted
average_precision_score_weighted
norm_macro_recall
precision_score_weighted|\n",
+ "|**iteration_timeout_minutes**|Time limit in minutes for each iteration.|\n",
+ "|**iterations**|Number of iterations. In each iteration AutoML trains a specific pipeline with the data.|\n",
+ "|**preprocess**|Setting this to *True* enables AutoML to perform preprocessing on the input to handle *missing data*, and to perform some common *feature extraction*.
**Note:** If input data is sparse, you cannot use *True*.|\n",
+ "|**X**|(sparse) array-like, shape = [n_samples, n_features]|\n",
+ "|**y**|(sparse) array-like, shape = [n_samples, ], [n_samples, n_classes]
Multi-class targets. An indicator matrix turns on multilabel classification. This should be an array of integers.|\n",
+ "|**X_valid**|(sparse) array-like, shape = [n_samples, n_features] for the custom validation set.|\n",
+ "|**y_valid**|(sparse) array-like, shape = [n_samples, ], [n_samples, n_classes]
Multi-class targets. An indicator matrix turns on multilabel classification for the custom validation set.|\n",
+ "|**path**|Relative path to the project folder. AutoML stores configuration files for the experiment under this folder. You can specify a new empty folder.|"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "automl_config = AutoMLConfig(task = 'classification',\n",
+ " debug_log = 'automl_errors.log',\n",
+ " primary_metric = 'AUC_weighted',\n",
+ " iteration_timeout_minutes = 60,\n",
+ " iterations = 5,\n",
+ " preprocess = False,\n",
+ " verbosity = logging.INFO,\n",
+ " X = X_train, \n",
+ " y = y_train,\n",
+ " X_valid = X_valid, \n",
+ " y_valid = y_valid, \n",
+ " path = project_folder)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Call the `submit` method on the experiment object and pass the run configuration. Execution of local runs is synchronous. Depending on the data and the number of iterations this can run for a while.\n",
+ "In this example, we specify `show_output = True` to print currently running iterations to the console."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "local_run = experiment.submit(automl_config, show_output=True)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "local_run"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Results"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Widget for Monitoring Runs\n",
+ "\n",
+ "The widget will first report a \"loading\" status while running the first iteration. After completing the first iteration, an auto-updating graph and table will be shown. The widget will refresh once per minute, so you should see the graph update as child runs complete.\n",
+ "\n",
+ "**Note:** The widget displays a link at the bottom. Use this link to open a web interface to explore the individual run details."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.widgets import RunDetails\n",
+ "RunDetails(local_run).show() "
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "\n",
+ "#### Retrieve All Child Runs\n",
+ "You can also use SDK methods to fetch all the child runs and see individual metrics that we log."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "children = list(local_run.get_children())\n",
+ "metricslist = {}\n",
+ "for run in children:\n",
+ " properties = run.get_properties()\n",
+ " metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)}\n",
+ " metricslist[int(properties['iteration'])] = metrics\n",
+ " \n",
+ "rundata = pd.DataFrame(metricslist).sort_index(1)\n",
+ "rundata"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Retrieve the Best Model\n",
+ "\n",
+ "Below we select the best pipeline from our iterations. The `get_output` method returns the best run and the fitted model. The Model includes the pipeline and any pre-processing. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "best_run, fitted_model = local_run.get_output()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Best Model Based on Any Other Metric\n",
+ "Show the run and the model which has the smallest `accuracy` value:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# lookup_metric = \"accuracy\"\n",
+ "# best_run, fitted_model = local_run.get_output(metric = lookup_metric)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Model from a Specific Iteration\n",
+ "Show the run and the model from the third iteration:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# iteration = 3\n",
+ "# best_run, fitted_model = local_run.get_output(iteration = iteration)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Test"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Load test data.\n",
+ "from pandas_ml import ConfusionMatrix\n",
+ "\n",
+ "data_test = fetch_20newsgroups(subset = 'test', categories = categories,\n",
+ " shuffle = True, random_state = 42,\n",
+ " remove = remove)\n",
+ "\n",
+ "X_test = vectorizer.transform(data_test.data)\n",
+ "y_test = data_test.target\n",
+ "\n",
+ "# Test our best pipeline.\n",
+ "\n",
+ "y_pred = fitted_model.predict(X_test)\n",
+ "y_pred_strings = [data_test.target_names[i] for i in y_pred]\n",
+ "y_test_strings = [data_test.target_names[i] for i in y_test]\n",
+ "\n",
+ "cm = ConfusionMatrix(y_test_strings, y_pred_strings)\n",
+ "print(cm)\n",
+ "cm.plot()"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "savitam"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.6"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/deployment/enable-app-insights-in-production-service/enable-app-insights-in-production-service.ipynb b/how-to-use-azureml/deployment/enable-app-insights-in-production-service/enable-app-insights-in-production-service.ipynb
index 25a82f1d..fb35d654 100644
--- a/how-to-use-azureml/deployment/enable-app-insights-in-production-service/enable-app-insights-in-production-service.ipynb
+++ b/how-to-use-azureml/deployment/enable-app-insights-in-production-service/enable-app-insights-in-production-service.ipynb
@@ -1,495 +1,495 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Enabling App Insights for Services in Production\n",
- "With this notebook, you can learn how to enable App Insights for standard service monitoring, plus, we provide examples for doing custom logging within a scoring files in a model. \n",
- "\n",
- "\n",
- "## What does Application Insights monitor?\n",
- "It monitors request rates, response times, failure rates, etc. For more information visit [App Insights docs.](https://docs.microsoft.com/en-us/azure/application-insights/app-insights-overview)\n",
- "\n",
- "\n",
- "## What is different compared to standard production deployment process?\n",
- "If you want to enable generic App Insights for a service run:\n",
- "```python\n",
- "aks_service= Webservice(ws, \"aks-w-dc2\")\n",
- "aks_service.update(enable_app_insights=True)```\n",
- "Where \"aks-w-dc2\" is your service name. You can also do this from the Azure Portal under your Workspace--> deployments--> Select deployment--> Edit--> Advanced Settings--> Select \"Enable AppInsights diagnostics\"\n",
- "\n",
- "If you want to log custom traces, you will follow the standard deplyment process for AKS and you will:\n",
- "1. Update scoring file.\n",
- "2. Update aks configuration.\n",
- "3. Build new image and deploy it. "
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## 1. Import your dependencies"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core import Workspace, Run\n",
- "from azureml.core.compute import AksCompute, ComputeTarget\n",
- "from azureml.core.webservice import Webservice, AksWebservice\n",
- "from azureml.core.image import Image\n",
- "from azureml.core.model import Model\n",
- "\n",
- "import azureml.core\n",
- "print(azureml.core.VERSION)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## 2. Set up your configuration and create a workspace\n"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "ws = Workspace.from_config()\n",
- "print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep = '\\n')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## 3. Register Model\n",
- "Register an existing trained model, add descirption and tags."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "#Register the model\n",
- "from azureml.core.model import Model\n",
- "model = Model.register(model_path = \"sklearn_regression_model.pkl\", # this points to a local file\n",
- " model_name = \"sklearn_regression_model.pkl\", # this is the name the model is registered as\n",
- " tags = {'area': \"diabetes\", 'type': \"regression\"},\n",
- " description = \"Ridge regression model to predict diabetes\",\n",
- " workspace = ws)\n",
- "\n",
- "print(model.name, model.description, model.version)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## 4. *Update your scoring file with custom print statements*\n",
- "Here is an example:\n",
- "### a. In your init function add:\n",
- "```python\n",
- "print (\"model initialized\" + time.strftime(\"%H:%M:%S\"))```\n",
- "\n",
- "### b. In your run function add:\n",
- "```python\n",
- "print (\"Prediction created\" + time.strftime(\"%H:%M:%S\"))```"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%writefile score.py\n",
- "import pickle\n",
- "import json\n",
- "import numpy \n",
- "from sklearn.externals import joblib\n",
- "from sklearn.linear_model import Ridge\n",
- "from azureml.core.model import Model\n",
- "import time\n",
- "\n",
- "def init():\n",
- " global model\n",
- " #Print statement for appinsights custom traces:\n",
- " print (\"model initialized\" + time.strftime(\"%H:%M:%S\"))\n",
- " \n",
- " # note here \"sklearn_regression_model.pkl\" is the name of the model registered under the workspace\n",
- " # this call should return the path to the model.pkl file on the local disk.\n",
- " model_path = Model.get_model_path(model_name = 'sklearn_regression_model.pkl')\n",
- " \n",
- " # deserialize the model file back into a sklearn model\n",
- " model = joblib.load(model_path)\n",
- " \n",
- "\n",
- "# note you can pass in multiple rows for scoring\n",
- "def run(raw_data):\n",
- " try:\n",
- " data = json.loads(raw_data)['data']\n",
- " data = numpy.array(data)\n",
- " result = model.predict(data)\n",
- " print (\"Prediction created\" + time.strftime(\"%H:%M:%S\"))\n",
- " # you can return any datatype as long as it is JSON-serializable\n",
- " return result.tolist()\n",
- " except Exception as e:\n",
- " error = str(e)\n",
- " print (error + time.strftime(\"%H:%M:%S\"))\n",
- " return error"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## 5. *Create myenv.yml file*"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.conda_dependencies import CondaDependencies \n",
- "\n",
- "myenv = CondaDependencies.create(conda_packages=['numpy','scikit-learn'])\n",
- "\n",
- "with open(\"myenv.yml\",\"w\") as f:\n",
- " f.write(myenv.serialize_to_string())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## 6. Create your new Image"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.image import ContainerImage\n",
- "\n",
- "image_config = ContainerImage.image_configuration(execution_script = \"score.py\",\n",
- " runtime = \"python\",\n",
- " conda_file = \"myenv.yml\",\n",
- " description = \"Image with ridge regression model\",\n",
- " tags = {'area': \"diabetes\", 'type': \"regression\"}\n",
- " )\n",
- "\n",
- "image = ContainerImage.create(name = \"myimage1\",\n",
- " # this is the model object\n",
- " models = [model],\n",
- " image_config = image_config,\n",
- " workspace = ws)\n",
- "\n",
- "image.wait_for_creation(show_output = True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Deploy to ACI (Optional)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.webservice import AciWebservice\n",
- "\n",
- "aciconfig = AciWebservice.deploy_configuration(cpu_cores = 1, \n",
- " memory_gb = 1, \n",
- " tags = {'area': \"diabetes\", 'type': \"regression\"}, \n",
- " description = 'Predict diabetes using regression model',\n",
- " enable_app_insights = True)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.webservice import Webservice\n",
- "\n",
- "aci_service_name = 'my-aci-service-4'\n",
- "print(aci_service_name)\n",
- "aci_service = Webservice.deploy_from_image(deployment_config = aciconfig,\n",
- " image = image,\n",
- " name = aci_service_name,\n",
- " workspace = ws)\n",
- "aci_service.wait_for_deployment(True)\n",
- "print(aci_service.state)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%time\n",
- "import json\n",
- "\n",
- "test_sample = json.dumps({'data': [\n",
- " [1,28,13,45,54,6,57,8,8,10], \n",
- " [101,9,8,37,6,45,4,3,2,41]\n",
- "]})\n",
- "test_sample = bytes(test_sample,encoding='utf8')"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "if aci_service.state == \"Healthy\":\n",
- " prediction = aci_service.run(input_data=test_sample)\n",
- " print(prediction)\n",
- "else:\n",
- " raise ValueError(\"Service deployment isn't healthy, can't call the service\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## 7. Deploy to AKS service"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create AKS compute if you haven't done so."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Use the default configuration (can also provide parameters to customize)\n",
- "prov_config = AksCompute.provisioning_configuration()\n",
- "\n",
- "aks_name = 'my-aks-test3' \n",
- "# Create the cluster\n",
- "aks_target = ComputeTarget.create(workspace = ws, \n",
- " name = aks_name, \n",
- " provisioning_configuration = prov_config)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%time\n",
- "aks_target.wait_for_completion(show_output = True)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "print(aks_target.provisioning_state)\n",
- "print(aks_target.provisioning_errors)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "If you already have a cluster you can attach the service to it:"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "```python \n",
- "%%time\n",
- "resource_id = '/subscriptions//resourcegroups//providers/Microsoft.ContainerService/managedClusters/'\n",
- "create_name= 'myaks4'\n",
- "attach_config = AksCompute.attach_configuration(resource_id=resource_id)\n",
- "aks_target = ComputeTarget.attach(workspace = ws, \n",
- " name = create_name, \n",
- " attach_configuration=attach_config)\n",
- "## Wait for the operation to complete\n",
- "aks_target.wait_for_provisioning(True)```"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### a. *Activate App Insights through updating AKS Webservice configuration*\n",
- "In order to enable App Insights in your service you will need to update your AKS configuration file:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "#Set the web service configuration\n",
- "aks_config = AksWebservice.deploy_configuration(enable_app_insights=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### b. Deploy your service"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "if aks_target.provisioning_state== \"Succeeded\": \n",
- " aks_service_name ='aks-w-dc5'\n",
- " aks_service = Webservice.deploy_from_image(workspace = ws, \n",
- " name = aks_service_name,\n",
- " image = image,\n",
- " deployment_config = aks_config,\n",
- " deployment_target = aks_target\n",
- " )\n",
- " aks_service.wait_for_deployment(show_output = True)\n",
- " print(aks_service.state)\n",
- "else:\n",
- " raise ValueError(\"AKS provisioning failed.\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## 8. Test your service "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%time\n",
- "import json\n",
- "\n",
- "test_sample = json.dumps({'data': [\n",
- " [1,28,13,45,54,6,57,8,8,10], \n",
- " [101,9,8,37,6,45,4,3,2,41]\n",
- "]})\n",
- "test_sample = bytes(test_sample,encoding='utf8')\n",
- "\n",
- "if aks_service.state == \"Healthy\":\n",
- " prediction = aks_service.run(input_data=test_sample)\n",
- " print(prediction)\n",
- "else:\n",
- " raise ValueError(\"Service deployment isn't healthy, can't call the service\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## 9. See your service telemetry in App Insights\n",
- "1. Go to the [Azure Portal](https://portal.azure.com/)\n",
- "2. All resources--> Select the subscription/resource group where you created your Workspace--> Select the App Insights type\n",
- "3. Click on the AppInsights resource. You'll see a highlevel dashboard with information on Requests, Server response time and availability.\n",
- "4. Click on the top banner \"Analytics\"\n",
- "5. In the \"Schema\" section select \"traces\" and run your query.\n",
- "6. Voila! All your custom traces should be there."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Disable App Insights"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "aks_service.update(enable_app_insights=False)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Clean up"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%time\n",
- "aks_service.delete()\n",
- "aci_service.delete()\n",
- "image.delete()\n",
- "model.delete()"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "marthalc"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Enabling App Insights for Services in Production\n",
+ "With this notebook, you can learn how to enable App Insights for standard service monitoring, plus, we provide examples for doing custom logging within a scoring files in a model. \n",
+ "\n",
+ "\n",
+ "## What does Application Insights monitor?\n",
+ "It monitors request rates, response times, failure rates, etc. For more information visit [App Insights docs.](https://docs.microsoft.com/en-us/azure/application-insights/app-insights-overview)\n",
+ "\n",
+ "\n",
+ "## What is different compared to standard production deployment process?\n",
+ "If you want to enable generic App Insights for a service run:\n",
+ "```python\n",
+ "aks_service= Webservice(ws, \"aks-w-dc2\")\n",
+ "aks_service.update(enable_app_insights=True)```\n",
+ "Where \"aks-w-dc2\" is your service name. You can also do this from the Azure Portal under your Workspace--> deployments--> Select deployment--> Edit--> Advanced Settings--> Select \"Enable AppInsights diagnostics\"\n",
+ "\n",
+ "If you want to log custom traces, you will follow the standard deplyment process for AKS and you will:\n",
+ "1. Update scoring file.\n",
+ "2. Update aks configuration.\n",
+ "3. Build new image and deploy it. "
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## 1. Import your dependencies"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core import Workspace, Run\n",
+ "from azureml.core.compute import AksCompute, ComputeTarget\n",
+ "from azureml.core.webservice import Webservice, AksWebservice\n",
+ "from azureml.core.image import Image\n",
+ "from azureml.core.model import Model\n",
+ "\n",
+ "import azureml.core\n",
+ "print(azureml.core.VERSION)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## 2. Set up your configuration and create a workspace\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "ws = Workspace.from_config()\n",
+ "print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep = '\\n')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## 3. Register Model\n",
+ "Register an existing trained model, add descirption and tags."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "#Register the model\n",
+ "from azureml.core.model import Model\n",
+ "model = Model.register(model_path = \"sklearn_regression_model.pkl\", # this points to a local file\n",
+ " model_name = \"sklearn_regression_model.pkl\", # this is the name the model is registered as\n",
+ " tags = {'area': \"diabetes\", 'type': \"regression\"},\n",
+ " description = \"Ridge regression model to predict diabetes\",\n",
+ " workspace = ws)\n",
+ "\n",
+ "print(model.name, model.description, model.version)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## 4. *Update your scoring file with custom print statements*\n",
+ "Here is an example:\n",
+ "### a. In your init function add:\n",
+ "```python\n",
+ "print (\"model initialized\" + time.strftime(\"%H:%M:%S\"))```\n",
+ "\n",
+ "### b. In your run function add:\n",
+ "```python\n",
+ "print (\"Prediction created\" + time.strftime(\"%H:%M:%S\"))```"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%writefile score.py\n",
+ "import pickle\n",
+ "import json\n",
+ "import numpy \n",
+ "from sklearn.externals import joblib\n",
+ "from sklearn.linear_model import Ridge\n",
+ "from azureml.core.model import Model\n",
+ "import time\n",
+ "\n",
+ "def init():\n",
+ " global model\n",
+ " #Print statement for appinsights custom traces:\n",
+ " print (\"model initialized\" + time.strftime(\"%H:%M:%S\"))\n",
+ " \n",
+ " # note here \"sklearn_regression_model.pkl\" is the name of the model registered under the workspace\n",
+ " # this call should return the path to the model.pkl file on the local disk.\n",
+ " model_path = Model.get_model_path(model_name = 'sklearn_regression_model.pkl')\n",
+ " \n",
+ " # deserialize the model file back into a sklearn model\n",
+ " model = joblib.load(model_path)\n",
+ " \n",
+ "\n",
+ "# note you can pass in multiple rows for scoring\n",
+ "def run(raw_data):\n",
+ " try:\n",
+ " data = json.loads(raw_data)['data']\n",
+ " data = numpy.array(data)\n",
+ " result = model.predict(data)\n",
+ " print (\"Prediction created\" + time.strftime(\"%H:%M:%S\"))\n",
+ " # you can return any datatype as long as it is JSON-serializable\n",
+ " return result.tolist()\n",
+ " except Exception as e:\n",
+ " error = str(e)\n",
+ " print (error + time.strftime(\"%H:%M:%S\"))\n",
+ " return error"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## 5. *Create myenv.yml file*"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.conda_dependencies import CondaDependencies \n",
+ "\n",
+ "myenv = CondaDependencies.create(conda_packages=['numpy','scikit-learn'])\n",
+ "\n",
+ "with open(\"myenv.yml\",\"w\") as f:\n",
+ " f.write(myenv.serialize_to_string())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## 6. Create your new Image"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.image import ContainerImage\n",
+ "\n",
+ "image_config = ContainerImage.image_configuration(execution_script = \"score.py\",\n",
+ " runtime = \"python\",\n",
+ " conda_file = \"myenv.yml\",\n",
+ " description = \"Image with ridge regression model\",\n",
+ " tags = {'area': \"diabetes\", 'type': \"regression\"}\n",
+ " )\n",
+ "\n",
+ "image = ContainerImage.create(name = \"myimage1\",\n",
+ " # this is the model object\n",
+ " models = [model],\n",
+ " image_config = image_config,\n",
+ " workspace = ws)\n",
+ "\n",
+ "image.wait_for_creation(show_output = True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Deploy to ACI (Optional)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.webservice import AciWebservice\n",
+ "\n",
+ "aciconfig = AciWebservice.deploy_configuration(cpu_cores = 1, \n",
+ " memory_gb = 1, \n",
+ " tags = {'area': \"diabetes\", 'type': \"regression\"}, \n",
+ " description = 'Predict diabetes using regression model',\n",
+ " enable_app_insights = True)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.webservice import Webservice\n",
+ "\n",
+ "aci_service_name = 'my-aci-service-4'\n",
+ "print(aci_service_name)\n",
+ "aci_service = Webservice.deploy_from_image(deployment_config = aciconfig,\n",
+ " image = image,\n",
+ " name = aci_service_name,\n",
+ " workspace = ws)\n",
+ "aci_service.wait_for_deployment(True)\n",
+ "print(aci_service.state)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%time\n",
+ "import json\n",
+ "\n",
+ "test_sample = json.dumps({'data': [\n",
+ " [1,28,13,45,54,6,57,8,8,10], \n",
+ " [101,9,8,37,6,45,4,3,2,41]\n",
+ "]})\n",
+ "test_sample = bytes(test_sample,encoding='utf8')"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "if aci_service.state == \"Healthy\":\n",
+ " prediction = aci_service.run(input_data=test_sample)\n",
+ " print(prediction)\n",
+ "else:\n",
+ " raise ValueError(\"Service deployment isn't healthy, can't call the service\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## 7. Deploy to AKS service"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create AKS compute if you haven't done so."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Use the default configuration (can also provide parameters to customize)\n",
+ "prov_config = AksCompute.provisioning_configuration()\n",
+ "\n",
+ "aks_name = 'my-aks-test3' \n",
+ "# Create the cluster\n",
+ "aks_target = ComputeTarget.create(workspace = ws, \n",
+ " name = aks_name, \n",
+ " provisioning_configuration = prov_config)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%time\n",
+ "aks_target.wait_for_completion(show_output = True)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "print(aks_target.provisioning_state)\n",
+ "print(aks_target.provisioning_errors)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "If you already have a cluster you can attach the service to it:"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "```python \n",
+ "%%time\n",
+ "resource_id = '/subscriptions//resourcegroups//providers/Microsoft.ContainerService/managedClusters/'\n",
+ "create_name= 'myaks4'\n",
+ "attach_config = AksCompute.attach_configuration(resource_id=resource_id)\n",
+ "aks_target = ComputeTarget.attach(workspace = ws, \n",
+ " name = create_name, \n",
+ " attach_configuration=attach_config)\n",
+ "## Wait for the operation to complete\n",
+ "aks_target.wait_for_provisioning(True)```"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### a. *Activate App Insights through updating AKS Webservice configuration*\n",
+ "In order to enable App Insights in your service you will need to update your AKS configuration file:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "#Set the web service configuration\n",
+ "aks_config = AksWebservice.deploy_configuration(enable_app_insights=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### b. Deploy your service"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "if aks_target.provisioning_state== \"Succeeded\": \n",
+ " aks_service_name ='aks-w-dc5'\n",
+ " aks_service = Webservice.deploy_from_image(workspace = ws, \n",
+ " name = aks_service_name,\n",
+ " image = image,\n",
+ " deployment_config = aks_config,\n",
+ " deployment_target = aks_target\n",
+ " )\n",
+ " aks_service.wait_for_deployment(show_output = True)\n",
+ " print(aks_service.state)\n",
+ "else:\n",
+ " raise ValueError(\"AKS provisioning failed.\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## 8. Test your service "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%time\n",
+ "import json\n",
+ "\n",
+ "test_sample = json.dumps({'data': [\n",
+ " [1,28,13,45,54,6,57,8,8,10], \n",
+ " [101,9,8,37,6,45,4,3,2,41]\n",
+ "]})\n",
+ "test_sample = bytes(test_sample,encoding='utf8')\n",
+ "\n",
+ "if aks_service.state == \"Healthy\":\n",
+ " prediction = aks_service.run(input_data=test_sample)\n",
+ " print(prediction)\n",
+ "else:\n",
+ " raise ValueError(\"Service deployment isn't healthy, can't call the service\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## 9. See your service telemetry in App Insights\n",
+ "1. Go to the [Azure Portal](https://portal.azure.com/)\n",
+ "2. All resources--> Select the subscription/resource group where you created your Workspace--> Select the App Insights type\n",
+ "3. Click on the AppInsights resource. You'll see a highlevel dashboard with information on Requests, Server response time and availability.\n",
+ "4. Click on the top banner \"Analytics\"\n",
+ "5. In the \"Schema\" section select \"traces\" and run your query.\n",
+ "6. Voila! All your custom traces should be there."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Disable App Insights"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "aks_service.update(enable_app_insights=False)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Clean up"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%time\n",
+ "aks_service.delete()\n",
+ "aci_service.delete()\n",
+ "image.delete()\n",
+ "model.delete()"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python [default]",
- "language": "python",
- "name": "python3"
+ "metadata": {
+ "authors": [
+ {
+ "name": "marthalc"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.5"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.5"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/deployment/enable-data-collection-for-models-in-aks/enable-data-collection-for-models-in-aks.ipynb b/how-to-use-azureml/deployment/enable-data-collection-for-models-in-aks/enable-data-collection-for-models-in-aks.ipynb
index d994e7ac..3a076523 100644
--- a/how-to-use-azureml/deployment/enable-data-collection-for-models-in-aks/enable-data-collection-for-models-in-aks.ipynb
+++ b/how-to-use-azureml/deployment/enable-data-collection-for-models-in-aks/enable-data-collection-for-models-in-aks.ipynb
@@ -1,477 +1,477 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Enabling Data Collection for Models in Production\n",
- "With this notebook, you can learn how to collect input model data from your Azure Machine Learning service in an Azure Blob storage. Once enabled, this data collected gives you the opportunity:\n",
- "\n",
- "* Monitor data drifts as production data enters your model\n",
- "* Make better decisions on when to retrain or optimize your model\n",
- "* Retrain your model with the data collected\n",
- "\n",
- "## What data is collected?\n",
- "* Model input data (voice, images, and video are not supported) from services deployed in Azure Kubernetes Cluster (AKS)\n",
- "* Model predictions using production input data.\n",
- "\n",
- "**Note:** pre-aggregation or pre-calculations on this data are done by user and not included in this version of the product.\n",
- "\n",
- "## What is different compared to standard production deployment process?\n",
- "1. Update scoring file.\n",
- "2. Update yml file with new dependency.\n",
- "3. Update aks configuration.\n",
- "4. Build new image and deploy it. "
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## 1. Import your dependencies"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core import Workspace, Run\n",
- "from azureml.core.compute import AksCompute, ComputeTarget\n",
- "from azureml.core.webservice import Webservice, AksWebservice\n",
- "from azureml.core.image import Image\n",
- "from azureml.core.model import Model\n",
- "\n",
- "import azureml.core\n",
- "print(azureml.core.VERSION)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## 2. Set up your configuration and create a workspace\n",
- "Follow Notebook 00 instructions to do this.\n"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "ws = Workspace.from_config()\n",
- "print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep = '\\n')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## 3. Register Model\n",
- "Register an existing trained model, add descirption and tags."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "#Register the model\n",
- "from azureml.core.model import Model\n",
- "model = Model.register(model_path = \"sklearn_regression_model.pkl\", # this points to a local file\n",
- " model_name = \"sklearn_regression_model.pkl\", # this is the name the model is registered as\n",
- " tags = {'area': \"diabetes\", 'type': \"regression\"},\n",
- " description = \"Ridge regression model to predict diabetes\",\n",
- " workspace = ws)\n",
- "\n",
- "print(model.name, model.description, model.version)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## 4. *Update your scoring file with Data Collection*\n",
- "The file below, compared to the file used in notebook 11, has the following changes:\n",
- "### a. Import the module\n",
- "```python \n",
- "from azureml.monitoring import ModelDataCollector```\n",
- "### b. In your init function add:\n",
- "```python \n",
- "global inputs_dc, prediction_d\n",
- "inputs_dc = ModelDataCollector(\"best_model\", identifier=\"inputs\", feature_names=[\"feat1\", \"feat2\", \"feat3\", \"feat4\", \"feat5\", \"Feat6\"])\n",
- "prediction_dc = ModelDataCollector(\"best_model\", identifier=\"predictions\", feature_names=[\"prediction1\", \"prediction2\"])```\n",
- " \n",
- "* Identifier: Identifier is later used for building the folder structure in your Blob, it can be used to divide \"raw\" data versus \"processed\".\n",
- "* CorrelationId: is an optional parameter, you do not need to set it up if your model doesn't require it. Having a correlationId in place does help you for easier mapping with other data. (Examples include: LoanNumber, CustomerId, etc.)\n",
- "* Feature Names: These need to be set up in the order of your features in order for them to have column names when the .csv is created.\n",
- "\n",
- "### c. In your run function add:\n",
- "```python\n",
- "inputs_dc.collect(data)\n",
- "prediction_dc.collect(result)```"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%writefile score.py\n",
- "import pickle\n",
- "import json\n",
- "import numpy \n",
- "from sklearn.externals import joblib\n",
- "from sklearn.linear_model import Ridge\n",
- "from azureml.core.model import Model\n",
- "from azureml.monitoring import ModelDataCollector\n",
- "import time\n",
- "\n",
- "def init():\n",
- " global model\n",
- " print (\"model initialized\" + time.strftime(\"%H:%M:%S\"))\n",
- " # note here \"sklearn_regression_model.pkl\" is the name of the model registered under the workspace\n",
- " # this call should return the path to the model.pkl file on the local disk.\n",
- " model_path = Model.get_model_path(model_name = 'sklearn_regression_model.pkl')\n",
- " # deserialize the model file back into a sklearn model\n",
- " model = joblib.load(model_path)\n",
- " global inputs_dc, prediction_dc\n",
- " # this setup will help us save our inputs under the \"inputs\" path in our Azure Blob\n",
- " inputs_dc = ModelDataCollector(model_name=\"sklearn_regression_model\", identifier=\"inputs\", feature_names=[\"feat1\", \"feat2\"]) \n",
- " # this setup will help us save our ipredictions under the \"predictions\" path in our Azure Blob\n",
- " prediction_dc = ModelDataCollector(\"sklearn_regression_model\", identifier=\"predictions\", feature_names=[\"prediction1\", \"prediction2\"]) \n",
- " \n",
- "# note you can pass in multiple rows for scoring\n",
- "def run(raw_data):\n",
- " global inputs_dc, prediction_dc\n",
- " try:\n",
- " data = json.loads(raw_data)['data']\n",
- " data = numpy.array(data)\n",
- " result = model.predict(data)\n",
- " print (\"saving input data\" + time.strftime(\"%H:%M:%S\"))\n",
- " inputs_dc.collect(data) #this call is saving our input data into our blob\n",
- " prediction_dc.collect(result)#this call is saving our prediction data into our blob\n",
- " print (\"saving prediction data\" + time.strftime(\"%H:%M:%S\"))\n",
- " # you can return any data type as long as it is JSON-serializable\n",
- " return result.tolist()\n",
- " except Exception as e:\n",
- " error = str(e)\n",
- " print (error + time.strftime(\"%H:%M:%S\"))\n",
- " return error"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## 5. *Update your myenv.yml file with the required module*"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.conda_dependencies import CondaDependencies \n",
- "\n",
- "myenv = CondaDependencies.create(conda_packages=['numpy','scikit-learn'])\n",
- "myenv.add_pip_package(\"azureml-monitoring\")\n",
- "\n",
- "with open(\"myenv.yml\",\"w\") as f:\n",
- " f.write(myenv.serialize_to_string())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## 6. Create your new Image"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.image import ContainerImage\n",
- "\n",
- "image_config = ContainerImage.image_configuration(execution_script = \"score.py\",\n",
- " runtime = \"python\",\n",
- " conda_file = \"myenv.yml\",\n",
- " description = \"Image with ridge regression model\",\n",
- " tags = {'area': \"diabetes\", 'type': \"regression\"}\n",
- " )\n",
- "\n",
- "image = ContainerImage.create(name = \"myimage1\",\n",
- " # this is the model object\n",
- " models = [model],\n",
- " image_config = image_config,\n",
- " workspace = ws)\n",
- "\n",
- "image.wait_for_creation(show_output = True)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "print(model.name, model.description, model.version)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## 7. Deploy to AKS service"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create AKS compute if you haven't done so."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Use the default configuration (can also provide parameters to customize)\n",
- "prov_config = AksCompute.provisioning_configuration()\n",
- "\n",
- "aks_name = 'my-aks-test1' \n",
- "# Create the cluster\n",
- "aks_target = ComputeTarget.create(workspace = ws, \n",
- " name = aks_name, \n",
- " provisioning_configuration = prov_config)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%time\n",
- "aks_target.wait_for_completion(show_output = True)\n",
- "print(aks_target.provisioning_state)\n",
- "print(aks_target.provisioning_errors)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "If you already have a cluster you can attach the service to it:"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {
- "scrolled": true
- },
- "source": [
- "```python \n",
- " %%time\n",
- " resource_id = '/subscriptions//resourcegroups//providers/Microsoft.ContainerService/managedClusters/'\n",
- " create_name= 'myaks4'\n",
- " attach_config = AksCompute.attach_configuration(resource_id=resource_id)\n",
- " aks_target = ComputeTarget.attach(workspace = ws, \n",
- " name = create_name, \n",
- " attach_configuration=attach_config)\n",
- " ## Wait for the operation to complete\n",
- " aks_target.wait_for_provisioning(True)```"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### a. *Activate Data Collection and App Insights through updating AKS Webservice configuration*\n",
- "In order to enable Data Collection and App Insights in your service you will need to update your AKS configuration file:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "#Set the web service configuration\n",
- "aks_config = AksWebservice.deploy_configuration(collect_model_data=True, enable_app_insights=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### b. Deploy your service"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "if aks_target.provisioning_state== \"Succeeded\": \n",
- " aks_service_name ='aks-w-dc0'\n",
- " aks_service = Webservice.deploy_from_image(workspace = ws, \n",
- " name = aks_service_name,\n",
- " image = image,\n",
- " deployment_config = aks_config,\n",
- " deployment_target = aks_target\n",
- " )\n",
- " aks_service.wait_for_deployment(show_output = True)\n",
- " print(aks_service.state)\n",
- "else: \n",
- " raise ValueError(\"aks provisioning failed, can't deploy service\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## 8. Test your service and send some data\n",
- "**Note**: It will take around 15 mins for your data to appear in your blob.\n",
- "The data will appear in your Azure Blob following this format:\n",
- "\n",
- "/modeldata/subscriptionid/resourcegroupname/workspacename/webservicename/modelname/modelversion/identifier/year/month/day/data.csv "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%time\n",
- "import json\n",
- "\n",
- "test_sample = json.dumps({'data': [\n",
- " [1,2,3,4,54,6,7,8,88,10], \n",
- " [10,9,8,37,36,45,4,33,2,1]\n",
- "]})\n",
- "test_sample = bytes(test_sample,encoding = 'utf8')\n",
- "\n",
- "if aks_service.state == \"Healthy\":\n",
- " prediction = aks_service.run(input_data=test_sample)\n",
- " print(prediction)\n",
- "else:\n",
- " raise ValueError(\"Service deployment isn't healthy, can't call the service\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## 9. Validate you data and analyze it\n",
- "You can look into your data following this path format in your Azure Blob (it takes up to 15 minutes for the data to appear):\n",
- "\n",
- "/modeldata/**subscriptionid>**/**resourcegroupname>**/**workspacename>**/**webservicename>**/**modelname>**/**modelversion>>**/**identifier>**/*year/month/day*/data.csv \n",
- "\n",
- "For doing further analysis you have multiple options:"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### a. Create DataBricks cluter and connect it to your blob\n",
- "https://docs.microsoft.com/en-us/azure/azure-databricks/quickstart-create-databricks-workspace-portal or in your databricks workspace you can look for the template \"Azure Blob Storage Import Example Notebook\".\n",
- "\n",
- "\n",
- "Here is an example for setting up the file location to extract the relevant data:\n",
- "\n",
- " file_location = \"wasbs://mycontainer@storageaccountname.blob.core.windows.net/unknown/unknown/unknown-bigdataset-unknown/my_iterate_parking_inputs/2018/°/°/data.csv\" \n",
- "file_type = \"csv\"\n"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### b. Connect Blob to Power Bi (Small Data only)\n",
- "1. Download and Open PowerBi Desktop\n",
- "2. Select “Get Data” and click on “Azure Blob Storage” >> Connect\n",
- "3. Add your storage account and enter your storage key.\n",
- "4. Select the container where your Data Collection is stored and click on Edit. \n",
- "5. In the query editor, click under “Name” column and add your Storage account Model path into the filter. Note: if you want to only look into files from a specific year or month, just expand the filter path. For example, just look into March data: /modeldata/subscriptionid>/resourcegroupname>/workspacename>/webservicename>/modelname>/modelversion>/identifier>/year>/3\n",
- "6. Click on the double arrow aside the “Content” column to combine the files. \n",
- "7. Click OK and the data will preload.\n",
- "8. You can now click Close and Apply and start building your custom reports on your Model Input data."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Disable Data Collection"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "aks_service.update(collect_model_data=False)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Clean up"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%time\n",
- "aks_service.delete()\n",
- "image.delete()\n",
- "model.delete()"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "marthalc"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Enabling Data Collection for Models in Production\n",
+ "With this notebook, you can learn how to collect input model data from your Azure Machine Learning service in an Azure Blob storage. Once enabled, this data collected gives you the opportunity:\n",
+ "\n",
+ "* Monitor data drifts as production data enters your model\n",
+ "* Make better decisions on when to retrain or optimize your model\n",
+ "* Retrain your model with the data collected\n",
+ "\n",
+ "## What data is collected?\n",
+ "* Model input data (voice, images, and video are not supported) from services deployed in Azure Kubernetes Cluster (AKS)\n",
+ "* Model predictions using production input data.\n",
+ "\n",
+ "**Note:** pre-aggregation or pre-calculations on this data are done by user and not included in this version of the product.\n",
+ "\n",
+ "## What is different compared to standard production deployment process?\n",
+ "1. Update scoring file.\n",
+ "2. Update yml file with new dependency.\n",
+ "3. Update aks configuration.\n",
+ "4. Build new image and deploy it. "
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## 1. Import your dependencies"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core import Workspace, Run\n",
+ "from azureml.core.compute import AksCompute, ComputeTarget\n",
+ "from azureml.core.webservice import Webservice, AksWebservice\n",
+ "from azureml.core.image import Image\n",
+ "from azureml.core.model import Model\n",
+ "\n",
+ "import azureml.core\n",
+ "print(azureml.core.VERSION)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## 2. Set up your configuration and create a workspace\n",
+ "Follow Notebook 00 instructions to do this.\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "ws = Workspace.from_config()\n",
+ "print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep = '\\n')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## 3. Register Model\n",
+ "Register an existing trained model, add descirption and tags."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "#Register the model\n",
+ "from azureml.core.model import Model\n",
+ "model = Model.register(model_path = \"sklearn_regression_model.pkl\", # this points to a local file\n",
+ " model_name = \"sklearn_regression_model.pkl\", # this is the name the model is registered as\n",
+ " tags = {'area': \"diabetes\", 'type': \"regression\"},\n",
+ " description = \"Ridge regression model to predict diabetes\",\n",
+ " workspace = ws)\n",
+ "\n",
+ "print(model.name, model.description, model.version)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## 4. *Update your scoring file with Data Collection*\n",
+ "The file below, compared to the file used in notebook 11, has the following changes:\n",
+ "### a. Import the module\n",
+ "```python \n",
+ "from azureml.monitoring import ModelDataCollector```\n",
+ "### b. In your init function add:\n",
+ "```python \n",
+ "global inputs_dc, prediction_d\n",
+ "inputs_dc = ModelDataCollector(\"best_model\", identifier=\"inputs\", feature_names=[\"feat1\", \"feat2\", \"feat3\", \"feat4\", \"feat5\", \"Feat6\"])\n",
+ "prediction_dc = ModelDataCollector(\"best_model\", identifier=\"predictions\", feature_names=[\"prediction1\", \"prediction2\"])```\n",
+ " \n",
+ "* Identifier: Identifier is later used for building the folder structure in your Blob, it can be used to divide \"raw\" data versus \"processed\".\n",
+ "* CorrelationId: is an optional parameter, you do not need to set it up if your model doesn't require it. Having a correlationId in place does help you for easier mapping with other data. (Examples include: LoanNumber, CustomerId, etc.)\n",
+ "* Feature Names: These need to be set up in the order of your features in order for them to have column names when the .csv is created.\n",
+ "\n",
+ "### c. In your run function add:\n",
+ "```python\n",
+ "inputs_dc.collect(data)\n",
+ "prediction_dc.collect(result)```"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%writefile score.py\n",
+ "import pickle\n",
+ "import json\n",
+ "import numpy \n",
+ "from sklearn.externals import joblib\n",
+ "from sklearn.linear_model import Ridge\n",
+ "from azureml.core.model import Model\n",
+ "from azureml.monitoring import ModelDataCollector\n",
+ "import time\n",
+ "\n",
+ "def init():\n",
+ " global model\n",
+ " print (\"model initialized\" + time.strftime(\"%H:%M:%S\"))\n",
+ " # note here \"sklearn_regression_model.pkl\" is the name of the model registered under the workspace\n",
+ " # this call should return the path to the model.pkl file on the local disk.\n",
+ " model_path = Model.get_model_path(model_name = 'sklearn_regression_model.pkl')\n",
+ " # deserialize the model file back into a sklearn model\n",
+ " model = joblib.load(model_path)\n",
+ " global inputs_dc, prediction_dc\n",
+ " # this setup will help us save our inputs under the \"inputs\" path in our Azure Blob\n",
+ " inputs_dc = ModelDataCollector(model_name=\"sklearn_regression_model\", identifier=\"inputs\", feature_names=[\"feat1\", \"feat2\"]) \n",
+ " # this setup will help us save our ipredictions under the \"predictions\" path in our Azure Blob\n",
+ " prediction_dc = ModelDataCollector(\"sklearn_regression_model\", identifier=\"predictions\", feature_names=[\"prediction1\", \"prediction2\"]) \n",
+ " \n",
+ "# note you can pass in multiple rows for scoring\n",
+ "def run(raw_data):\n",
+ " global inputs_dc, prediction_dc\n",
+ " try:\n",
+ " data = json.loads(raw_data)['data']\n",
+ " data = numpy.array(data)\n",
+ " result = model.predict(data)\n",
+ " print (\"saving input data\" + time.strftime(\"%H:%M:%S\"))\n",
+ " inputs_dc.collect(data) #this call is saving our input data into our blob\n",
+ " prediction_dc.collect(result)#this call is saving our prediction data into our blob\n",
+ " print (\"saving prediction data\" + time.strftime(\"%H:%M:%S\"))\n",
+ " # you can return any data type as long as it is JSON-serializable\n",
+ " return result.tolist()\n",
+ " except Exception as e:\n",
+ " error = str(e)\n",
+ " print (error + time.strftime(\"%H:%M:%S\"))\n",
+ " return error"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## 5. *Update your myenv.yml file with the required module*"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.conda_dependencies import CondaDependencies \n",
+ "\n",
+ "myenv = CondaDependencies.create(conda_packages=['numpy','scikit-learn'])\n",
+ "myenv.add_pip_package(\"azureml-monitoring\")\n",
+ "\n",
+ "with open(\"myenv.yml\",\"w\") as f:\n",
+ " f.write(myenv.serialize_to_string())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## 6. Create your new Image"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.image import ContainerImage\n",
+ "\n",
+ "image_config = ContainerImage.image_configuration(execution_script = \"score.py\",\n",
+ " runtime = \"python\",\n",
+ " conda_file = \"myenv.yml\",\n",
+ " description = \"Image with ridge regression model\",\n",
+ " tags = {'area': \"diabetes\", 'type': \"regression\"}\n",
+ " )\n",
+ "\n",
+ "image = ContainerImage.create(name = \"myimage1\",\n",
+ " # this is the model object\n",
+ " models = [model],\n",
+ " image_config = image_config,\n",
+ " workspace = ws)\n",
+ "\n",
+ "image.wait_for_creation(show_output = True)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "print(model.name, model.description, model.version)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## 7. Deploy to AKS service"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create AKS compute if you haven't done so."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Use the default configuration (can also provide parameters to customize)\n",
+ "prov_config = AksCompute.provisioning_configuration()\n",
+ "\n",
+ "aks_name = 'my-aks-test1' \n",
+ "# Create the cluster\n",
+ "aks_target = ComputeTarget.create(workspace = ws, \n",
+ " name = aks_name, \n",
+ " provisioning_configuration = prov_config)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%time\n",
+ "aks_target.wait_for_completion(show_output = True)\n",
+ "print(aks_target.provisioning_state)\n",
+ "print(aks_target.provisioning_errors)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "If you already have a cluster you can attach the service to it:"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "scrolled": true
+ },
+ "source": [
+ "```python \n",
+ " %%time\n",
+ " resource_id = '/subscriptions//resourcegroups//providers/Microsoft.ContainerService/managedClusters/'\n",
+ " create_name= 'myaks4'\n",
+ " attach_config = AksCompute.attach_configuration(resource_id=resource_id)\n",
+ " aks_target = ComputeTarget.attach(workspace = ws, \n",
+ " name = create_name, \n",
+ " attach_configuration=attach_config)\n",
+ " ## Wait for the operation to complete\n",
+ " aks_target.wait_for_provisioning(True)```"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### a. *Activate Data Collection and App Insights through updating AKS Webservice configuration*\n",
+ "In order to enable Data Collection and App Insights in your service you will need to update your AKS configuration file:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "#Set the web service configuration\n",
+ "aks_config = AksWebservice.deploy_configuration(collect_model_data=True, enable_app_insights=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### b. Deploy your service"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "if aks_target.provisioning_state== \"Succeeded\": \n",
+ " aks_service_name ='aks-w-dc0'\n",
+ " aks_service = Webservice.deploy_from_image(workspace = ws, \n",
+ " name = aks_service_name,\n",
+ " image = image,\n",
+ " deployment_config = aks_config,\n",
+ " deployment_target = aks_target\n",
+ " )\n",
+ " aks_service.wait_for_deployment(show_output = True)\n",
+ " print(aks_service.state)\n",
+ "else: \n",
+ " raise ValueError(\"aks provisioning failed, can't deploy service\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## 8. Test your service and send some data\n",
+ "**Note**: It will take around 15 mins for your data to appear in your blob.\n",
+ "The data will appear in your Azure Blob following this format:\n",
+ "\n",
+ "/modeldata/subscriptionid/resourcegroupname/workspacename/webservicename/modelname/modelversion/identifier/year/month/day/data.csv "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%time\n",
+ "import json\n",
+ "\n",
+ "test_sample = json.dumps({'data': [\n",
+ " [1,2,3,4,54,6,7,8,88,10], \n",
+ " [10,9,8,37,36,45,4,33,2,1]\n",
+ "]})\n",
+ "test_sample = bytes(test_sample,encoding = 'utf8')\n",
+ "\n",
+ "if aks_service.state == \"Healthy\":\n",
+ " prediction = aks_service.run(input_data=test_sample)\n",
+ " print(prediction)\n",
+ "else:\n",
+ " raise ValueError(\"Service deployment isn't healthy, can't call the service\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## 9. Validate you data and analyze it\n",
+ "You can look into your data following this path format in your Azure Blob (it takes up to 15 minutes for the data to appear):\n",
+ "\n",
+ "/modeldata/**subscriptionid>**/**resourcegroupname>**/**workspacename>**/**webservicename>**/**modelname>**/**modelversion>>**/**identifier>**/*year/month/day*/data.csv \n",
+ "\n",
+ "For doing further analysis you have multiple options:"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### a. Create DataBricks cluter and connect it to your blob\n",
+ "https://docs.microsoft.com/en-us/azure/azure-databricks/quickstart-create-databricks-workspace-portal or in your databricks workspace you can look for the template \"Azure Blob Storage Import Example Notebook\".\n",
+ "\n",
+ "\n",
+ "Here is an example for setting up the file location to extract the relevant data:\n",
+ "\n",
+ " file_location = \"wasbs://mycontainer@storageaccountname.blob.core.windows.net/unknown/unknown/unknown-bigdataset-unknown/my_iterate_parking_inputs/2018/°/°/data.csv\" \n",
+ "file_type = \"csv\"\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### b. Connect Blob to Power Bi (Small Data only)\n",
+ "1. Download and Open PowerBi Desktop\n",
+ "2. Select \u201cGet Data\u201d and click on \u201cAzure Blob Storage\u201d >> Connect\n",
+ "3. Add your storage account and enter your storage key.\n",
+ "4. Select the container where your Data Collection is stored and click on Edit. \n",
+ "5. In the query editor, click under \u201cName\u201d column and add your Storage account Model path into the filter. Note: if you want to only look into files from a specific year or month, just expand the filter path. For example, just look into March data: /modeldata/subscriptionid>/resourcegroupname>/workspacename>/webservicename>/modelname>/modelversion>/identifier>/year>/3\n",
+ "6. Click on the double arrow aside the \u201cContent\u201d column to combine the files. \n",
+ "7. Click OK and the data will preload.\n",
+ "8. You can now click Close and Apply and start building your custom reports on your Model Input data."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Disable Data Collection"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "aks_service.update(collect_model_data=False)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Clean up"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%time\n",
+ "aks_service.delete()\n",
+ "image.delete()\n",
+ "model.delete()"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python [default]",
- "language": "python",
- "name": "python3"
+ "metadata": {
+ "authors": [
+ {
+ "name": "marthalc"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.5"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.5"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/deployment/onnx/README.md b/how-to-use-azureml/deployment/onnx/README.md
index b16a7862..2e681e24 100644
--- a/how-to-use-azureml/deployment/onnx/README.md
+++ b/how-to-use-azureml/deployment/onnx/README.md
@@ -4,7 +4,7 @@ These tutorials show how to create and deploy Open Neural Network eXchange ([ONN
## Tutorials
-0. [Configure your Azure Machine Learning Workspace](https://github.com/Azure/MachineLearningNotebooks/blob/master/configuration.ipynb)
+0. [Configure your Azure Machine Learning Workspace](../../../configuration.ipynb)
#### Obtain models from the [ONNX Model Zoo](https://github.com/onnx/models) and deploy with ONNX Runtime Inference
1. [Handwritten Digit Classification (MNIST)](https://github.com/Azure/MachineLearningNotebooks/blob/master/how-to-use-azureml/deployment/onnx/onnx-inference-mnist-deploy.ipynb)
diff --git a/how-to-use-azureml/deployment/onnx/onnx-convert-aml-deploy-tinyyolo.ipynb b/how-to-use-azureml/deployment/onnx/onnx-convert-aml-deploy-tinyyolo.ipynb
index 9b27b555..4603e8ad 100644
--- a/how-to-use-azureml/deployment/onnx/onnx-convert-aml-deploy-tinyyolo.ipynb
+++ b/how-to-use-azureml/deployment/onnx/onnx-convert-aml-deploy-tinyyolo.ipynb
@@ -1,435 +1,435 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved. \n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# YOLO Real-time Object Detection using ONNX on AzureML\n",
- "\n",
- "This example shows how to convert the TinyYOLO model from CoreML to ONNX and operationalize it as a web service using Azure Machine Learning services and the ONNX Runtime.\n",
- "\n",
- "## What is ONNX\n",
- "ONNX is an open format for representing machine learning and deep learning models. ONNX enables open and interoperable AI by enabling data scientists and developers to use the tools of their choice without worrying about lock-in and flexibility to deploy to a variety of platforms. ONNX is developed and supported by a community of partners including Microsoft, Facebook, and Amazon. For more information, explore the [ONNX website](http://onnx.ai).\n",
- "\n",
- "## YOLO Details\n",
- "You Only Look Once (YOLO) is a state-of-the-art, real-time object detection system. For more information about YOLO, please visit the [YOLO website](https://pjreddie.com/darknet/yolo/)."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Prerequisites\n",
- "\n",
- "To make the best use of your time, make sure you have done the following:\n",
- "\n",
- "* Understand the [architecture and terms](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture) introduced by Azure Machine Learning\n",
- "* Go through the [00.configuration.ipynb](../00.configuration.ipynb) notebook to:\n",
- " * install the AML SDK\n",
- " * create a workspace and its configuration file (config.json)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Check core SDK version number\n",
- "import azureml.core\n",
- "\n",
- "print(\"SDK version:\", azureml.core.VERSION)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Install necessary packages\n",
- "\n",
- "You'll need to run the following commands to use this tutorial:\n",
- "\n",
- "```sh\n",
- "pip install onnxmltools\n",
- "pip install coremltools # use this on Linux and Mac\n",
- "pip install git+https://github.com/apple/coremltools # use this on Windows\n",
- "```"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Convert model to ONNX\n",
- "\n",
- "First we download the CoreML model. We use the CoreML model listed at https://coreml.store/tinyyolo. This may take a few minutes."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import urllib.request\n",
- "\n",
- "onnx_model_url = \"https://s3-us-west-2.amazonaws.com/coreml-models/TinyYOLO.mlmodel\"\n",
- "urllib.request.urlretrieve(onnx_model_url, filename=\"TinyYOLO.mlmodel\")\n"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Then we use ONNXMLTools to convert the model."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import onnxmltools\n",
- "import coremltools\n",
- "\n",
- "# Load a CoreML model\n",
- "coreml_model = coremltools.utils.load_spec('TinyYOLO.mlmodel')\n",
- "\n",
- "# Convert from CoreML into ONNX\n",
- "onnx_model = onnxmltools.convert_coreml(coreml_model, 'TinyYOLOv2')\n",
- "\n",
- "# Save ONNX model\n",
- "onnxmltools.utils.save_model(onnx_model, 'tinyyolov2.onnx')\n",
- "\n",
- "import os\n",
- "print(os.path.getsize('tinyyolov2.onnx'))"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Deploying as a web service with Azure ML\n",
- "\n",
- "### Load Azure ML workspace\n",
- "\n",
- "We begin by instantiating a workspace object from the existing workspace created earlier in the configuration notebook."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core import Workspace\n",
- "\n",
- "ws = Workspace.from_config()\n",
- "print(ws.name, ws.location, ws.resource_group, sep = '\\n')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Registering your model with Azure ML\n",
- "\n",
- "Now we upload the model and register it in the workspace."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.model import Model\n",
- "\n",
- "model = Model.register(model_path = \"tinyyolov2.onnx\",\n",
- " model_name = \"tinyyolov2\",\n",
- " tags = {\"onnx\": \"demo\"},\n",
- " description = \"TinyYOLO\",\n",
- " workspace = ws)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Displaying your registered models\n",
- "\n",
- "You can optionally list out all the models that you have registered in this workspace."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "models = ws.models\n",
- "for name, m in models.items():\n",
- " print(\"Name:\", name,\"\\tVersion:\", m.version, \"\\tDescription:\", m.description, m.tags)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Write scoring file\n",
- "\n",
- "We are now going to deploy our ONNX model on Azure ML using the ONNX Runtime. We begin by writing a score.py file that will be invoked by the web service call. The `init()` function is called once when the container is started so we load the model using the ONNX Runtime into a global session object."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%writefile score.py\n",
- "import json\n",
- "import time\n",
- "import sys\n",
- "import os\n",
- "from azureml.core.model import Model\n",
- "import numpy as np # we're going to use numpy to process input and output data\n",
- "import onnxruntime # to inference ONNX models, we use the ONNX Runtime\n",
- "\n",
- "def init():\n",
- " global session\n",
- " model = Model.get_model_path(model_name = 'tinyyolov2')\n",
- " session = onnxruntime.InferenceSession(model)\n",
- "\n",
- "def preprocess(input_data_json):\n",
- " # convert the JSON data into the tensor input\n",
- " return np.array(json.loads(input_data_json)['data']).astype('float32')\n",
- "\n",
- "def postprocess(result):\n",
- " return np.array(result).tolist()\n",
- "\n",
- "def run(input_data_json):\n",
- " try:\n",
- " start = time.time() # start timer\n",
- " input_data = preprocess(input_data_json)\n",
- " input_name = session.get_inputs()[0].name # get the id of the first input of the model \n",
- " result = session.run([], {input_name: input_data})\n",
- " end = time.time() # stop timer\n",
- " return {\"result\": postprocess(result),\n",
- " \"time\": end - start}\n",
- " except Exception as e:\n",
- " result = str(e)\n",
- " return {\"error\": result}"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create container image\n",
- "First we create a YAML file that specifies which dependencies we would like to see in our container."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.conda_dependencies import CondaDependencies \n",
- "\n",
- "myenv = CondaDependencies.create(pip_packages=[\"numpy\",\"onnxruntime\",\"azureml-core\"])\n",
- "\n",
- "with open(\"myenv.yml\",\"w\") as f:\n",
- " f.write(myenv.serialize_to_string())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Then we have Azure ML create the container. This step will likely take a few minutes."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.image import ContainerImage\n",
- "\n",
- "image_config = ContainerImage.image_configuration(execution_script = \"score.py\",\n",
- " runtime = \"python\",\n",
- " conda_file = \"myenv.yml\",\n",
- " description = \"TinyYOLO ONNX Demo\",\n",
- " tags = {\"demo\": \"onnx\"}\n",
- " )\n",
- "\n",
- "\n",
- "image = ContainerImage.create(name = \"onnxyolo\",\n",
- " models = [model],\n",
- " image_config = image_config,\n",
- " workspace = ws)\n",
- "\n",
- "image.wait_for_creation(show_output = True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "In case you need to debug your code, the next line of code accesses the log file."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "print(image.image_build_log_uri)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "We're all set! Let's get our model chugging.\n",
- "\n",
- "### Deploy the container image"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.webservice import AciWebservice\n",
- "\n",
- "aciconfig = AciWebservice.deploy_configuration(cpu_cores = 1, \n",
- " memory_gb = 1, \n",
- " tags = {'demo': 'onnx'}, \n",
- " description = 'web service for TinyYOLO ONNX model')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "The following cell will likely take a few minutes to run as well."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.webservice import Webservice\n",
- "from random import randint\n",
- "\n",
- "aci_service_name = 'onnx-tinyyolo'+str(randint(0,100))\n",
- "print(\"Service\", aci_service_name)\n",
- "\n",
- "aci_service = Webservice.deploy_from_image(deployment_config = aciconfig,\n",
- " image = image,\n",
- " name = aci_service_name,\n",
- " workspace = ws)\n",
- "\n",
- "aci_service.wait_for_deployment(True)\n",
- "print(aci_service.state)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "In case the deployment fails, you can check the logs. Make sure to delete your aci_service before trying again."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "if aci_service.state != 'Healthy':\n",
- " # run this command for debugging.\n",
- " print(aci_service.get_logs())\n",
- " aci_service.delete()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Success!\n",
- "\n",
- "If you've made it this far, you've deployed a working web service that does object detection using an ONNX model. You can get the URL for the webservice with the code below."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "print(aci_service.scoring_uri)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "When you are eventually done using the web service, remember to delete it."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "#aci_service.delete()"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "onnx"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved. \n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# YOLO Real-time Object Detection using ONNX on AzureML\n",
+ "\n",
+ "This example shows how to convert the TinyYOLO model from CoreML to ONNX and operationalize it as a web service using Azure Machine Learning services and the ONNX Runtime.\n",
+ "\n",
+ "## What is ONNX\n",
+ "ONNX is an open format for representing machine learning and deep learning models. ONNX enables open and interoperable AI by enabling data scientists and developers to use the tools of their choice without worrying about lock-in and flexibility to deploy to a variety of platforms. ONNX is developed and supported by a community of partners including Microsoft, Facebook, and Amazon. For more information, explore the [ONNX website](http://onnx.ai).\n",
+ "\n",
+ "## YOLO Details\n",
+ "You Only Look Once (YOLO) is a state-of-the-art, real-time object detection system. For more information about YOLO, please visit the [YOLO website](https://pjreddie.com/darknet/yolo/)."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Prerequisites\n",
+ "\n",
+ "To make the best use of your time, make sure you have done the following:\n",
+ "\n",
+ "* Understand the [architecture and terms](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture) introduced by Azure Machine Learning\n",
+ "* Go through the [configuration](../../../configuration.ipynb) notebook to:\n",
+ " * install the AML SDK\n",
+ " * create a workspace and its configuration file (config.json)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Check core SDK version number\n",
+ "import azureml.core\n",
+ "\n",
+ "print(\"SDK version:\", azureml.core.VERSION)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Install necessary packages\n",
+ "\n",
+ "You'll need to run the following commands to use this tutorial:\n",
+ "\n",
+ "```sh\n",
+ "pip install onnxmltools\n",
+ "pip install coremltools # use this on Linux and Mac\n",
+ "pip install git+https://github.com/apple/coremltools # use this on Windows\n",
+ "```"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Convert model to ONNX\n",
+ "\n",
+ "First we download the CoreML model. We use the CoreML model from [Matthijs Hollemans's tutorial](https://github.com/hollance/YOLO-CoreML-MPSNNGraph). This may take a few minutes."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import urllib.request\n",
+ "\n",
+ "coreml_model_url = \"https://github.com/hollance/YOLO-CoreML-MPSNNGraph/raw/master/TinyYOLO-CoreML/TinyYOLO-CoreML/TinyYOLO.mlmodel\"\n",
+ "urllib.request.urlretrieve(coreml_model_url, filename=\"TinyYOLO.mlmodel\")\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Then we use ONNXMLTools to convert the model."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import onnxmltools\n",
+ "import coremltools\n",
+ "\n",
+ "# Load a CoreML model\n",
+ "coreml_model = coremltools.utils.load_spec('TinyYOLO.mlmodel')\n",
+ "\n",
+ "# Convert from CoreML into ONNX\n",
+ "onnx_model = onnxmltools.convert_coreml(coreml_model, 'TinyYOLOv2')\n",
+ "\n",
+ "# Save ONNX model\n",
+ "onnxmltools.utils.save_model(onnx_model, 'tinyyolov2.onnx')\n",
+ "\n",
+ "import os\n",
+ "print(os.path.getsize('tinyyolov2.onnx'))"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Deploying as a web service with Azure ML\n",
+ "\n",
+ "### Load Azure ML workspace\n",
+ "\n",
+ "We begin by instantiating a workspace object from the existing workspace created earlier in the configuration notebook."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core import Workspace\n",
+ "\n",
+ "ws = Workspace.from_config()\n",
+ "print(ws.name, ws.location, ws.resource_group, sep = '\\n')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Registering your model with Azure ML\n",
+ "\n",
+ "Now we upload the model and register it in the workspace."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.model import Model\n",
+ "\n",
+ "model = Model.register(model_path = \"tinyyolov2.onnx\",\n",
+ " model_name = \"tinyyolov2\",\n",
+ " tags = {\"onnx\": \"demo\"},\n",
+ " description = \"TinyYOLO\",\n",
+ " workspace = ws)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Displaying your registered models\n",
+ "\n",
+ "You can optionally list out all the models that you have registered in this workspace."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "models = ws.models\n",
+ "for name, m in models.items():\n",
+ " print(\"Name:\", name,\"\\tVersion:\", m.version, \"\\tDescription:\", m.description, m.tags)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Write scoring file\n",
+ "\n",
+ "We are now going to deploy our ONNX model on Azure ML using the ONNX Runtime. We begin by writing a score.py file that will be invoked by the web service call. The `init()` function is called once when the container is started so we load the model using the ONNX Runtime into a global session object."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%writefile score.py\n",
+ "import json\n",
+ "import time\n",
+ "import sys\n",
+ "import os\n",
+ "from azureml.core.model import Model\n",
+ "import numpy as np # we're going to use numpy to process input and output data\n",
+ "import onnxruntime # to inference ONNX models, we use the ONNX Runtime\n",
+ "\n",
+ "def init():\n",
+ " global session\n",
+ " model = Model.get_model_path(model_name = 'tinyyolov2')\n",
+ " session = onnxruntime.InferenceSession(model)\n",
+ "\n",
+ "def preprocess(input_data_json):\n",
+ " # convert the JSON data into the tensor input\n",
+ " return np.array(json.loads(input_data_json)['data']).astype('float32')\n",
+ "\n",
+ "def postprocess(result):\n",
+ " return np.array(result).tolist()\n",
+ "\n",
+ "def run(input_data_json):\n",
+ " try:\n",
+ " start = time.time() # start timer\n",
+ " input_data = preprocess(input_data_json)\n",
+ " input_name = session.get_inputs()[0].name # get the id of the first input of the model \n",
+ " result = session.run([], {input_name: input_data})\n",
+ " end = time.time() # stop timer\n",
+ " return {\"result\": postprocess(result),\n",
+ " \"time\": end - start}\n",
+ " except Exception as e:\n",
+ " result = str(e)\n",
+ " return {\"error\": result}"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create container image\n",
+ "First we create a YAML file that specifies which dependencies we would like to see in our container."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.conda_dependencies import CondaDependencies \n",
+ "\n",
+ "myenv = CondaDependencies.create(pip_packages=[\"numpy\",\"onnxruntime\",\"azureml-core\"])\n",
+ "\n",
+ "with open(\"myenv.yml\",\"w\") as f:\n",
+ " f.write(myenv.serialize_to_string())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Then we have Azure ML create the container. This step will likely take a few minutes."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.image import ContainerImage\n",
+ "\n",
+ "image_config = ContainerImage.image_configuration(execution_script = \"score.py\",\n",
+ " runtime = \"python\",\n",
+ " conda_file = \"myenv.yml\",\n",
+ " description = \"TinyYOLO ONNX Demo\",\n",
+ " tags = {\"demo\": \"onnx\"}\n",
+ " )\n",
+ "\n",
+ "\n",
+ "image = ContainerImage.create(name = \"onnxyolo\",\n",
+ " models = [model],\n",
+ " image_config = image_config,\n",
+ " workspace = ws)\n",
+ "\n",
+ "image.wait_for_creation(show_output = True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "In case you need to debug your code, the next line of code accesses the log file."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "print(image.image_build_log_uri)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "We're all set! Let's get our model chugging.\n",
+ "\n",
+ "### Deploy the container image"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.webservice import AciWebservice\n",
+ "\n",
+ "aciconfig = AciWebservice.deploy_configuration(cpu_cores = 1, \n",
+ " memory_gb = 1, \n",
+ " tags = {'demo': 'onnx'}, \n",
+ " description = 'web service for TinyYOLO ONNX model')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "The following cell will likely take a few minutes to run as well."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.webservice import Webservice\n",
+ "from random import randint\n",
+ "\n",
+ "aci_service_name = 'onnx-tinyyolo'+str(randint(0,100))\n",
+ "print(\"Service\", aci_service_name)\n",
+ "\n",
+ "aci_service = Webservice.deploy_from_image(deployment_config = aciconfig,\n",
+ " image = image,\n",
+ " name = aci_service_name,\n",
+ " workspace = ws)\n",
+ "\n",
+ "aci_service.wait_for_deployment(True)\n",
+ "print(aci_service.state)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "In case the deployment fails, you can check the logs. Make sure to delete your aci_service before trying again."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "if aci_service.state != 'Healthy':\n",
+ " # run this command for debugging.\n",
+ " print(aci_service.get_logs())\n",
+ " aci_service.delete()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Success!\n",
+ "\n",
+ "If you've made it this far, you've deployed a working web service that does object detection using an ONNX model. You can get the URL for the webservice with the code below."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "print(aci_service.scoring_uri)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "When you are eventually done using the web service, remember to delete it."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "#aci_service.delete()"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "onnx"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.5.6"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.5.6"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/deployment/onnx/onnx-inference-facial-expression-recognition-deploy.ipynb b/how-to-use-azureml/deployment/onnx/onnx-inference-facial-expression-recognition-deploy.ipynb
index b067a21d..69936f1f 100644
--- a/how-to-use-azureml/deployment/onnx/onnx-inference-facial-expression-recognition-deploy.ipynb
+++ b/how-to-use-azureml/deployment/onnx/onnx-inference-facial-expression-recognition-deploy.ipynb
@@ -1,809 +1,809 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved. \n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Facial Expression Recognition (FER+) using ONNX Runtime on Azure ML\n",
- "\n",
- "This example shows how to deploy an image classification neural network using the Facial Expression Recognition ([FER](https://www.kaggle.com/c/challenges-in-representation-learning-facial-expression-recognition-challenge/data)) dataset and Open Neural Network eXchange format ([ONNX](http://aka.ms/onnxdocarticle)) on the Azure Machine Learning platform. This tutorial will show you how to deploy a FER+ model from the [ONNX model zoo](https://github.com/onnx/models), use it to make predictions using ONNX Runtime Inference, and deploy it as a web service in Azure.\n",
- "\n",
- "Throughout this tutorial, we will be referring to ONNX, a neural network exchange format used to represent deep learning models. With ONNX, AI developers can more easily move models between state-of-the-art tools (CNTK, PyTorch, Caffe, MXNet, TensorFlow) and choose the combination that is best for them. ONNX is developed and supported by a community of partners including Microsoft AI, Facebook, and Amazon. For more information, explore the [ONNX website](http://onnx.ai) and [open source files](https://github.com/onnx).\n",
- "\n",
- "[ONNX Runtime](https://aka.ms/onnxruntime-python) is the runtime engine that enables evaluation of trained machine learning (Traditional ML and Deep Learning) models with high performance and low resource utilization. We use the CPU version of ONNX Runtime in this tutorial, but will soon be releasing an additional tutorial for deploying this model using ONNX Runtime GPU.\n",
- "\n",
- "#### Tutorial Objectives:\n",
- "\n",
- "1. Describe the FER+ dataset and pretrained Convolutional Neural Net ONNX model for Emotion Recognition, stored in the ONNX model zoo.\n",
- "2. Deploy and run the pretrained FER+ ONNX model on an Azure Machine Learning instance\n",
- "3. Predict labels for test set data points in the cloud using ONNX Runtime and Azure ML"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Prerequisites\n",
- "\n",
- "### 1. Install Azure ML SDK and create a new workspace\n",
- "Please follow [Azure ML configuration notebook](https://github.com/Azure/MachineLearningNotebooks/blob/master/00.configuration.ipynb) to set up your environment.\n",
- "\n",
- "### 2. Install additional packages needed for this Notebook\n",
- "You need to install the popular plotting library `matplotlib`, the image manipulation library `opencv`, and the `onnx` library in the conda environment where Azure Maching Learning SDK is installed.\n",
- "\n",
- "```sh\n",
- "(myenv) $ pip install matplotlib onnx opencv-python\n",
- "```\n",
- "\n",
- "**Debugging tip**: Make sure that to activate your virtual environment (myenv) before you re-launch this notebook using the `jupyter notebook` comand. Choose the respective Python kernel for your new virtual environment using the `Kernel > Change Kernel` menu above. If you have completed the steps correctly, the upper right corner of your screen should state `Python [conda env:myenv]` instead of `Python [default]`.\n",
- "\n",
- "### 3. Download sample data and pre-trained ONNX model from ONNX Model Zoo.\n",
- "\n",
- "In the following lines of code, we download [the trained ONNX Emotion FER+ model and corresponding test data](https://github.com/onnx/models/tree/master/emotion_ferplus) and place them in the same folder as this tutorial notebook. For more information about the FER+ dataset, please visit Microsoft Researcher Emad Barsoum's [FER+ source data repository](https://github.com/ebarsoum/FERPlus)."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# urllib is a built-in Python library to download files from URLs\n",
- "\n",
- "# Objective: retrieve the latest version of the ONNX Emotion FER+ model files from the\n",
- "# ONNX Model Zoo and save it in the same folder as this tutorial\n",
- "\n",
- "import urllib.request\n",
- "\n",
- "onnx_model_url = \"https://www.cntk.ai/OnnxModels/emotion_ferplus/opset_7/emotion_ferplus.tar.gz\"\n",
- "\n",
- "urllib.request.urlretrieve(onnx_model_url, filename=\"emotion_ferplus.tar.gz\")\n",
- "\n",
- "# the ! magic command tells our jupyter notebook kernel to run the following line of \n",
- "# code from the command line instead of the notebook kernel\n",
- "\n",
- "# We use tar and xvcf to unzip the files we just retrieved from the ONNX model zoo\n",
- "\n",
- "!tar xvzf emotion_ferplus.tar.gz"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Deploy a VM with your ONNX model in the Cloud\n",
- "\n",
- "### Load Azure ML workspace\n",
- "\n",
- "We begin by instantiating a workspace object from the existing workspace created earlier in the configuration notebook."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Check core SDK version number\n",
- "import azureml.core\n",
- "\n",
- "print(\"SDK version:\", azureml.core.VERSION)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core import Workspace\n",
- "\n",
- "ws = Workspace.from_config()\n",
- "print(ws.name, ws.location, ws.resource_group, sep = '\\n')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Registering your model with Azure ML"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "model_dir = \"emotion_ferplus\" # replace this with the location of your model files\n",
- "\n",
- "# leave as is if it's in the same folder as this notebook"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.model import Model\n",
- "\n",
- "model = Model.register(model_path = model_dir + \"/\" + \"model.onnx\",\n",
- " model_name = \"onnx_emotion\",\n",
- " tags = {\"onnx\": \"demo\"},\n",
- " description = \"FER+ emotion recognition CNN from ONNX Model Zoo\",\n",
- " workspace = ws)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Optional: Displaying your registered models\n",
- "\n",
- "This step is not required, so feel free to skip it."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "models = ws.models\n",
- "for name, m in models.items():\n",
- " print(\"Name:\", name,\"\\tVersion:\", m.version, \"\\tDescription:\", m.description, m.tags)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### ONNX FER+ Model Methodology\n",
- "\n",
- "The image classification model we are using is pre-trained using Microsoft's deep learning cognitive toolkit, [CNTK](https://github.com/Microsoft/CNTK), from the [ONNX model zoo](http://github.com/onnx/models). The model zoo has many other models that can be deployed on cloud providers like AzureML without any additional training. To ensure that our cloud deployed model works, we use testing data from the well-known FER+ data set, provided as part of the [trained Emotion Recognition model](https://github.com/onnx/models/tree/master/emotion_ferplus) in the ONNX model zoo.\n",
- "\n",
- "The original Facial Emotion Recognition (FER) Dataset was released in 2013 by Pierre-Luc Carrier and Aaron Courville as part of a [Kaggle Competition](https://www.kaggle.com/c/challenges-in-representation-learning-facial-expression-recognition-challenge/data), but some of the labels are not entirely appropriate for the expression. In the FER+ Dataset, each photo was evaluated by at least 10 croud sourced reviewers, creating a more accurate basis for ground truth. \n",
- "\n",
- "You can see the difference of label quality in the sample model input below. The FER labels are the first word below each image, and the FER+ labels are the second word below each image.\n",
- "\n",
- "\n",
- "\n",
- "***Input: Photos of cropped faces from FER+ Dataset***\n",
- "\n",
- "***Task: Classify each facial image into its appropriate emotions in the emotion table***\n",
- "\n",
- "``` emotion_table = {'neutral':0, 'happiness':1, 'surprise':2, 'sadness':3, 'anger':4, 'disgust':5, 'fear':6, 'contempt':7} ```\n",
- "\n",
- "***Output: Emotion prediction for input image***\n",
- "\n",
- "\n",
- "Remember, once the application is deployed in Azure ML, you can use your own images as input for the model to classify."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# for images and plots in this notebook\n",
- "import matplotlib.pyplot as plt \n",
- "from IPython.display import Image\n",
- "\n",
- "# display images inline\n",
- "%matplotlib inline"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Model Description\n",
- "\n",
- "The FER+ model from the ONNX Model Zoo is summarized by the graphic below. You can see the entire workflow of our pre-trained model in the following image from Barsoum et. al's paper [\"Training Deep Networks for Facial Expression Recognition\n",
- "with Crowd-Sourced Label Distribution\"](https://arxiv.org/pdf/1608.01041.pdf), with our (64 x 64) input images and our output probabilities for each of the labels."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "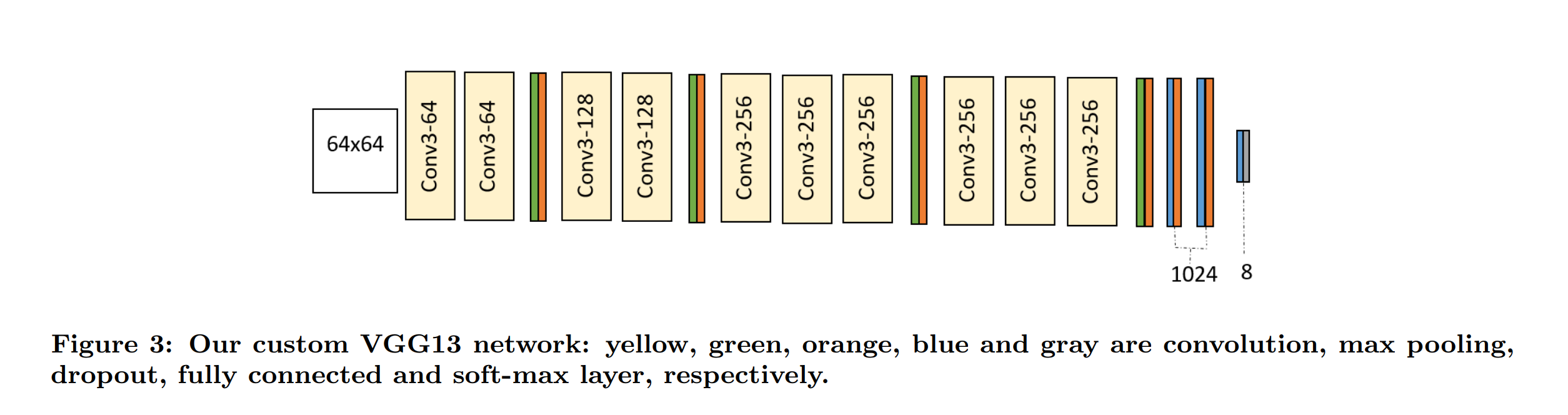"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Specify our Score and Environment Files"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "We are now going to deploy our ONNX Model on AML with inference in ONNX Runtime. We begin by writing a score.py file, which will help us run the model in our Azure ML virtual machine (VM), and then specify our environment by writing a yml file. You will also notice that we import the onnxruntime library to do runtime inference on our ONNX models (passing in input and evaluating out model's predicted output). More information on the API and commands can be found in the [ONNX Runtime documentation](https://aka.ms/onnxruntime).\n",
- "\n",
- "### Write Score File\n",
- "\n",
- "A score file is what tells our Azure cloud service what to do. After initializing our model using azureml.core.model, we start an ONNX Runtime inference session to evaluate the data passed in on our function calls."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%writefile score.py\n",
- "import json\n",
- "import numpy as np\n",
- "import onnxruntime\n",
- "import sys\n",
- "import os\n",
- "from azureml.core.model import Model\n",
- "import time\n",
- "\n",
- "def init():\n",
- " global session, input_name, output_name\n",
- " model = Model.get_model_path(model_name = 'onnx_emotion')\n",
- " session = onnxruntime.InferenceSession(model, None)\n",
- " input_name = session.get_inputs()[0].name\n",
- " output_name = session.get_outputs()[0].name \n",
- " \n",
- "def run(input_data):\n",
- " '''Purpose: evaluate test input in Azure Cloud using onnxruntime.\n",
- " We will call the run function later from our Jupyter Notebook \n",
- " so our azure service can evaluate our model input in the cloud. '''\n",
- "\n",
- " try:\n",
- " # load in our data, convert to readable format\n",
- " data = np.array(json.loads(input_data)['data']).astype('float32')\n",
- " \n",
- " start = time.time()\n",
- " r = session.run([output_name], {input_name : data})\n",
- " end = time.time()\n",
- " \n",
- " result = emotion_map(postprocess(r[0]))\n",
- " \n",
- " result_dict = {\"result\": result,\n",
- " \"time_in_sec\": [end - start]}\n",
- " except Exception as e:\n",
- " result_dict = {\"error\": str(e)}\n",
- " \n",
- " return json.dumps(result_dict)\n",
- "\n",
- "def emotion_map(classes, N=1):\n",
- " \"\"\"Take the most probable labels (output of postprocess) and returns the \n",
- " top N emotional labels that fit the picture.\"\"\"\n",
- " \n",
- " emotion_table = {'neutral':0, 'happiness':1, 'surprise':2, 'sadness':3, \n",
- " 'anger':4, 'disgust':5, 'fear':6, 'contempt':7}\n",
- " \n",
- " emotion_keys = list(emotion_table.keys())\n",
- " emotions = []\n",
- " for i in range(N):\n",
- " emotions.append(emotion_keys[classes[i]])\n",
- " return emotions\n",
- "\n",
- "def softmax(x):\n",
- " \"\"\"Compute softmax values (probabilities from 0 to 1) for each possible label.\"\"\"\n",
- " x = x.reshape(-1)\n",
- " e_x = np.exp(x - np.max(x))\n",
- " return e_x / e_x.sum(axis=0)\n",
- "\n",
- "def postprocess(scores):\n",
- " \"\"\"This function takes the scores generated by the network and \n",
- " returns the class IDs in decreasing order of probability.\"\"\"\n",
- " prob = softmax(scores)\n",
- " prob = np.squeeze(prob)\n",
- " classes = np.argsort(prob)[::-1]\n",
- " return classes"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Write Environment File"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.conda_dependencies import CondaDependencies \n",
- "\n",
- "myenv = CondaDependencies.create(pip_packages=[\"numpy\", \"onnxruntime\", \"azureml-core\"])\n",
- "\n",
- "with open(\"myenv.yml\",\"w\") as f:\n",
- " f.write(myenv.serialize_to_string())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create the Container Image\n",
- "\n",
- "This step will likely take a few minutes."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.image import ContainerImage\n",
- "\n",
- "image_config = ContainerImage.image_configuration(execution_script = \"score.py\",\n",
- " runtime = \"python\",\n",
- " conda_file = \"myenv.yml\",\n",
- " description = \"Emotion ONNX Runtime container\",\n",
- " tags = {\"demo\": \"onnx\"})\n",
- "\n",
- "\n",
- "image = ContainerImage.create(name = \"onnximage\",\n",
- " # this is the model object\n",
- " models = [model],\n",
- " image_config = image_config,\n",
- " workspace = ws)\n",
- "\n",
- "image.wait_for_creation(show_output = True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "In case you need to debug your code, the next line of code accesses the log file."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "print(image.image_build_log_uri)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "We're all done specifying what we want our virtual machine to do. Let's configure and deploy our container image.\n",
- "\n",
- "### Deploy the container image"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.webservice import AciWebservice\n",
- "\n",
- "aciconfig = AciWebservice.deploy_configuration(cpu_cores = 1, \n",
- " memory_gb = 1, \n",
- " tags = {'demo': 'onnx'}, \n",
- " description = 'ONNX for emotion recognition model')"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.webservice import Webservice\n",
- "\n",
- "aci_service_name = 'onnx-demo-emotion'\n",
- "print(\"Service\", aci_service_name)\n",
- "\n",
- "aci_service = Webservice.deploy_from_image(deployment_config = aciconfig,\n",
- " image = image,\n",
- " name = aci_service_name,\n",
- " workspace = ws)\n",
- "\n",
- "aci_service.wait_for_deployment(True)\n",
- "print(aci_service.state)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "The following cell will likely take a few minutes to run as well."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "if aci_service.state != 'Healthy':\n",
- " # run this command for debugging.\n",
- " print(aci_service.get_logs())\n",
- "\n",
- " # If your deployment fails, make sure to delete your aci_service before trying again!\n",
- " # aci_service.delete()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Success!\n",
- "\n",
- "If you've made it this far, you've deployed a working VM with a facial emotion recognition model running in the cloud using Azure ML. Congratulations!\n",
- "\n",
- "Let's see how well our model deals with our test images."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Testing and Evaluation\n",
- "\n",
- "### Useful Helper Functions\n",
- "\n",
- "We preprocess and postprocess our data (see score.py file) using the helper functions specified in the [ONNX FER+ Model page in the Model Zoo repository](https://github.com/onnx/models/tree/master/emotion_ferplus)."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "def emotion_map(classes, N=1):\n",
- " \"\"\"Take the most probable labels (output of postprocess) and returns the \n",
- " top N emotional labels that fit the picture.\"\"\"\n",
- " \n",
- " emotion_table = {'neutral':0, 'happiness':1, 'surprise':2, 'sadness':3, \n",
- " 'anger':4, 'disgust':5, 'fear':6, 'contempt':7}\n",
- " \n",
- " emotion_keys = list(emotion_table.keys())\n",
- " emotions = []\n",
- " for i in range(N):\n",
- " emotions.append(emotion_keys[classes[i]])\n",
- " return emotions\n",
- "\n",
- "def softmax(x):\n",
- " \"\"\"Compute softmax values (probabilities from 0 to 1) for each possible label.\"\"\"\n",
- " x = x.reshape(-1)\n",
- " e_x = np.exp(x - np.max(x))\n",
- " return e_x / e_x.sum(axis=0)\n",
- "\n",
- "def postprocess(scores):\n",
- " \"\"\"This function takes the scores generated by the network and \n",
- " returns the class IDs in decreasing order of probability.\"\"\"\n",
- " prob = softmax(scores)\n",
- " prob = np.squeeze(prob)\n",
- " classes = np.argsort(prob)[::-1]\n",
- " return classes"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Load Test Data\n",
- "\n",
- "These are already in your directory from your ONNX model download (from the model zoo).\n",
- "\n",
- "Notice that our Model Zoo files have a .pb extension. This is because they are [protobuf files (Protocol Buffers)](https://developers.google.com/protocol-buffers/docs/pythontutorial), so we need to read in our data through our ONNX TensorProto reader into a format we can work with, like numerical arrays."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# to manipulate our arrays\n",
- "import numpy as np \n",
- "\n",
- "# read in test data protobuf files included with the model\n",
- "import onnx\n",
- "from onnx import numpy_helper\n",
- "\n",
- "# to use parsers to read in our model/data\n",
- "import json\n",
- "import os\n",
- "\n",
- "test_inputs = []\n",
- "test_outputs = []\n",
- "\n",
- "# read in 3 testing images from .pb files\n",
- "test_data_size = 3\n",
- "\n",
- "for i in np.arange(test_data_size):\n",
- " input_test_data = os.path.join(model_dir, 'test_data_set_{0}'.format(i), 'input_0.pb')\n",
- " output_test_data = os.path.join(model_dir, 'test_data_set_{0}'.format(i), 'output_0.pb')\n",
- " \n",
- " # convert protobuf tensors to np arrays using the TensorProto reader from ONNX\n",
- " tensor = onnx.TensorProto()\n",
- " with open(input_test_data, 'rb') as f:\n",
- " tensor.ParseFromString(f.read())\n",
- " \n",
- " input_data = numpy_helper.to_array(tensor)\n",
- " test_inputs.append(input_data)\n",
- " \n",
- " with open(output_test_data, 'rb') as f:\n",
- " tensor.ParseFromString(f.read())\n",
- " \n",
- " output_data = numpy_helper.to_array(tensor)\n",
- " output_processed = emotion_map(postprocess(output_data[0]))[0]\n",
- " test_outputs.append(output_processed)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {
- "nbpresent": {
- "id": "c3f2f57c-7454-4d3e-b38d-b0946cf066ea"
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved. \n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Facial Expression Recognition (FER+) using ONNX Runtime on Azure ML\n",
+ "\n",
+ "This example shows how to deploy an image classification neural network using the Facial Expression Recognition ([FER](https://www.kaggle.com/c/challenges-in-representation-learning-facial-expression-recognition-challenge/data)) dataset and Open Neural Network eXchange format ([ONNX](http://aka.ms/onnxdocarticle)) on the Azure Machine Learning platform. This tutorial will show you how to deploy a FER+ model from the [ONNX model zoo](https://github.com/onnx/models), use it to make predictions using ONNX Runtime Inference, and deploy it as a web service in Azure.\n",
+ "\n",
+ "Throughout this tutorial, we will be referring to ONNX, a neural network exchange format used to represent deep learning models. With ONNX, AI developers can more easily move models between state-of-the-art tools (CNTK, PyTorch, Caffe, MXNet, TensorFlow) and choose the combination that is best for them. ONNX is developed and supported by a community of partners including Microsoft AI, Facebook, and Amazon. For more information, explore the [ONNX website](http://onnx.ai) and [open source files](https://github.com/onnx).\n",
+ "\n",
+ "[ONNX Runtime](https://aka.ms/onnxruntime-python) is the runtime engine that enables evaluation of trained machine learning (Traditional ML and Deep Learning) models with high performance and low resource utilization. We use the CPU version of ONNX Runtime in this tutorial, but will soon be releasing an additional tutorial for deploying this model using ONNX Runtime GPU.\n",
+ "\n",
+ "#### Tutorial Objectives:\n",
+ "\n",
+ "1. Describe the FER+ dataset and pretrained Convolutional Neural Net ONNX model for Emotion Recognition, stored in the ONNX model zoo.\n",
+ "2. Deploy and run the pretrained FER+ ONNX model on an Azure Machine Learning instance\n",
+ "3. Predict labels for test set data points in the cloud using ONNX Runtime and Azure ML"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Prerequisites\n",
+ "\n",
+ "### 1. Install Azure ML SDK and create a new workspace\n",
+ "Please follow [Azure ML configuration notebook](../../../configuration.ipynb) to set up your environment.\n",
+ "\n",
+ "### 2. Install additional packages needed for this Notebook\n",
+ "You need to install the popular plotting library `matplotlib`, the image manipulation library `opencv`, and the `onnx` library in the conda environment where Azure Maching Learning SDK is installed.\n",
+ "\n",
+ "```sh\n",
+ "(myenv) $ pip install matplotlib onnx opencv-python\n",
+ "```\n",
+ "\n",
+ "**Debugging tip**: Make sure that to activate your virtual environment (myenv) before you re-launch this notebook using the `jupyter notebook` comand. Choose the respective Python kernel for your new virtual environment using the `Kernel > Change Kernel` menu above. If you have completed the steps correctly, the upper right corner of your screen should state `Python [conda env:myenv]` instead of `Python [default]`.\n",
+ "\n",
+ "### 3. Download sample data and pre-trained ONNX model from ONNX Model Zoo.\n",
+ "\n",
+ "In the following lines of code, we download [the trained ONNX Emotion FER+ model and corresponding test data](https://github.com/onnx/models/tree/master/emotion_ferplus) and place them in the same folder as this tutorial notebook. For more information about the FER+ dataset, please visit Microsoft Researcher Emad Barsoum's [FER+ source data repository](https://github.com/ebarsoum/FERPlus)."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# urllib is a built-in Python library to download files from URLs\n",
+ "\n",
+ "# Objective: retrieve the latest version of the ONNX Emotion FER+ model files from the\n",
+ "# ONNX Model Zoo and save it in the same folder as this tutorial\n",
+ "\n",
+ "import urllib.request\n",
+ "\n",
+ "onnx_model_url = \"https://www.cntk.ai/OnnxModels/emotion_ferplus/opset_7/emotion_ferplus.tar.gz\"\n",
+ "\n",
+ "urllib.request.urlretrieve(onnx_model_url, filename=\"emotion_ferplus.tar.gz\")\n",
+ "\n",
+ "# the ! magic command tells our jupyter notebook kernel to run the following line of \n",
+ "# code from the command line instead of the notebook kernel\n",
+ "\n",
+ "# We use tar and xvcf to unzip the files we just retrieved from the ONNX model zoo\n",
+ "\n",
+ "!tar xvzf emotion_ferplus.tar.gz"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Deploy a VM with your ONNX model in the Cloud\n",
+ "\n",
+ "### Load Azure ML workspace\n",
+ "\n",
+ "We begin by instantiating a workspace object from the existing workspace created earlier in the configuration notebook."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Check core SDK version number\n",
+ "import azureml.core\n",
+ "\n",
+ "print(\"SDK version:\", azureml.core.VERSION)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core import Workspace\n",
+ "\n",
+ "ws = Workspace.from_config()\n",
+ "print(ws.name, ws.location, ws.resource_group, sep = '\\n')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Registering your model with Azure ML"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "model_dir = \"emotion_ferplus\" # replace this with the location of your model files\n",
+ "\n",
+ "# leave as is if it's in the same folder as this notebook"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.model import Model\n",
+ "\n",
+ "model = Model.register(model_path = model_dir + \"/\" + \"model.onnx\",\n",
+ " model_name = \"onnx_emotion\",\n",
+ " tags = {\"onnx\": \"demo\"},\n",
+ " description = \"FER+ emotion recognition CNN from ONNX Model Zoo\",\n",
+ " workspace = ws)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Optional: Displaying your registered models\n",
+ "\n",
+ "This step is not required, so feel free to skip it."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "models = ws.models\n",
+ "for name, m in models.items():\n",
+ " print(\"Name:\", name,\"\\tVersion:\", m.version, \"\\tDescription:\", m.description, m.tags)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### ONNX FER+ Model Methodology\n",
+ "\n",
+ "The image classification model we are using is pre-trained using Microsoft's deep learning cognitive toolkit, [CNTK](https://github.com/Microsoft/CNTK), from the [ONNX model zoo](http://github.com/onnx/models). The model zoo has many other models that can be deployed on cloud providers like AzureML without any additional training. To ensure that our cloud deployed model works, we use testing data from the well-known FER+ data set, provided as part of the [trained Emotion Recognition model](https://github.com/onnx/models/tree/master/emotion_ferplus) in the ONNX model zoo.\n",
+ "\n",
+ "The original Facial Emotion Recognition (FER) Dataset was released in 2013 by Pierre-Luc Carrier and Aaron Courville as part of a [Kaggle Competition](https://www.kaggle.com/c/challenges-in-representation-learning-facial-expression-recognition-challenge/data), but some of the labels are not entirely appropriate for the expression. In the FER+ Dataset, each photo was evaluated by at least 10 croud sourced reviewers, creating a more accurate basis for ground truth. \n",
+ "\n",
+ "You can see the difference of label quality in the sample model input below. The FER labels are the first word below each image, and the FER+ labels are the second word below each image.\n",
+ "\n",
+ "\n",
+ "\n",
+ "***Input: Photos of cropped faces from FER+ Dataset***\n",
+ "\n",
+ "***Task: Classify each facial image into its appropriate emotions in the emotion table***\n",
+ "\n",
+ "``` emotion_table = {'neutral':0, 'happiness':1, 'surprise':2, 'sadness':3, 'anger':4, 'disgust':5, 'fear':6, 'contempt':7} ```\n",
+ "\n",
+ "***Output: Emotion prediction for input image***\n",
+ "\n",
+ "\n",
+ "Remember, once the application is deployed in Azure ML, you can use your own images as input for the model to classify."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# for images and plots in this notebook\n",
+ "import matplotlib.pyplot as plt \n",
+ "from IPython.display import Image\n",
+ "\n",
+ "# display images inline\n",
+ "%matplotlib inline"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Model Description\n",
+ "\n",
+ "The FER+ model from the ONNX Model Zoo is summarized by the graphic below. You can see the entire workflow of our pre-trained model in the following image from Barsoum et. al's paper [\"Training Deep Networks for Facial Expression Recognition\n",
+ "with Crowd-Sourced Label Distribution\"](https://arxiv.org/pdf/1608.01041.pdf), with our (64 x 64) input images and our output probabilities for each of the labels."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ ""
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Specify our Score and Environment Files"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "We are now going to deploy our ONNX Model on AML with inference in ONNX Runtime. We begin by writing a score.py file, which will help us run the model in our Azure ML virtual machine (VM), and then specify our environment by writing a yml file. You will also notice that we import the onnxruntime library to do runtime inference on our ONNX models (passing in input and evaluating out model's predicted output). More information on the API and commands can be found in the [ONNX Runtime documentation](https://aka.ms/onnxruntime).\n",
+ "\n",
+ "### Write Score File\n",
+ "\n",
+ "A score file is what tells our Azure cloud service what to do. After initializing our model using azureml.core.model, we start an ONNX Runtime inference session to evaluate the data passed in on our function calls."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%writefile score.py\n",
+ "import json\n",
+ "import numpy as np\n",
+ "import onnxruntime\n",
+ "import sys\n",
+ "import os\n",
+ "from azureml.core.model import Model\n",
+ "import time\n",
+ "\n",
+ "def init():\n",
+ " global session, input_name, output_name\n",
+ " model = Model.get_model_path(model_name = 'onnx_emotion')\n",
+ " session = onnxruntime.InferenceSession(model, None)\n",
+ " input_name = session.get_inputs()[0].name\n",
+ " output_name = session.get_outputs()[0].name \n",
+ " \n",
+ "def run(input_data):\n",
+ " '''Purpose: evaluate test input in Azure Cloud using onnxruntime.\n",
+ " We will call the run function later from our Jupyter Notebook \n",
+ " so our azure service can evaluate our model input in the cloud. '''\n",
+ "\n",
+ " try:\n",
+ " # load in our data, convert to readable format\n",
+ " data = np.array(json.loads(input_data)['data']).astype('float32')\n",
+ " \n",
+ " start = time.time()\n",
+ " r = session.run([output_name], {input_name : data})\n",
+ " end = time.time()\n",
+ " \n",
+ " result = emotion_map(postprocess(r[0]))\n",
+ " \n",
+ " result_dict = {\"result\": result,\n",
+ " \"time_in_sec\": [end - start]}\n",
+ " except Exception as e:\n",
+ " result_dict = {\"error\": str(e)}\n",
+ " \n",
+ " return json.dumps(result_dict)\n",
+ "\n",
+ "def emotion_map(classes, N=1):\n",
+ " \"\"\"Take the most probable labels (output of postprocess) and returns the \n",
+ " top N emotional labels that fit the picture.\"\"\"\n",
+ " \n",
+ " emotion_table = {'neutral':0, 'happiness':1, 'surprise':2, 'sadness':3, \n",
+ " 'anger':4, 'disgust':5, 'fear':6, 'contempt':7}\n",
+ " \n",
+ " emotion_keys = list(emotion_table.keys())\n",
+ " emotions = []\n",
+ " for i in range(N):\n",
+ " emotions.append(emotion_keys[classes[i]])\n",
+ " return emotions\n",
+ "\n",
+ "def softmax(x):\n",
+ " \"\"\"Compute softmax values (probabilities from 0 to 1) for each possible label.\"\"\"\n",
+ " x = x.reshape(-1)\n",
+ " e_x = np.exp(x - np.max(x))\n",
+ " return e_x / e_x.sum(axis=0)\n",
+ "\n",
+ "def postprocess(scores):\n",
+ " \"\"\"This function takes the scores generated by the network and \n",
+ " returns the class IDs in decreasing order of probability.\"\"\"\n",
+ " prob = softmax(scores)\n",
+ " prob = np.squeeze(prob)\n",
+ " classes = np.argsort(prob)[::-1]\n",
+ " return classes"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Write Environment File"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.conda_dependencies import CondaDependencies \n",
+ "\n",
+ "myenv = CondaDependencies.create(pip_packages=[\"numpy\", \"onnxruntime\", \"azureml-core\"])\n",
+ "\n",
+ "with open(\"myenv.yml\",\"w\") as f:\n",
+ " f.write(myenv.serialize_to_string())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create the Container Image\n",
+ "\n",
+ "This step will likely take a few minutes."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.image import ContainerImage\n",
+ "\n",
+ "image_config = ContainerImage.image_configuration(execution_script = \"score.py\",\n",
+ " runtime = \"python\",\n",
+ " conda_file = \"myenv.yml\",\n",
+ " description = \"Emotion ONNX Runtime container\",\n",
+ " tags = {\"demo\": \"onnx\"})\n",
+ "\n",
+ "\n",
+ "image = ContainerImage.create(name = \"onnximage\",\n",
+ " # this is the model object\n",
+ " models = [model],\n",
+ " image_config = image_config,\n",
+ " workspace = ws)\n",
+ "\n",
+ "image.wait_for_creation(show_output = True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "In case you need to debug your code, the next line of code accesses the log file."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "print(image.image_build_log_uri)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "We're all done specifying what we want our virtual machine to do. Let's configure and deploy our container image.\n",
+ "\n",
+ "### Deploy the container image"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.webservice import AciWebservice\n",
+ "\n",
+ "aciconfig = AciWebservice.deploy_configuration(cpu_cores = 1, \n",
+ " memory_gb = 1, \n",
+ " tags = {'demo': 'onnx'}, \n",
+ " description = 'ONNX for emotion recognition model')"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.webservice import Webservice\n",
+ "\n",
+ "aci_service_name = 'onnx-demo-emotion'\n",
+ "print(\"Service\", aci_service_name)\n",
+ "\n",
+ "aci_service = Webservice.deploy_from_image(deployment_config = aciconfig,\n",
+ " image = image,\n",
+ " name = aci_service_name,\n",
+ " workspace = ws)\n",
+ "\n",
+ "aci_service.wait_for_deployment(True)\n",
+ "print(aci_service.state)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "The following cell will likely take a few minutes to run as well."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "if aci_service.state != 'Healthy':\n",
+ " # run this command for debugging.\n",
+ " print(aci_service.get_logs())\n",
+ "\n",
+ " # If your deployment fails, make sure to delete your aci_service before trying again!\n",
+ " # aci_service.delete()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Success!\n",
+ "\n",
+ "If you've made it this far, you've deployed a working VM with a facial emotion recognition model running in the cloud using Azure ML. Congratulations!\n",
+ "\n",
+ "Let's see how well our model deals with our test images."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Testing and Evaluation\n",
+ "\n",
+ "### Useful Helper Functions\n",
+ "\n",
+ "We preprocess and postprocess our data (see score.py file) using the helper functions specified in the [ONNX FER+ Model page in the Model Zoo repository](https://github.com/onnx/models/tree/master/emotion_ferplus)."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "def emotion_map(classes, N=1):\n",
+ " \"\"\"Take the most probable labels (output of postprocess) and returns the \n",
+ " top N emotional labels that fit the picture.\"\"\"\n",
+ " \n",
+ " emotion_table = {'neutral':0, 'happiness':1, 'surprise':2, 'sadness':3, \n",
+ " 'anger':4, 'disgust':5, 'fear':6, 'contempt':7}\n",
+ " \n",
+ " emotion_keys = list(emotion_table.keys())\n",
+ " emotions = []\n",
+ " for i in range(N):\n",
+ " emotions.append(emotion_keys[classes[i]])\n",
+ " return emotions\n",
+ "\n",
+ "def softmax(x):\n",
+ " \"\"\"Compute softmax values (probabilities from 0 to 1) for each possible label.\"\"\"\n",
+ " x = x.reshape(-1)\n",
+ " e_x = np.exp(x - np.max(x))\n",
+ " return e_x / e_x.sum(axis=0)\n",
+ "\n",
+ "def postprocess(scores):\n",
+ " \"\"\"This function takes the scores generated by the network and \n",
+ " returns the class IDs in decreasing order of probability.\"\"\"\n",
+ " prob = softmax(scores)\n",
+ " prob = np.squeeze(prob)\n",
+ " classes = np.argsort(prob)[::-1]\n",
+ " return classes"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Load Test Data\n",
+ "\n",
+ "These are already in your directory from your ONNX model download (from the model zoo).\n",
+ "\n",
+ "Notice that our Model Zoo files have a .pb extension. This is because they are [protobuf files (Protocol Buffers)](https://developers.google.com/protocol-buffers/docs/pythontutorial), so we need to read in our data through our ONNX TensorProto reader into a format we can work with, like numerical arrays."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# to manipulate our arrays\n",
+ "import numpy as np \n",
+ "\n",
+ "# read in test data protobuf files included with the model\n",
+ "import onnx\n",
+ "from onnx import numpy_helper\n",
+ "\n",
+ "# to use parsers to read in our model/data\n",
+ "import json\n",
+ "import os\n",
+ "\n",
+ "test_inputs = []\n",
+ "test_outputs = []\n",
+ "\n",
+ "# read in 3 testing images from .pb files\n",
+ "test_data_size = 3\n",
+ "\n",
+ "for i in np.arange(test_data_size):\n",
+ " input_test_data = os.path.join(model_dir, 'test_data_set_{0}'.format(i), 'input_0.pb')\n",
+ " output_test_data = os.path.join(model_dir, 'test_data_set_{0}'.format(i), 'output_0.pb')\n",
+ " \n",
+ " # convert protobuf tensors to np arrays using the TensorProto reader from ONNX\n",
+ " tensor = onnx.TensorProto()\n",
+ " with open(input_test_data, 'rb') as f:\n",
+ " tensor.ParseFromString(f.read())\n",
+ " \n",
+ " input_data = numpy_helper.to_array(tensor)\n",
+ " test_inputs.append(input_data)\n",
+ " \n",
+ " with open(output_test_data, 'rb') as f:\n",
+ " tensor.ParseFromString(f.read())\n",
+ " \n",
+ " output_data = numpy_helper.to_array(tensor)\n",
+ " output_processed = emotion_map(postprocess(output_data[0]))[0]\n",
+ " test_outputs.append(output_processed)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "nbpresent": {
+ "id": "c3f2f57c-7454-4d3e-b38d-b0946cf066ea"
+ }
+ },
+ "source": [
+ "### Show some sample images\n",
+ "We use `matplotlib` to plot 3 test images from the dataset."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "nbpresent": {
+ "id": "396d478b-34aa-4afa-9898-cdce8222a516"
+ }
+ },
+ "outputs": [],
+ "source": [
+ "plt.figure(figsize = (20, 20))\n",
+ "for test_image in np.arange(3):\n",
+ " test_inputs[test_image].reshape(1, 64, 64)\n",
+ " plt.subplot(1, 8, test_image+1)\n",
+ " plt.axhline('')\n",
+ " plt.axvline('')\n",
+ " plt.text(x = 10, y = -10, s = test_outputs[test_image], fontsize = 18)\n",
+ " plt.imshow(test_inputs[test_image].reshape(64, 64), cmap = plt.cm.gray)\n",
+ "plt.show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Run evaluation / prediction"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "plt.figure(figsize = (16, 6), frameon=False)\n",
+ "plt.subplot(1, 8, 1)\n",
+ "\n",
+ "plt.text(x = 0, y = -30, s = \"True Label: \", fontsize = 13, color = 'black')\n",
+ "plt.text(x = 0, y = -20, s = \"Result: \", fontsize = 13, color = 'black')\n",
+ "plt.text(x = 0, y = -10, s = \"Inference Time: \", fontsize = 13, color = 'black')\n",
+ "plt.text(x = 3, y = 14, s = \"Model Input\", fontsize = 12, color = 'black')\n",
+ "plt.text(x = 6, y = 18, s = \"(64 x 64)\", fontsize = 12, color = 'black')\n",
+ "plt.imshow(np.ones((28,28)), cmap=plt.cm.Greys) \n",
+ "\n",
+ "\n",
+ "for i in np.arange(test_data_size):\n",
+ " \n",
+ " input_data = json.dumps({'data': test_inputs[i].tolist()})\n",
+ "\n",
+ " # predict using the deployed model\n",
+ " r = json.loads(aci_service.run(input_data))\n",
+ " \n",
+ " if \"error\" in r:\n",
+ " print(r['error'])\n",
+ " break\n",
+ " \n",
+ " result = r['result'][0]\n",
+ " time_ms = np.round(r['time_in_sec'][0] * 1000, 2)\n",
+ " \n",
+ " ground_truth = test_outputs[i]\n",
+ " \n",
+ " # compare actual value vs. the predicted values:\n",
+ " plt.subplot(1, 8, i+2)\n",
+ " plt.axhline('')\n",
+ " plt.axvline('')\n",
+ "\n",
+ " # use different color for misclassified sample\n",
+ " font_color = 'red' if ground_truth != result else 'black'\n",
+ " clr_map = plt.cm.Greys if ground_truth != result else plt.cm.gray\n",
+ "\n",
+ " # ground truth labels are in blue\n",
+ " plt.text(x = 10, y = -70, s = ground_truth, fontsize = 18, color = 'blue')\n",
+ " \n",
+ " # predictions are in black if correct, red if incorrect\n",
+ " plt.text(x = 10, y = -45, s = result, fontsize = 18, color = font_color)\n",
+ " plt.text(x = 5, y = -22, s = str(time_ms) + ' ms', fontsize = 14, color = font_color)\n",
+ "\n",
+ " \n",
+ " plt.imshow(test_inputs[i].reshape(64, 64), cmap = clr_map)\n",
+ "\n",
+ "plt.show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Try classifying your own images!"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Preprocessing functions take your image and format it so it can be passed\n",
+ "# as input into our ONNX model\n",
+ "\n",
+ "import cv2\n",
+ "\n",
+ "def rgb2gray(rgb):\n",
+ " \"\"\"Convert the input image into grayscale\"\"\"\n",
+ " return np.dot(rgb[...,:3], [0.299, 0.587, 0.114])\n",
+ "\n",
+ "def resize_img(img):\n",
+ " \"\"\"Resize image to MNIST model input dimensions\"\"\"\n",
+ " img = cv2.resize(img, dsize=(64, 64), interpolation=cv2.INTER_AREA)\n",
+ " img.resize((1, 1, 64, 64))\n",
+ " return img\n",
+ "\n",
+ "def preprocess(img):\n",
+ " \"\"\"Resize input images and convert them to grayscale.\"\"\"\n",
+ " if img.shape == (64, 64):\n",
+ " img.resize((1, 1, 64, 64))\n",
+ " return img\n",
+ " \n",
+ " grayscale = rgb2gray(img)\n",
+ " processed_img = resize_img(grayscale)\n",
+ " return processed_img"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Replace the following string with your own path/test image\n",
+ "# Make sure your image is square and the dimensions are equal (i.e. 100 * 100 pixels or 28 * 28 pixels)\n",
+ "\n",
+ "# Any PNG or JPG image file should work\n",
+ "# Make sure to include the entire path with // instead of /\n",
+ "\n",
+ "# e.g. your_test_image = \"C:/Users/vinitra.swamy/Pictures/face.png\"\n",
+ "\n",
+ "your_test_image = \"\"\n",
+ "\n",
+ "import matplotlib.image as mpimg\n",
+ "\n",
+ "if your_test_image != \"\":\n",
+ " img = mpimg.imread(your_test_image)\n",
+ " plt.subplot(1,3,1)\n",
+ " plt.imshow(img, cmap = plt.cm.Greys)\n",
+ " print(\"Old Dimensions: \", img.shape)\n",
+ " img = preprocess(img)\n",
+ " print(\"New Dimensions: \", img.shape)\n",
+ "else:\n",
+ " img = None"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "if img is None:\n",
+ " print(\"Add the path for your image data.\")\n",
+ "else:\n",
+ " input_data = json.dumps({'data': img.tolist()})\n",
+ "\n",
+ " try:\n",
+ " r = json.loads(aci_service.run(input_data))\n",
+ " result = r['result'][0]\n",
+ " time_ms = np.round(r['time_in_sec'][0] * 1000, 2)\n",
+ " except Exception as e:\n",
+ " print(str(e))\n",
+ "\n",
+ " plt.figure(figsize = (16, 6))\n",
+ " plt.subplot(1,8,1)\n",
+ " plt.axhline('')\n",
+ " plt.axvline('')\n",
+ " plt.text(x = -10, y = -40, s = \"Model prediction: \", fontsize = 14)\n",
+ " plt.text(x = -10, y = -25, s = \"Inference time: \", fontsize = 14)\n",
+ " plt.text(x = 100, y = -40, s = str(result), fontsize = 14)\n",
+ " plt.text(x = 100, y = -25, s = str(time_ms) + \" ms\", fontsize = 14)\n",
+ " plt.text(x = -10, y = -10, s = \"Model Input image: \", fontsize = 14)\n",
+ " plt.imshow(img.reshape((64, 64)), cmap = plt.cm.gray) \n",
+ " "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# remember to delete your service after you are done using it!\n",
+ "\n",
+ "# aci_service.delete()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Conclusion\n",
+ "\n",
+ "Congratulations!\n",
+ "\n",
+ "In this tutorial, you have:\n",
+ "- familiarized yourself with ONNX Runtime inference and the pretrained models in the ONNX model zoo\n",
+ "- understood a state-of-the-art convolutional neural net image classification model (FER+ in ONNX) and deployed it in the Azure ML cloud\n",
+ "- ensured that your deep learning model is working perfectly (in the cloud) on test data, and checked it against some of your own!\n",
+ "\n",
+ "Next steps:\n",
+ "- If you have not already, check out another interesting ONNX/AML application that lets you set up a state-of-the-art [handwritten image classification model (MNIST)](https://github.com/Azure/MachineLearningNotebooks/tree/master/onnx/onnx-inference-mnist.ipynb) in the cloud! This tutorial deploys a pre-trained ONNX Computer Vision model for handwritten digit classification in an Azure ML virtual machine.\n",
+ "- Keep an eye out for an updated version of this tutorial that uses ONNX Runtime GPU.\n",
+ "- Contribute to our [open source ONNX repository on github](http://github.com/onnx/onnx) and/or add to our [ONNX model zoo](http://github.com/onnx/models)"
+ ]
}
- },
- "source": [
- "### Show some sample images\n",
- "We use `matplotlib` to plot 3 test images from the dataset."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "nbpresent": {
- "id": "396d478b-34aa-4afa-9898-cdce8222a516"
- }
- },
- "outputs": [],
- "source": [
- "plt.figure(figsize = (20, 20))\n",
- "for test_image in np.arange(3):\n",
- " test_inputs[test_image].reshape(1, 64, 64)\n",
- " plt.subplot(1, 8, test_image+1)\n",
- " plt.axhline('')\n",
- " plt.axvline('')\n",
- " plt.text(x = 10, y = -10, s = test_outputs[test_image], fontsize = 18)\n",
- " plt.imshow(test_inputs[test_image].reshape(64, 64), cmap = plt.cm.gray)\n",
- "plt.show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Run evaluation / prediction"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "plt.figure(figsize = (16, 6), frameon=False)\n",
- "plt.subplot(1, 8, 1)\n",
- "\n",
- "plt.text(x = 0, y = -30, s = \"True Label: \", fontsize = 13, color = 'black')\n",
- "plt.text(x = 0, y = -20, s = \"Result: \", fontsize = 13, color = 'black')\n",
- "plt.text(x = 0, y = -10, s = \"Inference Time: \", fontsize = 13, color = 'black')\n",
- "plt.text(x = 3, y = 14, s = \"Model Input\", fontsize = 12, color = 'black')\n",
- "plt.text(x = 6, y = 18, s = \"(64 x 64)\", fontsize = 12, color = 'black')\n",
- "plt.imshow(np.ones((28,28)), cmap=plt.cm.Greys) \n",
- "\n",
- "\n",
- "for i in np.arange(test_data_size):\n",
- " \n",
- " input_data = json.dumps({'data': test_inputs[i].tolist()})\n",
- "\n",
- " # predict using the deployed model\n",
- " r = json.loads(aci_service.run(input_data))\n",
- " \n",
- " if \"error\" in r:\n",
- " print(r['error'])\n",
- " break\n",
- " \n",
- " result = r['result'][0]\n",
- " time_ms = np.round(r['time_in_sec'][0] * 1000, 2)\n",
- " \n",
- " ground_truth = test_outputs[i]\n",
- " \n",
- " # compare actual value vs. the predicted values:\n",
- " plt.subplot(1, 8, i+2)\n",
- " plt.axhline('')\n",
- " plt.axvline('')\n",
- "\n",
- " # use different color for misclassified sample\n",
- " font_color = 'red' if ground_truth != result else 'black'\n",
- " clr_map = plt.cm.Greys if ground_truth != result else plt.cm.gray\n",
- "\n",
- " # ground truth labels are in blue\n",
- " plt.text(x = 10, y = -70, s = ground_truth, fontsize = 18, color = 'blue')\n",
- " \n",
- " # predictions are in black if correct, red if incorrect\n",
- " plt.text(x = 10, y = -45, s = result, fontsize = 18, color = font_color)\n",
- " plt.text(x = 5, y = -22, s = str(time_ms) + ' ms', fontsize = 14, color = font_color)\n",
- "\n",
- " \n",
- " plt.imshow(test_inputs[i].reshape(64, 64), cmap = clr_map)\n",
- "\n",
- "plt.show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Try classifying your own images!"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Preprocessing functions take your image and format it so it can be passed\n",
- "# as input into our ONNX model\n",
- "\n",
- "import cv2\n",
- "\n",
- "def rgb2gray(rgb):\n",
- " \"\"\"Convert the input image into grayscale\"\"\"\n",
- " return np.dot(rgb[...,:3], [0.299, 0.587, 0.114])\n",
- "\n",
- "def resize_img(img):\n",
- " \"\"\"Resize image to MNIST model input dimensions\"\"\"\n",
- " img = cv2.resize(img, dsize=(64, 64), interpolation=cv2.INTER_AREA)\n",
- " img.resize((1, 1, 64, 64))\n",
- " return img\n",
- "\n",
- "def preprocess(img):\n",
- " \"\"\"Resize input images and convert them to grayscale.\"\"\"\n",
- " if img.shape == (64, 64):\n",
- " img.resize((1, 1, 64, 64))\n",
- " return img\n",
- " \n",
- " grayscale = rgb2gray(img)\n",
- " processed_img = resize_img(grayscale)\n",
- " return processed_img"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Replace the following string with your own path/test image\n",
- "# Make sure your image is square and the dimensions are equal (i.e. 100 * 100 pixels or 28 * 28 pixels)\n",
- "\n",
- "# Any PNG or JPG image file should work\n",
- "# Make sure to include the entire path with // instead of /\n",
- "\n",
- "# e.g. your_test_image = \"C:/Users/vinitra.swamy/Pictures/face.png\"\n",
- "\n",
- "your_test_image = \"\"\n",
- "\n",
- "import matplotlib.image as mpimg\n",
- "\n",
- "if your_test_image != \"\":\n",
- " img = mpimg.imread(your_test_image)\n",
- " plt.subplot(1,3,1)\n",
- " plt.imshow(img, cmap = plt.cm.Greys)\n",
- " print(\"Old Dimensions: \", img.shape)\n",
- " img = preprocess(img)\n",
- " print(\"New Dimensions: \", img.shape)\n",
- "else:\n",
- " img = None"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "if img is None:\n",
- " print(\"Add the path for your image data.\")\n",
- "else:\n",
- " input_data = json.dumps({'data': img.tolist()})\n",
- "\n",
- " try:\n",
- " r = json.loads(aci_service.run(input_data))\n",
- " result = r['result'][0]\n",
- " time_ms = np.round(r['time_in_sec'][0] * 1000, 2)\n",
- " except Exception as e:\n",
- " print(str(e))\n",
- "\n",
- " plt.figure(figsize = (16, 6))\n",
- " plt.subplot(1,8,1)\n",
- " plt.axhline('')\n",
- " plt.axvline('')\n",
- " plt.text(x = -10, y = -40, s = \"Model prediction: \", fontsize = 14)\n",
- " plt.text(x = -10, y = -25, s = \"Inference time: \", fontsize = 14)\n",
- " plt.text(x = 100, y = -40, s = str(result), fontsize = 14)\n",
- " plt.text(x = 100, y = -25, s = str(time_ms) + \" ms\", fontsize = 14)\n",
- " plt.text(x = -10, y = -10, s = \"Model Input image: \", fontsize = 14)\n",
- " plt.imshow(img.reshape((64, 64)), cmap = plt.cm.gray) \n",
- " "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# remember to delete your service after you are done using it!\n",
- "\n",
- "# aci_service.delete()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Conclusion\n",
- "\n",
- "Congratulations!\n",
- "\n",
- "In this tutorial, you have:\n",
- "- familiarized yourself with ONNX Runtime inference and the pretrained models in the ONNX model zoo\n",
- "- understood a state-of-the-art convolutional neural net image classification model (FER+ in ONNX) and deployed it in the Azure ML cloud\n",
- "- ensured that your deep learning model is working perfectly (in the cloud) on test data, and checked it against some of your own!\n",
- "\n",
- "Next steps:\n",
- "- If you have not already, check out another interesting ONNX/AML application that lets you set up a state-of-the-art [handwritten image classification model (MNIST)](https://github.com/Azure/MachineLearningNotebooks/tree/master/onnx/onnx-inference-mnist.ipynb) in the cloud! This tutorial deploys a pre-trained ONNX Computer Vision model for handwritten digit classification in an Azure ML virtual machine.\n",
- "- Keep an eye out for an updated version of this tutorial that uses ONNX Runtime GPU.\n",
- "- Contribute to our [open source ONNX repository on github](http://github.com/onnx/onnx) and/or add to our [ONNX model zoo](http://github.com/onnx/models)"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "viswamy"
- }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "viswamy"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.6"
+ },
+ "msauthor": "vinitra.swamy"
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- },
- "msauthor": "vinitra.swamy"
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/deployment/onnx/onnx-inference-mnist-deploy.ipynb b/how-to-use-azureml/deployment/onnx/onnx-inference-mnist-deploy.ipynb
index f2ec51a4..68e7f80d 100644
--- a/how-to-use-azureml/deployment/onnx/onnx-inference-mnist-deploy.ipynb
+++ b/how-to-use-azureml/deployment/onnx/onnx-inference-mnist-deploy.ipynb
@@ -1,812 +1,812 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved. \n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Handwritten Digit Classification (MNIST) using ONNX Runtime on Azure ML\n",
- "\n",
- "This example shows how to deploy an image classification neural network using the Modified National Institute of Standards and Technology ([MNIST](http://yann.lecun.com/exdb/mnist/)) dataset and Open Neural Network eXchange format ([ONNX](http://aka.ms/onnxdocarticle)) on the Azure Machine Learning platform. MNIST is a popular dataset consisting of 70,000 grayscale images. Each image is a handwritten digit of 28x28 pixels, representing number from 0 to 9. This tutorial will show you how to deploy a MNIST model from the [ONNX model zoo](https://github.com/onnx/models), use it to make predictions using ONNX Runtime Inference, and deploy it as a web service in Azure.\n",
- "\n",
- "Throughout this tutorial, we will be referring to ONNX, a neural network exchange format used to represent deep learning models. With ONNX, AI developers can more easily move models between state-of-the-art tools (CNTK, PyTorch, Caffe, MXNet, TensorFlow) and choose the combination that is best for them. ONNX is developed and supported by a community of partners including Microsoft AI, Facebook, and Amazon. For more information, explore the [ONNX website](http://onnx.ai) and [open source files](https://github.com/onnx).\n",
- "\n",
- "[ONNX Runtime](https://aka.ms/onnxruntime-python) is the runtime engine that enables evaluation of trained machine learning (Traditional ML and Deep Learning) models with high performance and low resource utilization.\n",
- "\n",
- "#### Tutorial Objectives:\n",
- "\n",
- "- Describe the MNIST dataset and pretrained Convolutional Neural Net ONNX model, stored in the ONNX model zoo.\n",
- "- Deploy and run the pretrained MNIST ONNX model on an Azure Machine Learning instance\n",
- "- Predict labels for test set data points in the cloud using ONNX Runtime and Azure ML"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Prerequisites\n",
- "\n",
- "### 1. Install Azure ML SDK and create a new workspace\n",
- "Please follow [Azure ML configuration notebook](https://github.com/Azure/MachineLearningNotebooks/blob/master/00.configuration.ipynb) to set up your environment.\n",
- "\n",
- "### 2. Install additional packages needed for this tutorial notebook\n",
- "You need to install the popular plotting library `matplotlib`, the image manipulation library `opencv`, and the `onnx` library in the conda environment where Azure Maching Learning SDK is installed. \n",
- "\n",
- "```sh\n",
- "(myenv) $ pip install matplotlib onnx opencv-python\n",
- "```\n",
- "\n",
- "**Debugging tip**: Make sure that you run the \"jupyter notebook\" command to launch this notebook after activating your virtual environment. Choose the respective Python kernel for your new virtual environment using the `Kernel > Change Kernel` menu above. If you have completed the steps correctly, the upper right corner of your screen should state `Python [conda env:myenv]` instead of `Python [default]`.\n",
- "\n",
- "### 3. Download sample data and pre-trained ONNX model from ONNX Model Zoo.\n",
- "\n",
- "In the following lines of code, we download [the trained ONNX MNIST model and corresponding test data](https://github.com/onnx/models/tree/master/mnist) and place them in the same folder as this tutorial notebook. For more information about the MNIST dataset, please visit [Yan LeCun's website](http://yann.lecun.com/exdb/mnist/)."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# urllib is a built-in Python library to download files from URLs\n",
- "\n",
- "# Objective: retrieve the latest version of the ONNX MNIST model files from the\n",
- "# ONNX Model Zoo and save it in the same folder as this tutorial\n",
- "\n",
- "import urllib.request\n",
- "\n",
- "onnx_model_url = \"https://www.cntk.ai/OnnxModels/mnist/opset_7/mnist.tar.gz\"\n",
- "\n",
- "urllib.request.urlretrieve(onnx_model_url, filename=\"mnist.tar.gz\")"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# the ! magic command tells our jupyter notebook kernel to run the following line of \n",
- "# code from the command line instead of the notebook kernel\n",
- "\n",
- "# We use tar and xvcf to unzip the files we just retrieved from the ONNX model zoo\n",
- "\n",
- "!tar xvzf mnist.tar.gz"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Deploy a VM with your ONNX model in the Cloud\n",
- "\n",
- "### Load Azure ML workspace\n",
- "\n",
- "We begin by instantiating a workspace object from the existing workspace created earlier in the configuration notebook."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Check core SDK version number\n",
- "import azureml.core\n",
- "\n",
- "print(\"SDK version:\", azureml.core.VERSION)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core import Workspace\n",
- "\n",
- "ws = Workspace.from_config()\n",
- "print(ws.name, ws.resource_group, ws.location, sep = '\\n')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Registering your model with Azure ML"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "model_dir = \"mnist\" # replace this with the location of your model files\n",
- "\n",
- "# leave as is if it's in the same folder as this notebook"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.model import Model\n",
- "\n",
- "model = Model.register(workspace = ws,\n",
- " model_path = model_dir + \"/\" + \"model.onnx\",\n",
- " model_name = \"mnist_1\",\n",
- " tags = {\"onnx\": \"demo\"},\n",
- " description = \"MNIST image classification CNN from ONNX Model Zoo\",)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Optional: Displaying your registered models\n",
- "\n",
- "This step is not required, so feel free to skip it."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "models = ws.models\n",
- "for name, m in models.items():\n",
- " print(\"Name:\", name,\"\\tVersion:\", m.version, \"\\tDescription:\", m.description, m.tags)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {
- "nbpresent": {
- "id": "c3f2f57c-7454-4d3e-b38d-b0946cf066ea"
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved. \n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Handwritten Digit Classification (MNIST) using ONNX Runtime on Azure ML\n",
+ "\n",
+ "This example shows how to deploy an image classification neural network using the Modified National Institute of Standards and Technology ([MNIST](http://yann.lecun.com/exdb/mnist/)) dataset and Open Neural Network eXchange format ([ONNX](http://aka.ms/onnxdocarticle)) on the Azure Machine Learning platform. MNIST is a popular dataset consisting of 70,000 grayscale images. Each image is a handwritten digit of 28x28 pixels, representing number from 0 to 9. This tutorial will show you how to deploy a MNIST model from the [ONNX model zoo](https://github.com/onnx/models), use it to make predictions using ONNX Runtime Inference, and deploy it as a web service in Azure.\n",
+ "\n",
+ "Throughout this tutorial, we will be referring to ONNX, a neural network exchange format used to represent deep learning models. With ONNX, AI developers can more easily move models between state-of-the-art tools (CNTK, PyTorch, Caffe, MXNet, TensorFlow) and choose the combination that is best for them. ONNX is developed and supported by a community of partners including Microsoft AI, Facebook, and Amazon. For more information, explore the [ONNX website](http://onnx.ai) and [open source files](https://github.com/onnx).\n",
+ "\n",
+ "[ONNX Runtime](https://aka.ms/onnxruntime-python) is the runtime engine that enables evaluation of trained machine learning (Traditional ML and Deep Learning) models with high performance and low resource utilization.\n",
+ "\n",
+ "#### Tutorial Objectives:\n",
+ "\n",
+ "- Describe the MNIST dataset and pretrained Convolutional Neural Net ONNX model, stored in the ONNX model zoo.\n",
+ "- Deploy and run the pretrained MNIST ONNX model on an Azure Machine Learning instance\n",
+ "- Predict labels for test set data points in the cloud using ONNX Runtime and Azure ML"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Prerequisites\n",
+ "\n",
+ "### 1. Install Azure ML SDK and create a new workspace\n",
+ "Please follow [Azure ML configuration notebook](../../../configuration.ipynb) to set up your environment.\n",
+ "\n",
+ "### 2. Install additional packages needed for this tutorial notebook\n",
+ "You need to install the popular plotting library `matplotlib`, the image manipulation library `opencv`, and the `onnx` library in the conda environment where Azure Maching Learning SDK is installed. \n",
+ "\n",
+ "```sh\n",
+ "(myenv) $ pip install matplotlib onnx opencv-python\n",
+ "```\n",
+ "\n",
+ "**Debugging tip**: Make sure that you run the \"jupyter notebook\" command to launch this notebook after activating your virtual environment. Choose the respective Python kernel for your new virtual environment using the `Kernel > Change Kernel` menu above. If you have completed the steps correctly, the upper right corner of your screen should state `Python [conda env:myenv]` instead of `Python [default]`.\n",
+ "\n",
+ "### 3. Download sample data and pre-trained ONNX model from ONNX Model Zoo.\n",
+ "\n",
+ "In the following lines of code, we download [the trained ONNX MNIST model and corresponding test data](https://github.com/onnx/models/tree/master/mnist) and place them in the same folder as this tutorial notebook. For more information about the MNIST dataset, please visit [Yan LeCun's website](http://yann.lecun.com/exdb/mnist/)."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# urllib is a built-in Python library to download files from URLs\n",
+ "\n",
+ "# Objective: retrieve the latest version of the ONNX MNIST model files from the\n",
+ "# ONNX Model Zoo and save it in the same folder as this tutorial\n",
+ "\n",
+ "import urllib.request\n",
+ "\n",
+ "onnx_model_url = \"https://www.cntk.ai/OnnxModels/mnist/opset_7/mnist.tar.gz\"\n",
+ "\n",
+ "urllib.request.urlretrieve(onnx_model_url, filename=\"mnist.tar.gz\")"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# the ! magic command tells our jupyter notebook kernel to run the following line of \n",
+ "# code from the command line instead of the notebook kernel\n",
+ "\n",
+ "# We use tar and xvcf to unzip the files we just retrieved from the ONNX model zoo\n",
+ "\n",
+ "!tar xvzf mnist.tar.gz"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Deploy a VM with your ONNX model in the Cloud\n",
+ "\n",
+ "### Load Azure ML workspace\n",
+ "\n",
+ "We begin by instantiating a workspace object from the existing workspace created earlier in the configuration notebook."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Check core SDK version number\n",
+ "import azureml.core\n",
+ "\n",
+ "print(\"SDK version:\", azureml.core.VERSION)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core import Workspace\n",
+ "\n",
+ "ws = Workspace.from_config()\n",
+ "print(ws.name, ws.resource_group, ws.location, sep = '\\n')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Registering your model with Azure ML"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "model_dir = \"mnist\" # replace this with the location of your model files\n",
+ "\n",
+ "# leave as is if it's in the same folder as this notebook"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.model import Model\n",
+ "\n",
+ "model = Model.register(workspace = ws,\n",
+ " model_path = model_dir + \"/\" + \"model.onnx\",\n",
+ " model_name = \"mnist_1\",\n",
+ " tags = {\"onnx\": \"demo\"},\n",
+ " description = \"MNIST image classification CNN from ONNX Model Zoo\",)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Optional: Displaying your registered models\n",
+ "\n",
+ "This step is not required, so feel free to skip it."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "models = ws.models\n",
+ "for name, m in models.items():\n",
+ " print(\"Name:\", name,\"\\tVersion:\", m.version, \"\\tDescription:\", m.description, m.tags)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "nbpresent": {
+ "id": "c3f2f57c-7454-4d3e-b38d-b0946cf066ea"
+ }
+ },
+ "source": [
+ "### ONNX MNIST Model Methodology\n",
+ "\n",
+ "The image classification model we are using is pre-trained using Microsoft's deep learning cognitive toolkit, [CNTK](https://github.com/Microsoft/CNTK), from the [ONNX model zoo](http://github.com/onnx/models). The model zoo has many other models that can be deployed on cloud providers like AzureML without any additional training. To ensure that our cloud deployed model works, we use testing data from the famous MNIST data set, provided as part of the [trained MNIST model](https://github.com/onnx/models/tree/master/mnist) in the ONNX model zoo.\n",
+ "\n",
+ "***Input: Handwritten Images from MNIST Dataset***\n",
+ "\n",
+ "***Task: Classify each MNIST image into an appropriate digit***\n",
+ "\n",
+ "***Output: Digit prediction for input image***\n",
+ "\n",
+ "Run the cell below to look at some of the sample images from the MNIST dataset that we used to train this ONNX model. Remember, once the application is deployed in Azure ML, you can use your own images as input for the model to classify!"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# for images and plots in this notebook\n",
+ "import matplotlib.pyplot as plt \n",
+ "from IPython.display import Image\n",
+ "\n",
+ "# display images inline\n",
+ "%matplotlib inline"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "Image(url=\"http://3.bp.blogspot.com/_UpN7DfJA0j4/TJtUBWPk0SI/AAAAAAAAABY/oWPMtmqJn3k/s1600/mnist_originals.png\", width=200, height=200)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Specify our Score and Environment Files"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "We are now going to deploy our ONNX Model on AML with inference in ONNX Runtime. We begin by writing a score.py file, which will help us run the model in our Azure ML virtual machine (VM), and then specify our environment by writing a yml file. You will also notice that we import the onnxruntime library to do runtime inference on our ONNX models (passing in input and evaluating out model's predicted output). More information on the API and commands can be found in the [ONNX Runtime documentation](https://aka.ms/onnxruntime).\n",
+ "\n",
+ "### Write Score File\n",
+ "\n",
+ "A score file is what tells our Azure cloud service what to do. After initializing our model using azureml.core.model, we start an ONNX Runtime inference session to evaluate the data passed in on our function calls."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%writefile score.py\n",
+ "import json\n",
+ "import numpy as np\n",
+ "import onnxruntime\n",
+ "import sys\n",
+ "import os\n",
+ "from azureml.core.model import Model\n",
+ "import time\n",
+ "\n",
+ "\n",
+ "def init():\n",
+ " global session, input_name, output_name\n",
+ " model = Model.get_model_path(model_name = 'mnist_1')\n",
+ " session = onnxruntime.InferenceSession(model, None)\n",
+ " input_name = session.get_inputs()[0].name\n",
+ " output_name = session.get_outputs()[0].name \n",
+ " \n",
+ "\n",
+ "def preprocess(input_data_json):\n",
+ " # convert the JSON data into the tensor input\n",
+ " return np.array(json.loads(input_data_json)['data']).astype('float32')\n",
+ "\n",
+ "def postprocess(result):\n",
+ " # We use argmax to pick the highest confidence label\n",
+ " return int(np.argmax(np.array(result).squeeze(), axis=0))\n",
+ " \n",
+ "def run(input_data):\n",
+ "\n",
+ " try:\n",
+ " # load in our data, convert to readable format\n",
+ " data = preprocess(input_data)\n",
+ " \n",
+ " # start timer\n",
+ " start = time.time()\n",
+ " \n",
+ " r = session.run([output_name], {input_name: data})\n",
+ " \n",
+ " #end timer\n",
+ " end = time.time()\n",
+ " \n",
+ " result = postprocess(r)\n",
+ " result_dict = {\"result\": result,\n",
+ " \"time_in_sec\": end - start}\n",
+ " except Exception as e:\n",
+ " result_dict = {\"error\": str(e)}\n",
+ " \n",
+ " return result_dict\n",
+ "\n",
+ "def choose_class(result_prob):\n",
+ " \"\"\"We use argmax to determine the right label to choose from our output\"\"\"\n",
+ " return int(np.argmax(result_prob, axis=0))"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Write Environment File\n",
+ "\n",
+ "This step creates a YAML environment file that specifies which dependencies we would like to see in our Linux Virtual Machine."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.conda_dependencies import CondaDependencies \n",
+ "\n",
+ "myenv = CondaDependencies.create(pip_packages=[\"numpy\", \"onnxruntime\", \"azureml-core\"])\n",
+ "\n",
+ "with open(\"myenv.yml\",\"w\") as f:\n",
+ " f.write(myenv.serialize_to_string())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create the Container Image\n",
+ "This step will likely take a few minutes."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.image import ContainerImage\n",
+ "\n",
+ "image_config = ContainerImage.image_configuration(execution_script = \"score.py\",\n",
+ " runtime = \"python\",\n",
+ " conda_file = \"myenv.yml\",\n",
+ " description = \"MNIST ONNX Runtime container\",\n",
+ " tags = {\"demo\": \"onnx\"}) \n",
+ "\n",
+ "\n",
+ "image = ContainerImage.create(name = \"onnximage\",\n",
+ " # this is the model object\n",
+ " models = [model],\n",
+ " image_config = image_config,\n",
+ " workspace = ws)\n",
+ "\n",
+ "image.wait_for_creation(show_output = True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "In case you need to debug your code, the next line of code accesses the log file."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "print(image.image_build_log_uri)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "We're all done specifying what we want our virtual machine to do. Let's configure and deploy our container image.\n",
+ "\n",
+ "### Deploy the container image"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.webservice import AciWebservice\n",
+ "\n",
+ "aciconfig = AciWebservice.deploy_configuration(cpu_cores = 1, \n",
+ " memory_gb = 1, \n",
+ " tags = {'demo': 'onnx'}, \n",
+ " description = 'ONNX for mnist model')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "The following cell will likely take a few minutes to run as well."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.webservice import Webservice\n",
+ "\n",
+ "aci_service_name = 'onnx-demo-mnist'\n",
+ "print(\"Service\", aci_service_name)\n",
+ "\n",
+ "aci_service = Webservice.deploy_from_image(deployment_config = aciconfig,\n",
+ " image = image,\n",
+ " name = aci_service_name,\n",
+ " workspace = ws)\n",
+ "\n",
+ "aci_service.wait_for_deployment(True)\n",
+ "print(aci_service.state)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "if aci_service.state != 'Healthy':\n",
+ " # run this command for debugging.\n",
+ " print(aci_service.get_logs())\n",
+ "\n",
+ " # If your deployment fails, make sure to delete your aci_service or rename your service before trying again!\n",
+ " # aci_service.delete()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Success!\n",
+ "\n",
+ "If you've made it this far, you've deployed a working VM with a handwritten digit classifier running in the cloud using Azure ML. Congratulations!\n",
+ "\n",
+ "You can get the URL for the webservice with the code below. Let's now see how well our model deals with our test images."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "print(aci_service.scoring_uri)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Testing and Evaluation\n",
+ "\n",
+ "### Load Test Data\n",
+ "\n",
+ "These are already in your directory from your ONNX model download (from the model zoo).\n",
+ "\n",
+ "Notice that our Model Zoo files have a .pb extension. This is because they are [protobuf files (Protocol Buffers)](https://developers.google.com/protocol-buffers/docs/pythontutorial), so we need to read in our data through our ONNX TensorProto reader into a format we can work with, like numerical arrays."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# to manipulate our arrays\n",
+ "import numpy as np \n",
+ "\n",
+ "# read in test data protobuf files included with the model\n",
+ "import onnx\n",
+ "from onnx import numpy_helper\n",
+ "\n",
+ "# to use parsers to read in our model/data\n",
+ "import json\n",
+ "import os\n",
+ "\n",
+ "test_inputs = []\n",
+ "test_outputs = []\n",
+ "\n",
+ "# read in 3 testing images from .pb files\n",
+ "test_data_size = 3\n",
+ "\n",
+ "for i in np.arange(test_data_size):\n",
+ " input_test_data = os.path.join(model_dir, 'test_data_set_{0}'.format(i), 'input_0.pb')\n",
+ " output_test_data = os.path.join(model_dir, 'test_data_set_{0}'.format(i), 'output_0.pb')\n",
+ " \n",
+ " # convert protobuf tensors to np arrays using the TensorProto reader from ONNX\n",
+ " tensor = onnx.TensorProto()\n",
+ " with open(input_test_data, 'rb') as f:\n",
+ " tensor.ParseFromString(f.read())\n",
+ " \n",
+ " input_data = numpy_helper.to_array(tensor)\n",
+ " test_inputs.append(input_data)\n",
+ " \n",
+ " with open(output_test_data, 'rb') as f:\n",
+ " tensor.ParseFromString(f.read())\n",
+ " \n",
+ " output_data = numpy_helper.to_array(tensor)\n",
+ " test_outputs.append(output_data)\n",
+ " \n",
+ "if len(test_inputs) == test_data_size:\n",
+ " print('Test data loaded successfully.')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "nbpresent": {
+ "id": "c3f2f57c-7454-4d3e-b38d-b0946cf066ea"
+ }
+ },
+ "source": [
+ "### Show some sample images\n",
+ "We use `matplotlib` to plot 3 test images from the dataset."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "nbpresent": {
+ "id": "396d478b-34aa-4afa-9898-cdce8222a516"
+ }
+ },
+ "outputs": [],
+ "source": [
+ "plt.figure(figsize = (16, 6))\n",
+ "for test_image in np.arange(3):\n",
+ " plt.subplot(1, 15, test_image+1)\n",
+ " plt.axhline('')\n",
+ " plt.axvline('')\n",
+ " plt.imshow(test_inputs[test_image].reshape(28, 28), cmap = plt.cm.Greys)\n",
+ "plt.show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Run evaluation / prediction"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "plt.figure(figsize = (16, 6), frameon=False)\n",
+ "plt.subplot(1, 8, 1)\n",
+ "\n",
+ "plt.text(x = 0, y = -30, s = \"True Label: \", fontsize = 13, color = 'black')\n",
+ "plt.text(x = 0, y = -20, s = \"Result: \", fontsize = 13, color = 'black')\n",
+ "plt.text(x = 0, y = -10, s = \"Inference Time: \", fontsize = 13, color = 'black')\n",
+ "plt.text(x = 3, y = 14, s = \"Model Input\", fontsize = 12, color = 'black')\n",
+ "plt.text(x = 6, y = 18, s = \"(28 x 28)\", fontsize = 12, color = 'black')\n",
+ "plt.imshow(np.ones((28,28)), cmap=plt.cm.Greys) \n",
+ "\n",
+ "\n",
+ "for i in np.arange(test_data_size):\n",
+ " \n",
+ " input_data = json.dumps({'data': test_inputs[i].tolist()})\n",
+ " \n",
+ " # predict using the deployed model\n",
+ " r = aci_service.run(input_data)\n",
+ " \n",
+ " if \"error\" in r:\n",
+ " print(r['error'])\n",
+ " break\n",
+ " \n",
+ " result = r['result']\n",
+ " time_ms = np.round(r['time_in_sec'] * 1000, 2)\n",
+ " \n",
+ " ground_truth = int(np.argmax(test_outputs[i]))\n",
+ " \n",
+ " # compare actual value vs. the predicted values:\n",
+ " plt.subplot(1, 8, i+2)\n",
+ " plt.axhline('')\n",
+ " plt.axvline('')\n",
+ "\n",
+ " # use different color for misclassified sample\n",
+ " font_color = 'red' if ground_truth != result else 'black'\n",
+ " clr_map = plt.cm.gray if ground_truth != result else plt.cm.Greys\n",
+ "\n",
+ " # ground truth labels are in blue\n",
+ " plt.text(x = 10, y = -30, s = ground_truth, fontsize = 18, color = 'blue')\n",
+ " \n",
+ " # predictions are in black if correct, red if incorrect\n",
+ " plt.text(x = 10, y = -20, s = result, fontsize = 18, color = font_color)\n",
+ " plt.text(x = 5, y = -10, s = str(time_ms) + ' ms', fontsize = 14, color = font_color)\n",
+ "\n",
+ " \n",
+ " plt.imshow(test_inputs[i].reshape(28, 28), cmap = clr_map)\n",
+ "\n",
+ "plt.show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Try classifying your own images!\n",
+ "\n",
+ "Create your own handwritten image and pass it into the model."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Preprocessing functions take your image and format it so it can be passed\n",
+ "# as input into our ONNX model\n",
+ "\n",
+ "import cv2\n",
+ "\n",
+ "def rgb2gray(rgb):\n",
+ " \"\"\"Convert the input image into grayscale\"\"\"\n",
+ " return np.dot(rgb[...,:3], [0.299, 0.587, 0.114])\n",
+ "\n",
+ "def resize_img(img):\n",
+ " \"\"\"Resize image to MNIST model input dimensions\"\"\"\n",
+ " img = cv2.resize(img, dsize=(28, 28), interpolation=cv2.INTER_AREA)\n",
+ " img.resize((1, 1, 28, 28))\n",
+ " return img\n",
+ "\n",
+ "def preprocess(img):\n",
+ " \"\"\"Resize input images and convert them to grayscale.\"\"\"\n",
+ " if img.shape == (28, 28):\n",
+ " img.resize((1, 1, 28, 28))\n",
+ " return img\n",
+ " \n",
+ " grayscale = rgb2gray(img)\n",
+ " processed_img = resize_img(grayscale)\n",
+ " return processed_img"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Replace this string with your own path/test image\n",
+ "# Make sure your image is square and the dimensions are equal (i.e. 100 * 100 pixels or 28 * 28 pixels)\n",
+ "\n",
+ "# Any PNG or JPG image file should work\n",
+ "\n",
+ "your_test_image = \"\"\n",
+ "\n",
+ "# e.g. your_test_image = \"C:/Users/vinitra.swamy/Pictures/handwritten_digit.png\"\n",
+ "\n",
+ "import matplotlib.image as mpimg\n",
+ "\n",
+ "if your_test_image != \"\":\n",
+ " img = mpimg.imread(your_test_image)\n",
+ " plt.subplot(1,3,1)\n",
+ " plt.imshow(img, cmap = plt.cm.Greys)\n",
+ " print(\"Old Dimensions: \", img.shape)\n",
+ " img = preprocess(img)\n",
+ " print(\"New Dimensions: \", img.shape)\n",
+ "else:\n",
+ " img = None"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "if img is None:\n",
+ " print(\"Add the path for your image data.\")\n",
+ "else:\n",
+ " input_data = json.dumps({'data': img.tolist()})\n",
+ "\n",
+ " try:\n",
+ " r = aci_service.run(input_data)\n",
+ " result = r['result']\n",
+ " time_ms = np.round(r['time_in_sec'] * 1000, 2)\n",
+ " except Exception as e:\n",
+ " print(str(e))\n",
+ "\n",
+ " plt.figure(figsize = (16, 6))\n",
+ " plt.subplot(1, 15,1)\n",
+ " plt.axhline('')\n",
+ " plt.axvline('')\n",
+ " plt.text(x = -100, y = -20, s = \"Model prediction: \", fontsize = 14)\n",
+ " plt.text(x = -100, y = -10, s = \"Inference time: \", fontsize = 14)\n",
+ " plt.text(x = 0, y = -20, s = str(result), fontsize = 14)\n",
+ " plt.text(x = 0, y = -10, s = str(time_ms) + \" ms\", fontsize = 14)\n",
+ " plt.text(x = -100, y = 14, s = \"Input image: \", fontsize = 14)\n",
+ " plt.imshow(img.reshape(28, 28), cmap = plt.cm.gray) "
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Optional: How does our ONNX MNIST model work? \n",
+ "#### A brief explanation of Convolutional Neural Networks\n",
+ "\n",
+ "A [convolutional neural network](https://en.wikipedia.org/wiki/Convolutional_neural_network) (CNN, or ConvNet) is a type of [feed-forward](https://en.wikipedia.org/wiki/Feedforward_neural_network) artificial neural network made up of neurons that have learnable weights and biases. The CNNs take advantage of the spatial nature of the data. In nature, we perceive different objects by their shapes, size and colors. For example, objects in a natural scene are typically edges, corners/vertices (defined by two of more edges), color patches etc. These primitives are often identified using different detectors (e.g., edge detection, color detector) or combination of detectors interacting to facilitate image interpretation (object classification, region of interest detection, scene description etc.) in real world vision related tasks. These detectors are also known as filters. Convolution is a mathematical operator that takes an image and a filter as input and produces a filtered output (representing say edges, corners, or colors in the input image). \n",
+ "\n",
+ "Historically, these filters are a set of weights that were often hand crafted or modeled with mathematical functions (e.g., [Gaussian](https://en.wikipedia.org/wiki/Gaussian_filter) / [Laplacian](http://homepages.inf.ed.ac.uk/rbf/HIPR2/log.htm) / [Canny](https://en.wikipedia.org/wiki/Canny_edge_detector) filter). The filter outputs are mapped through non-linear activation functions mimicking human brain cells called [neurons](https://en.wikipedia.org/wiki/Neuron). Popular deep CNNs or ConvNets (such as [AlexNet](https://en.wikipedia.org/wiki/AlexNet), [VGG](https://arxiv.org/abs/1409.1556), [Inception](http://www.cv-foundation.org/openaccess/content_cvpr_2015/papers/Szegedy_Going_Deeper_With_2015_CVPR_paper.pdf), [ResNet](https://arxiv.org/pdf/1512.03385v1.pdf)) that are used for various [computer vision](https://en.wikipedia.org/wiki/Computer_vision) tasks have many of these architectural primitives (inspired from biology). \n",
+ "\n",
+ "### Convolution Layer\n",
+ "\n",
+ "A convolution layer is a set of filters. Each filter is defined by a weight (**W**) matrix, and bias ($b$).\n",
+ "\n",
+ "\n",
+ "\n",
+ "These filters are scanned across the image performing the dot product between the weights and corresponding input value ($x$). The bias value is added to the output of the dot product and the resulting sum is optionally mapped through an activation function. This process is illustrated in the following animation."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "Image(url=\"https://www.cntk.ai/jup/cntk103d_conv2d_final.gif\", width= 200)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Model Description\n",
+ "\n",
+ "The MNIST model from the ONNX Model Zoo uses maxpooling to update the weights in its convolutions, summarized by the graphic below. You can see the entire workflow of our pre-trained model in the following image, with our input images and our output probabilities of each of our 10 labels. If you're interested in exploring the logic behind creating a Deep Learning model further, please look at the [training tutorial for our ONNX MNIST Convolutional Neural Network](https://github.com/Microsoft/CNTK/blob/master/Tutorials/CNTK_103D_MNIST_ConvolutionalNeuralNetwork.ipynb). "
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Max-Pooling for Convolutional Neural Nets\n",
+ "\n",
+ ""
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Pre-Trained Model Architecture\n",
+ "\n",
+ ""
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# remember to delete your service after you are done using it!\n",
+ "\n",
+ "# aci_service.delete()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Conclusion\n",
+ "\n",
+ "Congratulations!\n",
+ "\n",
+ "In this tutorial, you have:\n",
+ "- familiarized yourself with ONNX Runtime inference and the pretrained models in the ONNX model zoo\n",
+ "- understood a state-of-the-art convolutional neural net image classification model (MNIST in ONNX) and deployed it in Azure ML cloud\n",
+ "- ensured that your deep learning model is working perfectly (in the cloud) on test data, and checked it against some of your own!\n",
+ "\n",
+ "Next steps:\n",
+ "- Check out another interesting application based on a Microsoft Research computer vision paper that lets you set up a [facial emotion recognition model](https://github.com/Azure/MachineLearningNotebooks/tree/master/onnx/onnx-inference-emotion-recognition.ipynb) in the cloud! This tutorial deploys a pre-trained ONNX Computer Vision model in an Azure ML virtual machine.\n",
+ "- Contribute to our [open source ONNX repository on github](http://github.com/onnx/onnx) and/or add to our [ONNX model zoo](http://github.com/onnx/models)"
+ ]
}
- },
- "source": [
- "### ONNX MNIST Model Methodology\n",
- "\n",
- "The image classification model we are using is pre-trained using Microsoft's deep learning cognitive toolkit, [CNTK](https://github.com/Microsoft/CNTK), from the [ONNX model zoo](http://github.com/onnx/models). The model zoo has many other models that can be deployed on cloud providers like AzureML without any additional training. To ensure that our cloud deployed model works, we use testing data from the famous MNIST data set, provided as part of the [trained MNIST model](https://github.com/onnx/models/tree/master/mnist) in the ONNX model zoo.\n",
- "\n",
- "***Input: Handwritten Images from MNIST Dataset***\n",
- "\n",
- "***Task: Classify each MNIST image into an appropriate digit***\n",
- "\n",
- "***Output: Digit prediction for input image***\n",
- "\n",
- "Run the cell below to look at some of the sample images from the MNIST dataset that we used to train this ONNX model. Remember, once the application is deployed in Azure ML, you can use your own images as input for the model to classify!"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# for images and plots in this notebook\n",
- "import matplotlib.pyplot as plt \n",
- "from IPython.display import Image\n",
- "\n",
- "# display images inline\n",
- "%matplotlib inline"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "Image(url=\"http://3.bp.blogspot.com/_UpN7DfJA0j4/TJtUBWPk0SI/AAAAAAAAABY/oWPMtmqJn3k/s1600/mnist_originals.png\", width=200, height=200)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Specify our Score and Environment Files"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "We are now going to deploy our ONNX Model on AML with inference in ONNX Runtime. We begin by writing a score.py file, which will help us run the model in our Azure ML virtual machine (VM), and then specify our environment by writing a yml file. You will also notice that we import the onnxruntime library to do runtime inference on our ONNX models (passing in input and evaluating out model's predicted output). More information on the API and commands can be found in the [ONNX Runtime documentation](https://aka.ms/onnxruntime).\n",
- "\n",
- "### Write Score File\n",
- "\n",
- "A score file is what tells our Azure cloud service what to do. After initializing our model using azureml.core.model, we start an ONNX Runtime inference session to evaluate the data passed in on our function calls."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%writefile score.py\n",
- "import json\n",
- "import numpy as np\n",
- "import onnxruntime\n",
- "import sys\n",
- "import os\n",
- "from azureml.core.model import Model\n",
- "import time\n",
- "\n",
- "\n",
- "def init():\n",
- " global session, input_name, output_name\n",
- " model = Model.get_model_path(model_name = 'mnist_1')\n",
- " session = onnxruntime.InferenceSession(model, None)\n",
- " input_name = session.get_inputs()[0].name\n",
- " output_name = session.get_outputs()[0].name \n",
- " \n",
- "\n",
- "def preprocess(input_data_json):\n",
- " # convert the JSON data into the tensor input\n",
- " return np.array(json.loads(input_data_json)['data']).astype('float32')\n",
- "\n",
- "def postprocess(result):\n",
- " # We use argmax to pick the highest confidence label\n",
- " return int(np.argmax(np.array(result).squeeze(), axis=0))\n",
- " \n",
- "def run(input_data):\n",
- "\n",
- " try:\n",
- " # load in our data, convert to readable format\n",
- " data = preprocess(input_data)\n",
- " \n",
- " # start timer\n",
- " start = time.time()\n",
- " \n",
- " r = session.run([output_name], {input_name: data})\n",
- " \n",
- " #end timer\n",
- " end = time.time()\n",
- " \n",
- " result = postprocess(r)\n",
- " result_dict = {\"result\": result,\n",
- " \"time_in_sec\": end - start}\n",
- " except Exception as e:\n",
- " result_dict = {\"error\": str(e)}\n",
- " \n",
- " return result_dict\n",
- "\n",
- "def choose_class(result_prob):\n",
- " \"\"\"We use argmax to determine the right label to choose from our output\"\"\"\n",
- " return int(np.argmax(result_prob, axis=0))"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Write Environment File\n",
- "\n",
- "This step creates a YAML environment file that specifies which dependencies we would like to see in our Linux Virtual Machine."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.conda_dependencies import CondaDependencies \n",
- "\n",
- "myenv = CondaDependencies.create(pip_packages=[\"numpy\", \"onnxruntime\", \"azureml-core\"])\n",
- "\n",
- "with open(\"myenv.yml\",\"w\") as f:\n",
- " f.write(myenv.serialize_to_string())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create the Container Image\n",
- "This step will likely take a few minutes."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.image import ContainerImage\n",
- "\n",
- "image_config = ContainerImage.image_configuration(execution_script = \"score.py\",\n",
- " runtime = \"python\",\n",
- " conda_file = \"myenv.yml\",\n",
- " description = \"MNIST ONNX Runtime container\",\n",
- " tags = {\"demo\": \"onnx\"}) \n",
- "\n",
- "\n",
- "image = ContainerImage.create(name = \"onnximage\",\n",
- " # this is the model object\n",
- " models = [model],\n",
- " image_config = image_config,\n",
- " workspace = ws)\n",
- "\n",
- "image.wait_for_creation(show_output = True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "In case you need to debug your code, the next line of code accesses the log file."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "print(image.image_build_log_uri)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "We're all done specifying what we want our virtual machine to do. Let's configure and deploy our container image.\n",
- "\n",
- "### Deploy the container image"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.webservice import AciWebservice\n",
- "\n",
- "aciconfig = AciWebservice.deploy_configuration(cpu_cores = 1, \n",
- " memory_gb = 1, \n",
- " tags = {'demo': 'onnx'}, \n",
- " description = 'ONNX for mnist model')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "The following cell will likely take a few minutes to run as well."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.webservice import Webservice\n",
- "\n",
- "aci_service_name = 'onnx-demo-mnist'\n",
- "print(\"Service\", aci_service_name)\n",
- "\n",
- "aci_service = Webservice.deploy_from_image(deployment_config = aciconfig,\n",
- " image = image,\n",
- " name = aci_service_name,\n",
- " workspace = ws)\n",
- "\n",
- "aci_service.wait_for_deployment(True)\n",
- "print(aci_service.state)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "if aci_service.state != 'Healthy':\n",
- " # run this command for debugging.\n",
- " print(aci_service.get_logs())\n",
- "\n",
- " # If your deployment fails, make sure to delete your aci_service or rename your service before trying again!\n",
- " # aci_service.delete()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Success!\n",
- "\n",
- "If you've made it this far, you've deployed a working VM with a handwritten digit classifier running in the cloud using Azure ML. Congratulations!\n",
- "\n",
- "You can get the URL for the webservice with the code below. Let's now see how well our model deals with our test images."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "print(aci_service.scoring_uri)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Testing and Evaluation\n",
- "\n",
- "### Load Test Data\n",
- "\n",
- "These are already in your directory from your ONNX model download (from the model zoo).\n",
- "\n",
- "Notice that our Model Zoo files have a .pb extension. This is because they are [protobuf files (Protocol Buffers)](https://developers.google.com/protocol-buffers/docs/pythontutorial), so we need to read in our data through our ONNX TensorProto reader into a format we can work with, like numerical arrays."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# to manipulate our arrays\n",
- "import numpy as np \n",
- "\n",
- "# read in test data protobuf files included with the model\n",
- "import onnx\n",
- "from onnx import numpy_helper\n",
- "\n",
- "# to use parsers to read in our model/data\n",
- "import json\n",
- "import os\n",
- "\n",
- "test_inputs = []\n",
- "test_outputs = []\n",
- "\n",
- "# read in 3 testing images from .pb files\n",
- "test_data_size = 3\n",
- "\n",
- "for i in np.arange(test_data_size):\n",
- " input_test_data = os.path.join(model_dir, 'test_data_set_{0}'.format(i), 'input_0.pb')\n",
- " output_test_data = os.path.join(model_dir, 'test_data_set_{0}'.format(i), 'output_0.pb')\n",
- " \n",
- " # convert protobuf tensors to np arrays using the TensorProto reader from ONNX\n",
- " tensor = onnx.TensorProto()\n",
- " with open(input_test_data, 'rb') as f:\n",
- " tensor.ParseFromString(f.read())\n",
- " \n",
- " input_data = numpy_helper.to_array(tensor)\n",
- " test_inputs.append(input_data)\n",
- " \n",
- " with open(output_test_data, 'rb') as f:\n",
- " tensor.ParseFromString(f.read())\n",
- " \n",
- " output_data = numpy_helper.to_array(tensor)\n",
- " test_outputs.append(output_data)\n",
- " \n",
- "if len(test_inputs) == test_data_size:\n",
- " print('Test data loaded successfully.')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {
- "nbpresent": {
- "id": "c3f2f57c-7454-4d3e-b38d-b0946cf066ea"
- }
- },
- "source": [
- "### Show some sample images\n",
- "We use `matplotlib` to plot 3 test images from the dataset."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "nbpresent": {
- "id": "396d478b-34aa-4afa-9898-cdce8222a516"
- }
- },
- "outputs": [],
- "source": [
- "plt.figure(figsize = (16, 6))\n",
- "for test_image in np.arange(3):\n",
- " plt.subplot(1, 15, test_image+1)\n",
- " plt.axhline('')\n",
- " plt.axvline('')\n",
- " plt.imshow(test_inputs[test_image].reshape(28, 28), cmap = plt.cm.Greys)\n",
- "plt.show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Run evaluation / prediction"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "plt.figure(figsize = (16, 6), frameon=False)\n",
- "plt.subplot(1, 8, 1)\n",
- "\n",
- "plt.text(x = 0, y = -30, s = \"True Label: \", fontsize = 13, color = 'black')\n",
- "plt.text(x = 0, y = -20, s = \"Result: \", fontsize = 13, color = 'black')\n",
- "plt.text(x = 0, y = -10, s = \"Inference Time: \", fontsize = 13, color = 'black')\n",
- "plt.text(x = 3, y = 14, s = \"Model Input\", fontsize = 12, color = 'black')\n",
- "plt.text(x = 6, y = 18, s = \"(28 x 28)\", fontsize = 12, color = 'black')\n",
- "plt.imshow(np.ones((28,28)), cmap=plt.cm.Greys) \n",
- "\n",
- "\n",
- "for i in np.arange(test_data_size):\n",
- " \n",
- " input_data = json.dumps({'data': test_inputs[i].tolist()})\n",
- " \n",
- " # predict using the deployed model\n",
- " r = aci_service.run(input_data)\n",
- " \n",
- " if \"error\" in r:\n",
- " print(r['error'])\n",
- " break\n",
- " \n",
- " result = r['result']\n",
- " time_ms = np.round(r['time_in_sec'] * 1000, 2)\n",
- " \n",
- " ground_truth = int(np.argmax(test_outputs[i]))\n",
- " \n",
- " # compare actual value vs. the predicted values:\n",
- " plt.subplot(1, 8, i+2)\n",
- " plt.axhline('')\n",
- " plt.axvline('')\n",
- "\n",
- " # use different color for misclassified sample\n",
- " font_color = 'red' if ground_truth != result else 'black'\n",
- " clr_map = plt.cm.gray if ground_truth != result else plt.cm.Greys\n",
- "\n",
- " # ground truth labels are in blue\n",
- " plt.text(x = 10, y = -30, s = ground_truth, fontsize = 18, color = 'blue')\n",
- " \n",
- " # predictions are in black if correct, red if incorrect\n",
- " plt.text(x = 10, y = -20, s = result, fontsize = 18, color = font_color)\n",
- " plt.text(x = 5, y = -10, s = str(time_ms) + ' ms', fontsize = 14, color = font_color)\n",
- "\n",
- " \n",
- " plt.imshow(test_inputs[i].reshape(28, 28), cmap = clr_map)\n",
- "\n",
- "plt.show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Try classifying your own images!\n",
- "\n",
- "Create your own handwritten image and pass it into the model."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Preprocessing functions take your image and format it so it can be passed\n",
- "# as input into our ONNX model\n",
- "\n",
- "import cv2\n",
- "\n",
- "def rgb2gray(rgb):\n",
- " \"\"\"Convert the input image into grayscale\"\"\"\n",
- " return np.dot(rgb[...,:3], [0.299, 0.587, 0.114])\n",
- "\n",
- "def resize_img(img):\n",
- " \"\"\"Resize image to MNIST model input dimensions\"\"\"\n",
- " img = cv2.resize(img, dsize=(28, 28), interpolation=cv2.INTER_AREA)\n",
- " img.resize((1, 1, 28, 28))\n",
- " return img\n",
- "\n",
- "def preprocess(img):\n",
- " \"\"\"Resize input images and convert them to grayscale.\"\"\"\n",
- " if img.shape == (28, 28):\n",
- " img.resize((1, 1, 28, 28))\n",
- " return img\n",
- " \n",
- " grayscale = rgb2gray(img)\n",
- " processed_img = resize_img(grayscale)\n",
- " return processed_img"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Replace this string with your own path/test image\n",
- "# Make sure your image is square and the dimensions are equal (i.e. 100 * 100 pixels or 28 * 28 pixels)\n",
- "\n",
- "# Any PNG or JPG image file should work\n",
- "\n",
- "your_test_image = \"\"\n",
- "\n",
- "# e.g. your_test_image = \"C:/Users/vinitra.swamy/Pictures/handwritten_digit.png\"\n",
- "\n",
- "import matplotlib.image as mpimg\n",
- "\n",
- "if your_test_image != \"\":\n",
- " img = mpimg.imread(your_test_image)\n",
- " plt.subplot(1,3,1)\n",
- " plt.imshow(img, cmap = plt.cm.Greys)\n",
- " print(\"Old Dimensions: \", img.shape)\n",
- " img = preprocess(img)\n",
- " print(\"New Dimensions: \", img.shape)\n",
- "else:\n",
- " img = None"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "if img is None:\n",
- " print(\"Add the path for your image data.\")\n",
- "else:\n",
- " input_data = json.dumps({'data': img.tolist()})\n",
- "\n",
- " try:\n",
- " r = aci_service.run(input_data)\n",
- " result = r['result']\n",
- " time_ms = np.round(r['time_in_sec'] * 1000, 2)\n",
- " except Exception as e:\n",
- " print(str(e))\n",
- "\n",
- " plt.figure(figsize = (16, 6))\n",
- " plt.subplot(1, 15,1)\n",
- " plt.axhline('')\n",
- " plt.axvline('')\n",
- " plt.text(x = -100, y = -20, s = \"Model prediction: \", fontsize = 14)\n",
- " plt.text(x = -100, y = -10, s = \"Inference time: \", fontsize = 14)\n",
- " plt.text(x = 0, y = -20, s = str(result), fontsize = 14)\n",
- " plt.text(x = 0, y = -10, s = str(time_ms) + \" ms\", fontsize = 14)\n",
- " plt.text(x = -100, y = 14, s = \"Input image: \", fontsize = 14)\n",
- " plt.imshow(img.reshape(28, 28), cmap = plt.cm.gray) "
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Optional: How does our ONNX MNIST model work? \n",
- "#### A brief explanation of Convolutional Neural Networks\n",
- "\n",
- "A [convolutional neural network](https://en.wikipedia.org/wiki/Convolutional_neural_network) (CNN, or ConvNet) is a type of [feed-forward](https://en.wikipedia.org/wiki/Feedforward_neural_network) artificial neural network made up of neurons that have learnable weights and biases. The CNNs take advantage of the spatial nature of the data. In nature, we perceive different objects by their shapes, size and colors. For example, objects in a natural scene are typically edges, corners/vertices (defined by two of more edges), color patches etc. These primitives are often identified using different detectors (e.g., edge detection, color detector) or combination of detectors interacting to facilitate image interpretation (object classification, region of interest detection, scene description etc.) in real world vision related tasks. These detectors are also known as filters. Convolution is a mathematical operator that takes an image and a filter as input and produces a filtered output (representing say edges, corners, or colors in the input image). \n",
- "\n",
- "Historically, these filters are a set of weights that were often hand crafted or modeled with mathematical functions (e.g., [Gaussian](https://en.wikipedia.org/wiki/Gaussian_filter) / [Laplacian](http://homepages.inf.ed.ac.uk/rbf/HIPR2/log.htm) / [Canny](https://en.wikipedia.org/wiki/Canny_edge_detector) filter). The filter outputs are mapped through non-linear activation functions mimicking human brain cells called [neurons](https://en.wikipedia.org/wiki/Neuron). Popular deep CNNs or ConvNets (such as [AlexNet](https://en.wikipedia.org/wiki/AlexNet), [VGG](https://arxiv.org/abs/1409.1556), [Inception](http://www.cv-foundation.org/openaccess/content_cvpr_2015/papers/Szegedy_Going_Deeper_With_2015_CVPR_paper.pdf), [ResNet](https://arxiv.org/pdf/1512.03385v1.pdf)) that are used for various [computer vision](https://en.wikipedia.org/wiki/Computer_vision) tasks have many of these architectural primitives (inspired from biology). \n",
- "\n",
- "### Convolution Layer\n",
- "\n",
- "A convolution layer is a set of filters. Each filter is defined by a weight (**W**) matrix, and bias ($b$).\n",
- "\n",
- "\n",
- "\n",
- "These filters are scanned across the image performing the dot product between the weights and corresponding input value ($x$). The bias value is added to the output of the dot product and the resulting sum is optionally mapped through an activation function. This process is illustrated in the following animation."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "Image(url=\"https://www.cntk.ai/jup/cntk103d_conv2d_final.gif\", width= 200)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Model Description\n",
- "\n",
- "The MNIST model from the ONNX Model Zoo uses maxpooling to update the weights in its convolutions, summarized by the graphic below. You can see the entire workflow of our pre-trained model in the following image, with our input images and our output probabilities of each of our 10 labels. If you're interested in exploring the logic behind creating a Deep Learning model further, please look at the [training tutorial for our ONNX MNIST Convolutional Neural Network](https://github.com/Microsoft/CNTK/blob/master/Tutorials/CNTK_103D_MNIST_ConvolutionalNeuralNetwork.ipynb). "
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Max-Pooling for Convolutional Neural Nets\n",
- "\n",
- ""
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Pre-Trained Model Architecture\n",
- "\n",
- ""
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# remember to delete your service after you are done using it!\n",
- "\n",
- "# aci_service.delete()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Conclusion\n",
- "\n",
- "Congratulations!\n",
- "\n",
- "In this tutorial, you have:\n",
- "- familiarized yourself with ONNX Runtime inference and the pretrained models in the ONNX model zoo\n",
- "- understood a state-of-the-art convolutional neural net image classification model (MNIST in ONNX) and deployed it in Azure ML cloud\n",
- "- ensured that your deep learning model is working perfectly (in the cloud) on test data, and checked it against some of your own!\n",
- "\n",
- "Next steps:\n",
- "- Check out another interesting application based on a Microsoft Research computer vision paper that lets you set up a [facial emotion recognition model](https://github.com/Azure/MachineLearningNotebooks/tree/master/onnx/onnx-inference-emotion-recognition.ipynb) in the cloud! This tutorial deploys a pre-trained ONNX Computer Vision model in an Azure ML virtual machine.\n",
- "- Contribute to our [open source ONNX repository on github](http://github.com/onnx/onnx) and/or add to our [ONNX model zoo](http://github.com/onnx/models)"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "viswamy"
- }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "viswamy"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.5"
+ },
+ "msauthor": "vinitra.swamy"
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.5"
- },
- "msauthor": "vinitra.swamy"
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/deployment/onnx/onnx-modelzoo-aml-deploy-resnet50.ipynb b/how-to-use-azureml/deployment/onnx/onnx-modelzoo-aml-deploy-resnet50.ipynb
index 1d587a6d..f1c5858a 100644
--- a/how-to-use-azureml/deployment/onnx/onnx-modelzoo-aml-deploy-resnet50.ipynb
+++ b/how-to-use-azureml/deployment/onnx/onnx-modelzoo-aml-deploy-resnet50.ipynb
@@ -1,419 +1,419 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved. \n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# ResNet50 Image Classification using ONNX and AzureML\n",
- "\n",
- "This example shows how to deploy the ResNet50 ONNX model as a web service using Azure Machine Learning services and the ONNX Runtime.\n",
- "\n",
- "## What is ONNX\n",
- "ONNX is an open format for representing machine learning and deep learning models. ONNX enables open and interoperable AI by enabling data scientists and developers to use the tools of their choice without worrying about lock-in and flexibility to deploy to a variety of platforms. ONNX is developed and supported by a community of partners including Microsoft, Facebook, and Amazon. For more information, explore the [ONNX website](http://onnx.ai).\n",
- "\n",
- "## ResNet50 Details\n",
- "ResNet classifies the major object in an input image into a set of 1000 pre-defined classes. For more information about the ResNet50 model and how it was created can be found on the [ONNX Model Zoo github](https://github.com/onnx/models/tree/master/models/image_classification/resnet). "
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Prerequisites\n",
- "\n",
- "To make the best use of your time, make sure you have done the following:\n",
- "\n",
- "* Understand the [architecture and terms](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture) introduced by Azure Machine Learning\n",
- "* Go through the [00.configuration.ipynb](../00.configuration.ipynb) notebook to:\n",
- " * install the AML SDK\n",
- " * create a workspace and its configuration file (config.json)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Check core SDK version number\n",
- "import azureml.core\n",
- "\n",
- "print(\"SDK version:\", azureml.core.VERSION)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Download pre-trained ONNX model from ONNX Model Zoo.\n",
- "\n",
- "Download the [ResNet50v2 model and test data](https://s3.amazonaws.com/onnx-model-zoo/resnet/resnet50v2/resnet50v2.tar.gz) and extract it in the same folder as this tutorial notebook.\n"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import urllib.request\n",
- "\n",
- "onnx_model_url = \"https://s3.amazonaws.com/onnx-model-zoo/resnet/resnet50v2/resnet50v2.tar.gz\"\n",
- "urllib.request.urlretrieve(onnx_model_url, filename=\"resnet50v2.tar.gz\")\n",
- "\n",
- "!tar xvzf resnet50v2.tar.gz"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Deploying as a web service with Azure ML"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Load your Azure ML workspace\n",
- "\n",
- "We begin by instantiating a workspace object from the existing workspace created earlier in the configuration notebook."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core import Workspace\n",
- "\n",
- "ws = Workspace.from_config()\n",
- "print(ws.name, ws.location, ws.resource_group, sep = '\\n')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Register your model with Azure ML\n",
- "\n",
- "Now we upload the model and register it in the workspace."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.model import Model\n",
- "\n",
- "model = Model.register(model_path = \"resnet50v2/resnet50v2.onnx\",\n",
- " model_name = \"resnet50v2\",\n",
- " tags = {\"onnx\": \"demo\"},\n",
- " description = \"ResNet50v2 from ONNX Model Zoo\",\n",
- " workspace = ws)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Displaying your registered models\n",
- "\n",
- "You can optionally list out all the models that you have registered in this workspace."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "models = ws.models\n",
- "for name, m in models.items():\n",
- " print(\"Name:\", name,\"\\tVersion:\", m.version, \"\\tDescription:\", m.description, m.tags)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Write scoring file\n",
- "\n",
- "We are now going to deploy our ONNX model on Azure ML using the ONNX Runtime. We begin by writing a score.py file that will be invoked by the web service call. The `init()` function is called once when the container is started so we load the model using the ONNX Runtime into a global session object."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%writefile score.py\n",
- "import json\n",
- "import time\n",
- "import sys\n",
- "import os\n",
- "from azureml.core.model import Model\n",
- "import numpy as np # we're going to use numpy to process input and output data\n",
- "import onnxruntime # to inference ONNX models, we use the ONNX Runtime\n",
- "\n",
- "def softmax(x):\n",
- " x = x.reshape(-1)\n",
- " e_x = np.exp(x - np.max(x))\n",
- " return e_x / e_x.sum(axis=0)\n",
- "\n",
- "def init():\n",
- " global session\n",
- " model = Model.get_model_path(model_name = 'resnet50v2')\n",
- " session = onnxruntime.InferenceSession(model, None)\n",
- "\n",
- "def preprocess(input_data_json):\n",
- " # convert the JSON data into the tensor input\n",
- " img_data = np.array(json.loads(input_data_json)['data']).astype('float32')\n",
- " \n",
- " #normalize\n",
- " mean_vec = np.array([0.485, 0.456, 0.406])\n",
- " stddev_vec = np.array([0.229, 0.224, 0.225])\n",
- " norm_img_data = np.zeros(img_data.shape).astype('float32')\n",
- " for i in range(img_data.shape[0]):\n",
- " norm_img_data[i,:,:] = (img_data[i,:,:]/255 - mean_vec[i]) / stddev_vec[i]\n",
- "\n",
- " return norm_img_data\n",
- "\n",
- "def postprocess(result):\n",
- " return softmax(np.array(result)).tolist()\n",
- "\n",
- "def run(input_data_json):\n",
- " try:\n",
- " start = time.time()\n",
- " # load in our data which is expected as NCHW 224x224 image\n",
- " input_data = preprocess(input_data_json)\n",
- " input_name = session.get_inputs()[0].name # get the id of the first input of the model \n",
- " result = session.run([], {input_name: input_data})\n",
- " end = time.time() # stop timer\n",
- " return {\"result\": postprocess(result),\n",
- " \"time\": end - start}\n",
- " except Exception as e:\n",
- " result = str(e)\n",
- " return {\"error\": result}"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create container image"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "First we create a YAML file that specifies which dependencies we would like to see in our container."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.conda_dependencies import CondaDependencies \n",
- "\n",
- "myenv = CondaDependencies.create(pip_packages=[\"numpy\",\"onnxruntime\",\"azureml-core\"])\n",
- "\n",
- "with open(\"myenv.yml\",\"w\") as f:\n",
- " f.write(myenv.serialize_to_string())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Then we have Azure ML create the container. This step will likely take a few minutes."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.image import ContainerImage\n",
- "\n",
- "image_config = ContainerImage.image_configuration(execution_script = \"score.py\",\n",
- " runtime = \"python\",\n",
- " conda_file = \"myenv.yml\",\n",
- " description = \"ONNX ResNet50 Demo\",\n",
- " tags = {\"demo\": \"onnx\"}\n",
- " )\n",
- "\n",
- "\n",
- "image = ContainerImage.create(name = \"onnxresnet50v2\",\n",
- " models = [model],\n",
- " image_config = image_config,\n",
- " workspace = ws)\n",
- "\n",
- "image.wait_for_creation(show_output = True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "In case you need to debug your code, the next line of code accesses the log file."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "print(image.image_build_log_uri)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "We're all set! Let's get our model chugging.\n",
- "\n",
- "### Deploy the container image"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.webservice import AciWebservice\n",
- "\n",
- "aciconfig = AciWebservice.deploy_configuration(cpu_cores = 1, \n",
- " memory_gb = 1, \n",
- " tags = {'demo': 'onnx'}, \n",
- " description = 'web service for ResNet50 ONNX model')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "The following cell will likely take a few minutes to run as well."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.webservice import Webservice\n",
- "from random import randint\n",
- "\n",
- "aci_service_name = 'onnx-demo-resnet50'+str(randint(0,100))\n",
- "print(\"Service\", aci_service_name)\n",
- "\n",
- "aci_service = Webservice.deploy_from_image(deployment_config = aciconfig,\n",
- " image = image,\n",
- " name = aci_service_name,\n",
- " workspace = ws)\n",
- "\n",
- "aci_service.wait_for_deployment(True)\n",
- "print(aci_service.state)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "In case the deployment fails, you can check the logs. Make sure to delete your aci_service before trying again."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "if aci_service.state != 'Healthy':\n",
- " # run this command for debugging.\n",
- " print(aci_service.get_logs())\n",
- " aci_service.delete()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Success!\n",
- "\n",
- "If you've made it this far, you've deployed a working web service that does image classification using an ONNX model. You can get the URL for the webservice with the code below."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "print(aci_service.scoring_uri)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "When you are eventually done using the web service, remember to delete it."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "#aci_service.delete()"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "onnx"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved. \n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# ResNet50 Image Classification using ONNX and AzureML\n",
+ "\n",
+ "This example shows how to deploy the ResNet50 ONNX model as a web service using Azure Machine Learning services and the ONNX Runtime.\n",
+ "\n",
+ "## What is ONNX\n",
+ "ONNX is an open format for representing machine learning and deep learning models. ONNX enables open and interoperable AI by enabling data scientists and developers to use the tools of their choice without worrying about lock-in and flexibility to deploy to a variety of platforms. ONNX is developed and supported by a community of partners including Microsoft, Facebook, and Amazon. For more information, explore the [ONNX website](http://onnx.ai).\n",
+ "\n",
+ "## ResNet50 Details\n",
+ "ResNet classifies the major object in an input image into a set of 1000 pre-defined classes. For more information about the ResNet50 model and how it was created can be found on the [ONNX Model Zoo github](https://github.com/onnx/models/tree/master/models/image_classification/resnet). "
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Prerequisites\n",
+ "\n",
+ "To make the best use of your time, make sure you have done the following:\n",
+ "\n",
+ "* Understand the [architecture and terms](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture) introduced by Azure Machine Learning\n",
+ "* Go through the [configuration notebook](../../../configuration.ipynb) to:\n",
+ " * install the AML SDK\n",
+ " * create a workspace and its configuration file (config.json)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Check core SDK version number\n",
+ "import azureml.core\n",
+ "\n",
+ "print(\"SDK version:\", azureml.core.VERSION)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Download pre-trained ONNX model from ONNX Model Zoo.\n",
+ "\n",
+ "Download the [ResNet50v2 model and test data](https://s3.amazonaws.com/onnx-model-zoo/resnet/resnet50v2/resnet50v2.tar.gz) and extract it in the same folder as this tutorial notebook.\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import urllib.request\n",
+ "\n",
+ "onnx_model_url = \"https://s3.amazonaws.com/onnx-model-zoo/resnet/resnet50v2/resnet50v2.tar.gz\"\n",
+ "urllib.request.urlretrieve(onnx_model_url, filename=\"resnet50v2.tar.gz\")\n",
+ "\n",
+ "!tar xvzf resnet50v2.tar.gz"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Deploying as a web service with Azure ML"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Load your Azure ML workspace\n",
+ "\n",
+ "We begin by instantiating a workspace object from the existing workspace created earlier in the configuration notebook."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core import Workspace\n",
+ "\n",
+ "ws = Workspace.from_config()\n",
+ "print(ws.name, ws.location, ws.resource_group, sep = '\\n')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Register your model with Azure ML\n",
+ "\n",
+ "Now we upload the model and register it in the workspace."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.model import Model\n",
+ "\n",
+ "model = Model.register(model_path = \"resnet50v2/resnet50v2.onnx\",\n",
+ " model_name = \"resnet50v2\",\n",
+ " tags = {\"onnx\": \"demo\"},\n",
+ " description = \"ResNet50v2 from ONNX Model Zoo\",\n",
+ " workspace = ws)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Displaying your registered models\n",
+ "\n",
+ "You can optionally list out all the models that you have registered in this workspace."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "models = ws.models\n",
+ "for name, m in models.items():\n",
+ " print(\"Name:\", name,\"\\tVersion:\", m.version, \"\\tDescription:\", m.description, m.tags)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Write scoring file\n",
+ "\n",
+ "We are now going to deploy our ONNX model on Azure ML using the ONNX Runtime. We begin by writing a score.py file that will be invoked by the web service call. The `init()` function is called once when the container is started so we load the model using the ONNX Runtime into a global session object."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%writefile score.py\n",
+ "import json\n",
+ "import time\n",
+ "import sys\n",
+ "import os\n",
+ "from azureml.core.model import Model\n",
+ "import numpy as np # we're going to use numpy to process input and output data\n",
+ "import onnxruntime # to inference ONNX models, we use the ONNX Runtime\n",
+ "\n",
+ "def softmax(x):\n",
+ " x = x.reshape(-1)\n",
+ " e_x = np.exp(x - np.max(x))\n",
+ " return e_x / e_x.sum(axis=0)\n",
+ "\n",
+ "def init():\n",
+ " global session\n",
+ " model = Model.get_model_path(model_name = 'resnet50v2')\n",
+ " session = onnxruntime.InferenceSession(model, None)\n",
+ "\n",
+ "def preprocess(input_data_json):\n",
+ " # convert the JSON data into the tensor input\n",
+ " img_data = np.array(json.loads(input_data_json)['data']).astype('float32')\n",
+ " \n",
+ " #normalize\n",
+ " mean_vec = np.array([0.485, 0.456, 0.406])\n",
+ " stddev_vec = np.array([0.229, 0.224, 0.225])\n",
+ " norm_img_data = np.zeros(img_data.shape).astype('float32')\n",
+ " for i in range(img_data.shape[0]):\n",
+ " norm_img_data[i,:,:] = (img_data[i,:,:]/255 - mean_vec[i]) / stddev_vec[i]\n",
+ "\n",
+ " return norm_img_data\n",
+ "\n",
+ "def postprocess(result):\n",
+ " return softmax(np.array(result)).tolist()\n",
+ "\n",
+ "def run(input_data_json):\n",
+ " try:\n",
+ " start = time.time()\n",
+ " # load in our data which is expected as NCHW 224x224 image\n",
+ " input_data = preprocess(input_data_json)\n",
+ " input_name = session.get_inputs()[0].name # get the id of the first input of the model \n",
+ " result = session.run([], {input_name: input_data})\n",
+ " end = time.time() # stop timer\n",
+ " return {\"result\": postprocess(result),\n",
+ " \"time\": end - start}\n",
+ " except Exception as e:\n",
+ " result = str(e)\n",
+ " return {\"error\": result}"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create container image"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "First we create a YAML file that specifies which dependencies we would like to see in our container."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.conda_dependencies import CondaDependencies \n",
+ "\n",
+ "myenv = CondaDependencies.create(pip_packages=[\"numpy\",\"onnxruntime\",\"azureml-core\"])\n",
+ "\n",
+ "with open(\"myenv.yml\",\"w\") as f:\n",
+ " f.write(myenv.serialize_to_string())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Then we have Azure ML create the container. This step will likely take a few minutes."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.image import ContainerImage\n",
+ "\n",
+ "image_config = ContainerImage.image_configuration(execution_script = \"score.py\",\n",
+ " runtime = \"python\",\n",
+ " conda_file = \"myenv.yml\",\n",
+ " description = \"ONNX ResNet50 Demo\",\n",
+ " tags = {\"demo\": \"onnx\"}\n",
+ " )\n",
+ "\n",
+ "\n",
+ "image = ContainerImage.create(name = \"onnxresnet50v2\",\n",
+ " models = [model],\n",
+ " image_config = image_config,\n",
+ " workspace = ws)\n",
+ "\n",
+ "image.wait_for_creation(show_output = True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "In case you need to debug your code, the next line of code accesses the log file."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "print(image.image_build_log_uri)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "We're all set! Let's get our model chugging.\n",
+ "\n",
+ "### Deploy the container image"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.webservice import AciWebservice\n",
+ "\n",
+ "aciconfig = AciWebservice.deploy_configuration(cpu_cores = 1, \n",
+ " memory_gb = 1, \n",
+ " tags = {'demo': 'onnx'}, \n",
+ " description = 'web service for ResNet50 ONNX model')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "The following cell will likely take a few minutes to run as well."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.webservice import Webservice\n",
+ "from random import randint\n",
+ "\n",
+ "aci_service_name = 'onnx-demo-resnet50'+str(randint(0,100))\n",
+ "print(\"Service\", aci_service_name)\n",
+ "\n",
+ "aci_service = Webservice.deploy_from_image(deployment_config = aciconfig,\n",
+ " image = image,\n",
+ " name = aci_service_name,\n",
+ " workspace = ws)\n",
+ "\n",
+ "aci_service.wait_for_deployment(True)\n",
+ "print(aci_service.state)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "In case the deployment fails, you can check the logs. Make sure to delete your aci_service before trying again."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "if aci_service.state != 'Healthy':\n",
+ " # run this command for debugging.\n",
+ " print(aci_service.get_logs())\n",
+ " aci_service.delete()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Success!\n",
+ "\n",
+ "If you've made it this far, you've deployed a working web service that does image classification using an ONNX model. You can get the URL for the webservice with the code below."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "print(aci_service.scoring_uri)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "When you are eventually done using the web service, remember to delete it."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "#aci_service.delete()"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "onnx"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.5.6"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.5.6"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/deployment/production-deploy-to-aks/production-deploy-to-aks.ipynb b/how-to-use-azureml/deployment/production-deploy-to-aks/production-deploy-to-aks.ipynb
index b7a0f545..f1b8d285 100644
--- a/how-to-use-azureml/deployment/production-deploy-to-aks/production-deploy-to-aks.ipynb
+++ b/how-to-use-azureml/deployment/production-deploy-to-aks/production-deploy-to-aks.ipynb
@@ -1,343 +1,343 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Deploying a web service to Azure Kubernetes Service (AKS)\n",
- "This notebook shows the steps for deploying a service: registering a model, creating an image, provisioning a cluster (one time action), and deploying a service to it. \n",
- "We then test and delete the service, image and model."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core import Workspace\n",
- "from azureml.core.compute import AksCompute, ComputeTarget\n",
- "from azureml.core.webservice import Webservice, AksWebservice\n",
- "from azureml.core.image import Image\n",
- "from azureml.core.model import Model"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import azureml.core\n",
- "print(azureml.core.VERSION)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Get workspace\n",
- "Load existing workspace from the config file info."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.workspace import Workspace\n",
- "\n",
- "ws = Workspace.from_config()\n",
- "print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep = '\\n')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Register the model\n",
- "Register an existing trained model, add descirption and tags."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "#Register the model\n",
- "from azureml.core.model import Model\n",
- "model = Model.register(model_path = \"sklearn_regression_model.pkl\", # this points to a local file\n",
- " model_name = \"sklearn_regression_model.pkl\", # this is the name the model is registered as\n",
- " tags = {'area': \"diabetes\", 'type': \"regression\"},\n",
- " description = \"Ridge regression model to predict diabetes\",\n",
- " workspace = ws)\n",
- "\n",
- "print(model.name, model.description, model.version)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Create an image\n",
- "Create an image using the registered model the script that will load and run the model."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%writefile score.py\n",
- "import pickle\n",
- "import json\n",
- "import numpy\n",
- "from sklearn.externals import joblib\n",
- "from sklearn.linear_model import Ridge\n",
- "from azureml.core.model import Model\n",
- "\n",
- "def init():\n",
- " global model\n",
- " # note here \"sklearn_regression_model.pkl\" is the name of the model registered under\n",
- " # this is a different behavior than before when the code is run locally, even though the code is the same.\n",
- " model_path = Model.get_model_path('sklearn_regression_model.pkl')\n",
- " # deserialize the model file back into a sklearn model\n",
- " model = joblib.load(model_path)\n",
- "\n",
- "# note you can pass in multiple rows for scoring\n",
- "def run(raw_data):\n",
- " try:\n",
- " data = json.loads(raw_data)['data']\n",
- " data = numpy.array(data)\n",
- " result = model.predict(data)\n",
- " # you can return any data type as long as it is JSON-serializable\n",
- " return result.tolist()\n",
- " except Exception as e:\n",
- " error = str(e)\n",
- " return error"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.conda_dependencies import CondaDependencies \n",
- "\n",
- "myenv = CondaDependencies.create(conda_packages=['numpy','scikit-learn'])\n",
- "\n",
- "with open(\"myenv.yml\",\"w\") as f:\n",
- " f.write(myenv.serialize_to_string())"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.image import ContainerImage\n",
- "\n",
- "image_config = ContainerImage.image_configuration(execution_script = \"score.py\",\n",
- " runtime = \"python\",\n",
- " conda_file = \"myenv.yml\",\n",
- " description = \"Image with ridge regression model\",\n",
- " tags = {'area': \"diabetes\", 'type': \"regression\"}\n",
- " )\n",
- "\n",
- "image = ContainerImage.create(name = \"myimage1\",\n",
- " # this is the model object\n",
- " models = [model],\n",
- " image_config = image_config,\n",
- " workspace = ws)\n",
- "\n",
- "image.wait_for_creation(show_output = True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Provision the AKS Cluster\n",
- "This is a one time setup. You can reuse this cluster for multiple deployments after it has been created. If you delete the cluster or the resource group that contains it, then you would have to recreate it."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Use the default configuration (can also provide parameters to customize)\n",
- "prov_config = AksCompute.provisioning_configuration()\n",
- "\n",
- "aks_name = 'my-aks-9' \n",
- "# Create the cluster\n",
- "aks_target = ComputeTarget.create(workspace = ws, \n",
- " name = aks_name, \n",
- " provisioning_configuration = prov_config)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%time\n",
- "aks_target.wait_for_completion(show_output = True)\n",
- "print(aks_target.provisioning_state)\n",
- "print(aks_target.provisioning_errors)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Optional step: Attach existing AKS cluster\n",
- "\n",
- "If you have existing AKS cluster in your Azure subscription, you can attach it to the Workspace."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "'''\n",
- "# Use the default configuration (can also provide parameters to customize)\n",
- "resource_id = '/subscriptions/92c76a2f-0e1c-4216-b65e-abf7a3f34c1e/resourcegroups/raymondsdk0604/providers/Microsoft.ContainerService/managedClusters/my-aks-0605d37425356b7d01'\n",
- "\n",
- "create_name='my-existing-aks' \n",
- "# Create the cluster\n",
- "attach_config = AksCompute.attach_configuration(resource_id=resource_id)\n",
- "aks_target = ComputeTarget.attach(workspace=ws, name=create_name, attach_configuration=attach_config)\n",
- "# Wait for the operation to complete\n",
- "aks_target.wait_for_completion(True)\n",
- "'''"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Deploy web service to AKS"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "#Set the web service configuration (using default here)\n",
- "aks_config = AksWebservice.deploy_configuration()"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%time\n",
- "aks_service_name ='aks-service-1'\n",
- "\n",
- "aks_service = Webservice.deploy_from_image(workspace = ws, \n",
- " name = aks_service_name,\n",
- " image = image,\n",
- " deployment_config = aks_config,\n",
- " deployment_target = aks_target)\n",
- "aks_service.wait_for_deployment(show_output = True)\n",
- "print(aks_service.state)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Test the web service\n",
- "We test the web sevice by passing data."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%time\n",
- "import json\n",
- "\n",
- "test_sample = json.dumps({'data': [\n",
- " [1,2,3,4,5,6,7,8,9,10], \n",
- " [10,9,8,7,6,5,4,3,2,1]\n",
- "]})\n",
- "test_sample = bytes(test_sample,encoding = 'utf8')\n",
- "\n",
- "prediction = aks_service.run(input_data = test_sample)\n",
- "print(prediction)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Clean up\n",
- "Delete the service, image and model."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%time\n",
- "aks_service.delete()\n",
- "image.delete()\n",
- "model.delete()"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "raymondl"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Deploying a web service to Azure Kubernetes Service (AKS)\n",
+ "This notebook shows the steps for deploying a service: registering a model, creating an image, provisioning a cluster (one time action), and deploying a service to it. \n",
+ "We then test and delete the service, image and model."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core import Workspace\n",
+ "from azureml.core.compute import AksCompute, ComputeTarget\n",
+ "from azureml.core.webservice import Webservice, AksWebservice\n",
+ "from azureml.core.image import Image\n",
+ "from azureml.core.model import Model"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import azureml.core\n",
+ "print(azureml.core.VERSION)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Get workspace\n",
+ "Load existing workspace from the config file info."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.workspace import Workspace\n",
+ "\n",
+ "ws = Workspace.from_config()\n",
+ "print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep = '\\n')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Register the model\n",
+ "Register an existing trained model, add descirption and tags."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "#Register the model\n",
+ "from azureml.core.model import Model\n",
+ "model = Model.register(model_path = \"sklearn_regression_model.pkl\", # this points to a local file\n",
+ " model_name = \"sklearn_regression_model.pkl\", # this is the name the model is registered as\n",
+ " tags = {'area': \"diabetes\", 'type': \"regression\"},\n",
+ " description = \"Ridge regression model to predict diabetes\",\n",
+ " workspace = ws)\n",
+ "\n",
+ "print(model.name, model.description, model.version)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Create an image\n",
+ "Create an image using the registered model the script that will load and run the model."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%writefile score.py\n",
+ "import pickle\n",
+ "import json\n",
+ "import numpy\n",
+ "from sklearn.externals import joblib\n",
+ "from sklearn.linear_model import Ridge\n",
+ "from azureml.core.model import Model\n",
+ "\n",
+ "def init():\n",
+ " global model\n",
+ " # note here \"sklearn_regression_model.pkl\" is the name of the model registered under\n",
+ " # this is a different behavior than before when the code is run locally, even though the code is the same.\n",
+ " model_path = Model.get_model_path('sklearn_regression_model.pkl')\n",
+ " # deserialize the model file back into a sklearn model\n",
+ " model = joblib.load(model_path)\n",
+ "\n",
+ "# note you can pass in multiple rows for scoring\n",
+ "def run(raw_data):\n",
+ " try:\n",
+ " data = json.loads(raw_data)['data']\n",
+ " data = numpy.array(data)\n",
+ " result = model.predict(data)\n",
+ " # you can return any data type as long as it is JSON-serializable\n",
+ " return result.tolist()\n",
+ " except Exception as e:\n",
+ " error = str(e)\n",
+ " return error"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.conda_dependencies import CondaDependencies \n",
+ "\n",
+ "myenv = CondaDependencies.create(conda_packages=['numpy','scikit-learn'])\n",
+ "\n",
+ "with open(\"myenv.yml\",\"w\") as f:\n",
+ " f.write(myenv.serialize_to_string())"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.image import ContainerImage\n",
+ "\n",
+ "image_config = ContainerImage.image_configuration(execution_script = \"score.py\",\n",
+ " runtime = \"python\",\n",
+ " conda_file = \"myenv.yml\",\n",
+ " description = \"Image with ridge regression model\",\n",
+ " tags = {'area': \"diabetes\", 'type': \"regression\"}\n",
+ " )\n",
+ "\n",
+ "image = ContainerImage.create(name = \"myimage1\",\n",
+ " # this is the model object\n",
+ " models = [model],\n",
+ " image_config = image_config,\n",
+ " workspace = ws)\n",
+ "\n",
+ "image.wait_for_creation(show_output = True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Provision the AKS Cluster\n",
+ "This is a one time setup. You can reuse this cluster for multiple deployments after it has been created. If you delete the cluster or the resource group that contains it, then you would have to recreate it."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Use the default configuration (can also provide parameters to customize)\n",
+ "prov_config = AksCompute.provisioning_configuration()\n",
+ "\n",
+ "aks_name = 'my-aks-9' \n",
+ "# Create the cluster\n",
+ "aks_target = ComputeTarget.create(workspace = ws, \n",
+ " name = aks_name, \n",
+ " provisioning_configuration = prov_config)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%time\n",
+ "aks_target.wait_for_completion(show_output = True)\n",
+ "print(aks_target.provisioning_state)\n",
+ "print(aks_target.provisioning_errors)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Optional step: Attach existing AKS cluster\n",
+ "\n",
+ "If you have existing AKS cluster in your Azure subscription, you can attach it to the Workspace."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "'''\n",
+ "# Use the default configuration (can also provide parameters to customize)\n",
+ "resource_id = '/subscriptions/92c76a2f-0e1c-4216-b65e-abf7a3f34c1e/resourcegroups/raymondsdk0604/providers/Microsoft.ContainerService/managedClusters/my-aks-0605d37425356b7d01'\n",
+ "\n",
+ "create_name='my-existing-aks' \n",
+ "# Create the cluster\n",
+ "attach_config = AksCompute.attach_configuration(resource_id=resource_id)\n",
+ "aks_target = ComputeTarget.attach(workspace=ws, name=create_name, attach_configuration=attach_config)\n",
+ "# Wait for the operation to complete\n",
+ "aks_target.wait_for_completion(True)\n",
+ "'''"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Deploy web service to AKS"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "#Set the web service configuration (using default here)\n",
+ "aks_config = AksWebservice.deploy_configuration()"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%time\n",
+ "aks_service_name ='aks-service-1'\n",
+ "\n",
+ "aks_service = Webservice.deploy_from_image(workspace = ws, \n",
+ " name = aks_service_name,\n",
+ " image = image,\n",
+ " deployment_config = aks_config,\n",
+ " deployment_target = aks_target)\n",
+ "aks_service.wait_for_deployment(show_output = True)\n",
+ "print(aks_service.state)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Test the web service\n",
+ "We test the web sevice by passing data."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%time\n",
+ "import json\n",
+ "\n",
+ "test_sample = json.dumps({'data': [\n",
+ " [1,2,3,4,5,6,7,8,9,10], \n",
+ " [10,9,8,7,6,5,4,3,2,1]\n",
+ "]})\n",
+ "test_sample = bytes(test_sample,encoding = 'utf8')\n",
+ "\n",
+ "prediction = aks_service.run(input_data = test_sample)\n",
+ "print(prediction)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Clean up\n",
+ "Delete the service, image and model."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%time\n",
+ "aks_service.delete()\n",
+ "image.delete()\n",
+ "model.delete()"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "raymondl"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.6"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/deployment/register-model-create-image-deploy-service/register-model-create-image-deploy-service.ipynb b/how-to-use-azureml/deployment/register-model-create-image-deploy-service/register-model-create-image-deploy-service.ipynb
index ef226a20..0afe9a3a 100644
--- a/how-to-use-azureml/deployment/register-model-create-image-deploy-service/register-model-create-image-deploy-service.ipynb
+++ b/how-to-use-azureml/deployment/register-model-create-image-deploy-service/register-model-create-image-deploy-service.ipynb
@@ -1,420 +1,420 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## 10. Register Model, Create Image and Deploy Service\n",
- "\n",
- "This example shows how to deploy a web service in step-by-step fashion:\n",
- "\n",
- " 1. Register model\n",
- " 2. Query versions of models and select one to deploy\n",
- " 3. Create Docker image\n",
- " 4. Query versions of images\n",
- " 5. Deploy the image as web service\n",
- " \n",
- "**IMPORTANT**:\n",
- " * This notebook requires you to first complete \"01.SDK-101-Train-and-Deploy-to-ACI.ipynb\" Notebook\n",
- " \n",
- "The 101 Notebook taught you how to deploy a web service directly from model in one step. This Notebook shows a more advanced approach that gives you more control over model versions and Docker image versions. "
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Prerequisites\n",
- "Make sure you go through the [00. Installation and Configuration](00.configuration.ipynb) Notebook first if you haven't."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Check core SDK version number\n",
- "import azureml.core\n",
- "\n",
- "print(\"SDK version:\", azureml.core.VERSION)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Initialize Workspace\n",
- "\n",
- "Initialize a workspace object from persisted configuration."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "create workspace"
- ]
- },
- "outputs": [],
- "source": [
- "from azureml.core import Workspace\n",
- "\n",
- "ws = Workspace.from_config()\n",
- "print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep = '\\n')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Register Model"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "You can add tags and descriptions to your models. Note you need to have a `sklearn_linreg_model.pkl` file in the current directory. This file is generated by the 01 notebook. The below call registers that file as a model with the same name `sklearn_linreg_model.pkl` in the workspace.\n",
- "\n",
- "Using tags, you can track useful information such as the name and version of the machine learning library used to train the model. Note that tags must be alphanumeric."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "register model from file"
- ]
- },
- "outputs": [],
- "source": [
- "from azureml.core.model import Model\n",
- "import sklearn\n",
- "\n",
- "library_version = \"sklearn\"+sklearn.__version__.replace(\".\",\"x\")\n",
- "\n",
- "model = Model.register(model_path = \"sklearn_regression_model.pkl\",\n",
- " model_name = \"sklearn_regression_model.pkl\",\n",
- " tags = {'area': \"diabetes\", 'type': \"regression\", 'version': library_version},\n",
- " description = \"Ridge regression model to predict diabetes\",\n",
- " workspace = ws)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "You can explore the registered models within your workspace and query by tag. Models are versioned. If you call the register_model command many times with same model name, you will get multiple versions of the model with increasing version numbers."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "register model from file"
- ]
- },
- "outputs": [],
- "source": [
- "regression_models = Model.list(workspace=ws, tags=['area'])\n",
- "for m in regression_models:\n",
- " print(\"Name:\", m.name,\"\\tVersion:\", m.version, \"\\tDescription:\", m.description, m.tags)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "You can pick a specific model to deploy"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "print(model.name, model.description, model.version, sep = '\\t')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create Docker Image"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Show `score.py`. Note that the `sklearn_regression_model.pkl` in the `get_model_path` call is referring to a model named `sklearn_linreg_model.pkl` registered under the workspace. It is NOT referenceing the local file."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%writefile score.py\n",
- "import pickle\n",
- "import json\n",
- "import numpy\n",
- "from sklearn.externals import joblib\n",
- "from sklearn.linear_model import Ridge\n",
- "from azureml.core.model import Model\n",
- "\n",
- "def init():\n",
- " global model\n",
- " # note here \"sklearn_regression_model.pkl\" is the name of the model registered under\n",
- " # this is a different behavior than before when the code is run locally, even though the code is the same.\n",
- " model_path = Model.get_model_path('sklearn_regression_model.pkl')\n",
- " # deserialize the model file back into a sklearn model\n",
- " model = joblib.load(model_path)\n",
- "\n",
- "# note you can pass in multiple rows for scoring\n",
- "def run(raw_data):\n",
- " try:\n",
- " data = json.loads(raw_data)['data']\n",
- " data = numpy.array(data)\n",
- " result = model.predict(data)\n",
- " # you can return any datatype as long as it is JSON-serializable\n",
- " return result.tolist()\n",
- " except Exception as e:\n",
- " error = str(e)\n",
- " return error"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.conda_dependencies import CondaDependencies \n",
- "\n",
- "myenv = CondaDependencies.create(conda_packages=['numpy','scikit-learn'])\n",
- "\n",
- "with open(\"myenv.yml\",\"w\") as f:\n",
- " f.write(myenv.serialize_to_string())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Note that following command can take few minutes. \n",
- "\n",
- "You can add tags and descriptions to images. Also, an image can contain multiple models."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "create image"
- ]
- },
- "outputs": [],
- "source": [
- "from azureml.core.image import Image, ContainerImage\n",
- "\n",
- "image_config = ContainerImage.image_configuration(runtime= \"python\",\n",
- " execution_script=\"score.py\",\n",
- " conda_file=\"myenv.yml\",\n",
- " tags = {'area': \"diabetes\", 'type': \"regression\"},\n",
- " description = \"Image with ridge regression model\")\n",
- "\n",
- "image = Image.create(name = \"myimage1\",\n",
- " # this is the model object \n",
- " models = [model],\n",
- " image_config = image_config, \n",
- " workspace = ws)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "create image"
- ]
- },
- "outputs": [],
- "source": [
- "image.wait_for_creation(show_output = True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "List images by tag and find out the detailed build log for debugging."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "create image"
- ]
- },
- "outputs": [],
- "source": [
- "for i in Image.list(workspace = ws,tags = [\"area\"]):\n",
- " print('{}(v.{} [{}]) stored at {} with build log {}'.format(i.name, i.version, i.creation_state, i.image_location, i.image_build_log_uri))"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Deploy image as web service on Azure Container Instance\n",
- "\n",
- "Note that the service creation can take few minutes."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "deploy service",
- "aci"
- ]
- },
- "outputs": [],
- "source": [
- "from azureml.core.webservice import AciWebservice\n",
- "\n",
- "aciconfig = AciWebservice.deploy_configuration(cpu_cores = 1, \n",
- " memory_gb = 1, \n",
- " tags = {'area': \"diabetes\", 'type': \"regression\"}, \n",
- " description = 'Predict diabetes using regression model')"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "deploy service",
- "aci"
- ]
- },
- "outputs": [],
- "source": [
- "from azureml.core.webservice import Webservice\n",
- "\n",
- "aci_service_name = 'my-aci-service-2'\n",
- "print(aci_service_name)\n",
- "aci_service = Webservice.deploy_from_image(deployment_config = aciconfig,\n",
- " image = image,\n",
- " name = aci_service_name,\n",
- " workspace = ws)\n",
- "aci_service.wait_for_deployment(True)\n",
- "print(aci_service.state)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Test web service"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Call the web service with some dummy input data to get a prediction."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "deploy service",
- "aci"
- ]
- },
- "outputs": [],
- "source": [
- "import json\n",
- "\n",
- "test_sample = json.dumps({'data': [\n",
- " [1,2,3,4,5,6,7,8,9,10], \n",
- " [10,9,8,7,6,5,4,3,2,1]\n",
- "]})\n",
- "test_sample = bytes(test_sample,encoding = 'utf8')\n",
- "\n",
- "prediction = aci_service.run(input_data=test_sample)\n",
- "print(prediction)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Delete ACI to clean up"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "deploy service",
- "aci"
- ]
- },
- "outputs": [],
- "source": [
- "aci_service.delete()"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "raymondl"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Register Model, Create Image and Deploy Service\n",
+ "\n",
+ "This example shows how to deploy a web service in step-by-step fashion:\n",
+ "\n",
+ " 1. Register model\n",
+ " 2. Query versions of models and select one to deploy\n",
+ " 3. Create Docker image\n",
+ " 4. Query versions of images\n",
+ " 5. Deploy the image as web service\n",
+ " \n",
+ "**IMPORTANT**:\n",
+ " * This notebook requires you to first complete [train-within-notebook](../../training/train-within-notebook/train-within-notebook.ipynb) example\n",
+ " \n",
+ "The train-within-notebook example taught you how to deploy a web service directly from model in one step. This Notebook shows a more advanced approach that gives you more control over model versions and Docker image versions. "
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Prerequisites\n",
+ "Make sure you go through the [configuration](../../../configuration.ipynb) Notebook first if you haven't."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Check core SDK version number\n",
+ "import azureml.core\n",
+ "\n",
+ "print(\"SDK version:\", azureml.core.VERSION)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Initialize Workspace\n",
+ "\n",
+ "Initialize a workspace object from persisted configuration."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "create workspace"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "from azureml.core import Workspace\n",
+ "\n",
+ "ws = Workspace.from_config()\n",
+ "print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep = '\\n')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Register Model"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "You can add tags and descriptions to your models. Note you need to have a `sklearn_linreg_model.pkl` file in the current directory. This file is generated by the 01 notebook. The below call registers that file as a model with the same name `sklearn_linreg_model.pkl` in the workspace.\n",
+ "\n",
+ "Using tags, you can track useful information such as the name and version of the machine learning library used to train the model. Note that tags must be alphanumeric."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "register model from file"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "from azureml.core.model import Model\n",
+ "import sklearn\n",
+ "\n",
+ "library_version = \"sklearn\"+sklearn.__version__.replace(\".\",\"x\")\n",
+ "\n",
+ "model = Model.register(model_path = \"sklearn_regression_model.pkl\",\n",
+ " model_name = \"sklearn_regression_model.pkl\",\n",
+ " tags = {'area': \"diabetes\", 'type': \"regression\", 'version': library_version},\n",
+ " description = \"Ridge regression model to predict diabetes\",\n",
+ " workspace = ws)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "You can explore the registered models within your workspace and query by tag. Models are versioned. If you call the register_model command many times with same model name, you will get multiple versions of the model with increasing version numbers."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "register model from file"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "regression_models = Model.list(workspace=ws, tags=['area'])\n",
+ "for m in regression_models:\n",
+ " print(\"Name:\", m.name,\"\\tVersion:\", m.version, \"\\tDescription:\", m.description, m.tags)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "You can pick a specific model to deploy"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "print(model.name, model.description, model.version, sep = '\\t')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create Docker Image"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Show `score.py`. Note that the `sklearn_regression_model.pkl` in the `get_model_path` call is referring to a model named `sklearn_linreg_model.pkl` registered under the workspace. It is NOT referenceing the local file."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%writefile score.py\n",
+ "import pickle\n",
+ "import json\n",
+ "import numpy\n",
+ "from sklearn.externals import joblib\n",
+ "from sklearn.linear_model import Ridge\n",
+ "from azureml.core.model import Model\n",
+ "\n",
+ "def init():\n",
+ " global model\n",
+ " # note here \"sklearn_regression_model.pkl\" is the name of the model registered under\n",
+ " # this is a different behavior than before when the code is run locally, even though the code is the same.\n",
+ " model_path = Model.get_model_path('sklearn_regression_model.pkl')\n",
+ " # deserialize the model file back into a sklearn model\n",
+ " model = joblib.load(model_path)\n",
+ "\n",
+ "# note you can pass in multiple rows for scoring\n",
+ "def run(raw_data):\n",
+ " try:\n",
+ " data = json.loads(raw_data)['data']\n",
+ " data = numpy.array(data)\n",
+ " result = model.predict(data)\n",
+ " # you can return any datatype as long as it is JSON-serializable\n",
+ " return result.tolist()\n",
+ " except Exception as e:\n",
+ " error = str(e)\n",
+ " return error"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.conda_dependencies import CondaDependencies \n",
+ "\n",
+ "myenv = CondaDependencies.create(conda_packages=['numpy','scikit-learn'])\n",
+ "\n",
+ "with open(\"myenv.yml\",\"w\") as f:\n",
+ " f.write(myenv.serialize_to_string())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Note that following command can take few minutes. \n",
+ "\n",
+ "You can add tags and descriptions to images. Also, an image can contain multiple models."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "create image"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "from azureml.core.image import Image, ContainerImage\n",
+ "\n",
+ "image_config = ContainerImage.image_configuration(runtime= \"python\",\n",
+ " execution_script=\"score.py\",\n",
+ " conda_file=\"myenv.yml\",\n",
+ " tags = {'area': \"diabetes\", 'type': \"regression\"},\n",
+ " description = \"Image with ridge regression model\")\n",
+ "\n",
+ "image = Image.create(name = \"myimage1\",\n",
+ " # this is the model object \n",
+ " models = [model],\n",
+ " image_config = image_config, \n",
+ " workspace = ws)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "create image"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "image.wait_for_creation(show_output = True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "List images by tag and find out the detailed build log for debugging."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "create image"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "for i in Image.list(workspace = ws,tags = [\"area\"]):\n",
+ " print('{}(v.{} [{}]) stored at {} with build log {}'.format(i.name, i.version, i.creation_state, i.image_location, i.image_build_log_uri))"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Deploy image as web service on Azure Container Instance\n",
+ "\n",
+ "Note that the service creation can take few minutes."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "deploy service",
+ "aci"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "from azureml.core.webservice import AciWebservice\n",
+ "\n",
+ "aciconfig = AciWebservice.deploy_configuration(cpu_cores = 1, \n",
+ " memory_gb = 1, \n",
+ " tags = {'area': \"diabetes\", 'type': \"regression\"}, \n",
+ " description = 'Predict diabetes using regression model')"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "deploy service",
+ "aci"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "from azureml.core.webservice import Webservice\n",
+ "\n",
+ "aci_service_name = 'my-aci-service-2'\n",
+ "print(aci_service_name)\n",
+ "aci_service = Webservice.deploy_from_image(deployment_config = aciconfig,\n",
+ " image = image,\n",
+ " name = aci_service_name,\n",
+ " workspace = ws)\n",
+ "aci_service.wait_for_deployment(True)\n",
+ "print(aci_service.state)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Test web service"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Call the web service with some dummy input data to get a prediction."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "deploy service",
+ "aci"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "import json\n",
+ "\n",
+ "test_sample = json.dumps({'data': [\n",
+ " [1,2,3,4,5,6,7,8,9,10], \n",
+ " [10,9,8,7,6,5,4,3,2,1]\n",
+ "]})\n",
+ "test_sample = bytes(test_sample,encoding = 'utf8')\n",
+ "\n",
+ "prediction = aci_service.run(input_data=test_sample)\n",
+ "print(prediction)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Delete ACI to clean up"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "deploy service",
+ "aci"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "aci_service.delete()"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "raymondl"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.6"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/machine-learning-pipelines/aml-pipelines-concept.png b/how-to-use-azureml/machine-learning-pipelines/aml-pipelines-concept.png
deleted file mode 100644
index b01526da..00000000
Binary files a/how-to-use-azureml/machine-learning-pipelines/aml-pipelines-concept.png and /dev/null differ
diff --git a/how-to-use-azureml/machine-learning-pipelines/intro-to-pipelines/aml-pipelines-data-transfer.ipynb b/how-to-use-azureml/machine-learning-pipelines/intro-to-pipelines/aml-pipelines-data-transfer.ipynb
index 70ee7677..640e879d 100644
--- a/how-to-use-azureml/machine-learning-pipelines/intro-to-pipelines/aml-pipelines-data-transfer.ipynb
+++ b/how-to-use-azureml/machine-learning-pipelines/intro-to-pipelines/aml-pipelines-data-transfer.ipynb
@@ -1,332 +1,332 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved. \n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Azure Machine Learning Pipeline with DataTranferStep\n",
- "This notebook is used to demonstrate the use of DataTranferStep in Azure Machine Learning Pipeline.\n",
- "\n",
- "In certain cases, you will need to transfer data from one data location to another. For example, your data may be in Files storage and you may want to move it to Blob storage. Or, if your data is in an ADLS account and you want to make it available in the Blob storage. The built-in **DataTransferStep** class helps you transfer data in these situations.\n",
- "\n",
- "The below example shows how to move data in an ADLS account to Blob storage."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Azure Machine Learning and Pipeline SDK-specific imports"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import os\n",
- "import azureml.core\n",
- "from azureml.core.compute import ComputeTarget, DatabricksCompute, DataFactoryCompute\n",
- "from azureml.exceptions import ComputeTargetException\n",
- "from azureml.core import Workspace, Run, Experiment\n",
- "from azureml.pipeline.core import Pipeline, PipelineData\n",
- "from azureml.pipeline.steps import AdlaStep\n",
- "from azureml.core.datastore import Datastore\n",
- "from azureml.data.data_reference import DataReference\n",
- "from azureml.data.sql_data_reference import SqlDataReference\n",
- "from azureml.core import attach_legacy_compute_target\n",
- "from azureml.data.stored_procedure_parameter import StoredProcedureParameter, StoredProcedureParameterType\n",
- "from azureml.pipeline.steps import DataTransferStep\n",
- "\n",
- "# Check core SDK version number\n",
- "print(\"SDK version:\", azureml.core.VERSION)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Initialize Workspace\n",
- "\n",
- "Initialize a workspace object from persisted configuration. Make sure the config file is present at .\\config.json\n",
- "\n",
- "If you don't have a config.json file, please go through the configuration Notebook located here:\n",
- "https://github.com/Azure/MachineLearningNotebooks. \n",
- "\n",
- "This sets you up with a working config file that has information on your workspace, subscription id, etc. "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "create workspace"
- ]
- },
- "outputs": [],
- "source": [
- "ws = Workspace.from_config()\n",
- "print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep = '\\n')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Register Datastores\n",
- "\n",
- "In the code cell below, you will need to fill in the appropriate values for the workspace name, datastore name, subscription id, resource group, store name, tenant id, client id, and client secret that are associated with your ADLS datastore. \n",
- "\n",
- "For background on registering your data store, consult this article:\n",
- "\n",
- "https://docs.microsoft.com/en-us/azure/data-lake-store/data-lake-store-service-to-service-authenticate-using-active-directory"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "\n",
- "workspace = ws.name\n",
- "datastore_name='MyAdlsDatastore'\n",
- "subscription_id=os.getenv(\"ADL_SUBSCRIPTION_62\", \"\") # subscription id of ADLS account\n",
- "resource_group=os.getenv(\"ADL_RESOURCE_GROUP_62\", \"\") # resource group of ADLS account\n",
- "store_name=os.getenv(\"ADL_STORENAME_62\", \"\") # ADLS account name\n",
- "tenant_id=os.getenv(\"ADL_TENANT_62\", \"\") # tenant id of service principal\n",
- "client_id=os.getenv(\"ADL_CLIENTID_62\", \"\") # client id of service principal\n",
- "client_secret=os.getenv(\"ADL_CLIENT_SECRET_62\", \"\") # the secret of service principal\n",
- "\n",
- "try:\n",
- " adls_datastore = Datastore.get(ws, datastore_name)\n",
- " print(\"found datastore with name: %s\" % datastore_name)\n",
- "except:\n",
- " adls_datastore = Datastore.register_azure_data_lake(\n",
- " workspace=ws,\n",
- " datastore_name=datastore_name,\n",
- " subscription_id=subscription_id, # subscription id of ADLS account\n",
- " resource_group=resource_group, # resource group of ADLS account\n",
- " store_name=store_name, # ADLS account name\n",
- " tenant_id=tenant_id, # tenant id of service principal\n",
- " client_id=client_id, # client id of service principal\n",
- " client_secret=client_secret) # the secret of service principal\n",
- " print(\"registered datastore with name: %s\" % datastore_name)\n",
- "\n",
- "\n",
- "\n",
- "blob_datastore_name='MyBlobDatastore'\n",
- "account_name=os.getenv(\"BLOB_ACCOUNTNAME_62\", \"\") # Storage account name\n",
- "container_name=os.getenv(\"BLOB_CONTAINER_62\", \"\") # Name of Azure blob container\n",
- "account_key=os.getenv(\"BLOB_ACCOUNT_KEY_62\", \"\") # Storage account key\n",
- "\n",
- "try:\n",
- " blob_datastore = Datastore.get(ws, blob_datastore_name)\n",
- " print(\"found blob datastore with name: %s\" % blob_datastore_name)\n",
- "except:\n",
- " blob_datastore = Datastore.register_azure_blob_container(\n",
- " workspace=ws,\n",
- " datastore_name=blob_datastore_name,\n",
- " account_name=account_name, # Storage account name\n",
- " container_name=container_name, # Name of Azure blob container\n",
- " account_key=account_key) # Storage account key\"\n",
- " print(\"registered blob datastore with name: %s\" % blob_datastore_name)\n",
- "\n",
- "# CLI:\n",
- "# az ml datastore register-blob -n -a -c -k [-t ]"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Create DataReferences"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "adls_datastore = Datastore(workspace=ws, name=\"MyAdlsDatastore\")\n",
- "\n",
- "# adls\n",
- "adls_data_ref = DataReference(\n",
- " datastore=adls_datastore,\n",
- " data_reference_name=\"adls_test_data\",\n",
- " path_on_datastore=\"testdata\")\n",
- "\n",
- "blob_datastore = Datastore(workspace=ws, name=\"MyBlobDatastore\")\n",
- "\n",
- "# blob data\n",
- "blob_data_ref = DataReference(\n",
- " datastore=blob_datastore,\n",
- " data_reference_name=\"blob_test_data\",\n",
- " path_on_datastore=\"testdata\")\n",
- "\n",
- "print(\"obtained adls, blob data references\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Setup Data Factory Account"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "data_factory_name = 'adftest'\n",
- "\n",
- "def get_or_create_data_factory(workspace, factory_name):\n",
- " try:\n",
- " return DataFactoryCompute(workspace, factory_name)\n",
- " except ComputeTargetException as e:\n",
- " if 'ComputeTargetNotFound' in e.message:\n",
- " print('Data factory not found, creating...')\n",
- " provisioning_config = DataFactoryCompute.provisioning_configuration()\n",
- " data_factory = ComputeTarget.create(workspace, factory_name, provisioning_config)\n",
- " data_factory.wait_for_completion()\n",
- " return data_factory\n",
- " else:\n",
- " raise e\n",
- " \n",
- "data_factory_compute = get_or_create_data_factory(ws, data_factory_name)\n",
- "\n",
- "print(\"setup data factory account complete\")\n",
- "\n",
- "# CLI:\n",
- "# Create: az ml computetarget setup datafactory -n \n",
- "# BYOC: az ml computetarget attach datafactory -n -i "
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Create a DataTransferStep"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "**DataTransferStep** is used to transfer data between Azure Blob, Azure Data Lake Store, and Azure SQL database.\n",
- "\n",
- "- **name:** Name of module\n",
- "- **source_data_reference:** Input connection that serves as source of data transfer operation.\n",
- "- **destination_data_reference:** Input connection that serves as destination of data transfer operation.\n",
- "- **compute_target:** Azure Data Factory to use for transferring data.\n",
- "- **allow_reuse:** Whether the step should reuse results of previous DataTransferStep when run with same inputs. Set as False to force data to be transferred again.\n",
- "\n",
- "Optional arguments to explicitly specify whether a path corresponds to a file or a directory. These are useful when storage contains both file and directory with the same name or when creating a new destination path.\n",
- "\n",
- "- **source_reference_type:** An optional string specifying the type of source_data_reference. Possible values include: 'file', 'directory'. When not specified, we use the type of existing path or directory if it's a new path.\n",
- "- **destination_reference_type:** An optional string specifying the type of destination_data_reference. Possible values include: 'file', 'directory'. When not specified, we use the type of existing path or directory if it's a new path."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "transfer_adls_to_blob = DataTransferStep(\n",
- " name=\"transfer_adls_to_blob\",\n",
- " source_data_reference=adls_data_ref,\n",
- " destination_data_reference=blob_data_ref,\n",
- " compute_target=data_factory_compute)\n",
- "\n",
- "print(\"data transfer step created\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Build and Submit the Experiment"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "pipeline = Pipeline(\n",
- " description=\"data_transfer_101\",\n",
- " workspace=ws,\n",
- " steps=[transfer_adls_to_blob])\n",
- "\n",
- "pipeline_run = Experiment(ws, \"Data_Transfer_example\").submit(pipeline)\n",
- "pipeline_run.wait_for_completion()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### View Run Details"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.widgets import RunDetails\n",
- "RunDetails(pipeline_run).show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Next: Databricks as a Compute Target\n",
- "To use Databricks as a compute target from Azure Machine Learning Pipeline, a DatabricksStep is used. This [notebook](./aml-pipelines-use-databricks-as-compute-target.ipynb) demonstrates the use of a DatabricksStep in an Azure Machine Learning Pipeline."
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "diray"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved. \n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Azure Machine Learning Pipeline with DataTranferStep\n",
+ "This notebook is used to demonstrate the use of DataTranferStep in Azure Machine Learning Pipeline.\n",
+ "\n",
+ "In certain cases, you will need to transfer data from one data location to another. For example, your data may be in Files storage and you may want to move it to Blob storage. Or, if your data is in an ADLS account and you want to make it available in the Blob storage. The built-in **DataTransferStep** class helps you transfer data in these situations.\n",
+ "\n",
+ "The below example shows how to move data in an ADLS account to Blob storage."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Azure Machine Learning and Pipeline SDK-specific imports"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import os\n",
+ "import azureml.core\n",
+ "from azureml.core.compute import ComputeTarget, DatabricksCompute, DataFactoryCompute\n",
+ "from azureml.exceptions import ComputeTargetException\n",
+ "from azureml.core import Workspace, Run, Experiment\n",
+ "from azureml.pipeline.core import Pipeline, PipelineData\n",
+ "from azureml.pipeline.steps import AdlaStep\n",
+ "from azureml.core.datastore import Datastore\n",
+ "from azureml.data.data_reference import DataReference\n",
+ "from azureml.data.sql_data_reference import SqlDataReference\n",
+ "from azureml.core import attach_legacy_compute_target\n",
+ "from azureml.data.stored_procedure_parameter import StoredProcedureParameter, StoredProcedureParameterType\n",
+ "from azureml.pipeline.steps import DataTransferStep\n",
+ "\n",
+ "# Check core SDK version number\n",
+ "print(\"SDK version:\", azureml.core.VERSION)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Initialize Workspace\n",
+ "\n",
+ "Initialize a workspace object from persisted configuration. Make sure the config file is present at .\\config.json\n",
+ "\n",
+ "If you don't have a config.json file, please go through the configuration Notebook located here:\n",
+ "https://github.com/Azure/MachineLearningNotebooks. \n",
+ "\n",
+ "This sets you up with a working config file that has information on your workspace, subscription id, etc. "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "create workspace"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "ws = Workspace.from_config()\n",
+ "print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep = '\\n')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Register Datastores\n",
+ "\n",
+ "In the code cell below, you will need to fill in the appropriate values for the workspace name, datastore name, subscription id, resource group, store name, tenant id, client id, and client secret that are associated with your ADLS datastore. \n",
+ "\n",
+ "For background on registering your data store, consult this article:\n",
+ "\n",
+ "https://docs.microsoft.com/en-us/azure/data-lake-store/data-lake-store-service-to-service-authenticate-using-active-directory"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "\n",
+ "workspace = ws.name\n",
+ "datastore_name='MyAdlsDatastore'\n",
+ "subscription_id=os.getenv(\"ADL_SUBSCRIPTION_62\", \"\") # subscription id of ADLS account\n",
+ "resource_group=os.getenv(\"ADL_RESOURCE_GROUP_62\", \"\") # resource group of ADLS account\n",
+ "store_name=os.getenv(\"ADL_STORENAME_62\", \"\") # ADLS account name\n",
+ "tenant_id=os.getenv(\"ADL_TENANT_62\", \"\") # tenant id of service principal\n",
+ "client_id=os.getenv(\"ADL_CLIENTID_62\", \"\") # client id of service principal\n",
+ "client_secret=os.getenv(\"ADL_CLIENT_SECRET_62\", \"\") # the secret of service principal\n",
+ "\n",
+ "try:\n",
+ " adls_datastore = Datastore.get(ws, datastore_name)\n",
+ " print(\"found datastore with name: %s\" % datastore_name)\n",
+ "except:\n",
+ " adls_datastore = Datastore.register_azure_data_lake(\n",
+ " workspace=ws,\n",
+ " datastore_name=datastore_name,\n",
+ " subscription_id=subscription_id, # subscription id of ADLS account\n",
+ " resource_group=resource_group, # resource group of ADLS account\n",
+ " store_name=store_name, # ADLS account name\n",
+ " tenant_id=tenant_id, # tenant id of service principal\n",
+ " client_id=client_id, # client id of service principal\n",
+ " client_secret=client_secret) # the secret of service principal\n",
+ " print(\"registered datastore with name: %s\" % datastore_name)\n",
+ "\n",
+ "\n",
+ "\n",
+ "blob_datastore_name='MyBlobDatastore'\n",
+ "account_name=os.getenv(\"BLOB_ACCOUNTNAME_62\", \"\") # Storage account name\n",
+ "container_name=os.getenv(\"BLOB_CONTAINER_62\", \"\") # Name of Azure blob container\n",
+ "account_key=os.getenv(\"BLOB_ACCOUNT_KEY_62\", \"\") # Storage account key\n",
+ "\n",
+ "try:\n",
+ " blob_datastore = Datastore.get(ws, blob_datastore_name)\n",
+ " print(\"found blob datastore with name: %s\" % blob_datastore_name)\n",
+ "except:\n",
+ " blob_datastore = Datastore.register_azure_blob_container(\n",
+ " workspace=ws,\n",
+ " datastore_name=blob_datastore_name,\n",
+ " account_name=account_name, # Storage account name\n",
+ " container_name=container_name, # Name of Azure blob container\n",
+ " account_key=account_key) # Storage account key\"\n",
+ " print(\"registered blob datastore with name: %s\" % blob_datastore_name)\n",
+ "\n",
+ "# CLI:\n",
+ "# az ml datastore register-blob -n -a -c -k [-t ]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Create DataReferences"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "adls_datastore = Datastore(workspace=ws, name=\"MyAdlsDatastore\")\n",
+ "\n",
+ "# adls\n",
+ "adls_data_ref = DataReference(\n",
+ " datastore=adls_datastore,\n",
+ " data_reference_name=\"adls_test_data\",\n",
+ " path_on_datastore=\"testdata\")\n",
+ "\n",
+ "blob_datastore = Datastore(workspace=ws, name=\"MyBlobDatastore\")\n",
+ "\n",
+ "# blob data\n",
+ "blob_data_ref = DataReference(\n",
+ " datastore=blob_datastore,\n",
+ " data_reference_name=\"blob_test_data\",\n",
+ " path_on_datastore=\"testdata\")\n",
+ "\n",
+ "print(\"obtained adls, blob data references\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Setup Data Factory Account"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "data_factory_name = 'adftest'\n",
+ "\n",
+ "def get_or_create_data_factory(workspace, factory_name):\n",
+ " try:\n",
+ " return DataFactoryCompute(workspace, factory_name)\n",
+ " except ComputeTargetException as e:\n",
+ " if 'ComputeTargetNotFound' in e.message:\n",
+ " print('Data factory not found, creating...')\n",
+ " provisioning_config = DataFactoryCompute.provisioning_configuration()\n",
+ " data_factory = ComputeTarget.create(workspace, factory_name, provisioning_config)\n",
+ " data_factory.wait_for_completion()\n",
+ " return data_factory\n",
+ " else:\n",
+ " raise e\n",
+ " \n",
+ "data_factory_compute = get_or_create_data_factory(ws, data_factory_name)\n",
+ "\n",
+ "print(\"setup data factory account complete\")\n",
+ "\n",
+ "# CLI:\n",
+ "# Create: az ml computetarget setup datafactory -n \n",
+ "# BYOC: az ml computetarget attach datafactory -n -i "
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Create a DataTransferStep"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "**DataTransferStep** is used to transfer data between Azure Blob, Azure Data Lake Store, and Azure SQL database.\n",
+ "\n",
+ "- **name:** Name of module\n",
+ "- **source_data_reference:** Input connection that serves as source of data transfer operation.\n",
+ "- **destination_data_reference:** Input connection that serves as destination of data transfer operation.\n",
+ "- **compute_target:** Azure Data Factory to use for transferring data.\n",
+ "- **allow_reuse:** Whether the step should reuse results of previous DataTransferStep when run with same inputs. Set as False to force data to be transferred again.\n",
+ "\n",
+ "Optional arguments to explicitly specify whether a path corresponds to a file or a directory. These are useful when storage contains both file and directory with the same name or when creating a new destination path.\n",
+ "\n",
+ "- **source_reference_type:** An optional string specifying the type of source_data_reference. Possible values include: 'file', 'directory'. When not specified, we use the type of existing path or directory if it's a new path.\n",
+ "- **destination_reference_type:** An optional string specifying the type of destination_data_reference. Possible values include: 'file', 'directory'. When not specified, we use the type of existing path or directory if it's a new path."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "transfer_adls_to_blob = DataTransferStep(\n",
+ " name=\"transfer_adls_to_blob\",\n",
+ " source_data_reference=adls_data_ref,\n",
+ " destination_data_reference=blob_data_ref,\n",
+ " compute_target=data_factory_compute)\n",
+ "\n",
+ "print(\"data transfer step created\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Build and Submit the Experiment"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "pipeline = Pipeline(\n",
+ " description=\"data_transfer_101\",\n",
+ " workspace=ws,\n",
+ " steps=[transfer_adls_to_blob])\n",
+ "\n",
+ "pipeline_run = Experiment(ws, \"Data_Transfer_example\").submit(pipeline)\n",
+ "pipeline_run.wait_for_completion()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### View Run Details"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.widgets import RunDetails\n",
+ "RunDetails(pipeline_run).show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Next: Databricks as a Compute Target\n",
+ "To use Databricks as a compute target from Azure Machine Learning Pipeline, a DatabricksStep is used. This [notebook](./aml-pipelines-use-databricks-as-compute-target.ipynb) demonstrates the use of a DatabricksStep in an Azure Machine Learning Pipeline."
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3",
- "language": "python",
- "name": "python3"
+ "metadata": {
+ "authors": [
+ {
+ "name": "diray"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.7"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.7"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/machine-learning-pipelines/intro-to-pipelines/aml-pipelines-getting-started.ipynb b/how-to-use-azureml/machine-learning-pipelines/intro-to-pipelines/aml-pipelines-getting-started.ipynb
index a6965896..4bb7fd68 100644
--- a/how-to-use-azureml/machine-learning-pipelines/intro-to-pipelines/aml-pipelines-getting-started.ipynb
+++ b/how-to-use-azureml/machine-learning-pipelines/intro-to-pipelines/aml-pipelines-getting-started.ipynb
@@ -1,606 +1,606 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved. \n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Azure Machine Learning Pipelines: Getting Started\n",
- "\n",
- "## Overview\n",
- "\n",
- "Read [Azure Machine Learning Pipelines](https://docs.microsoft.com/en-us/azure/machine-learning/service/concept-ml-pipelines) overview, or the [readme article](../README.md) on Azure Machine Learning Pipelines to get more information.\n",
- " \n",
- "\n",
- "This Notebook shows basic construction of a **pipeline** that runs jobs unattended in different compute clusters. "
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Prerequisites and Azure Machine Learning Basics\n",
- "Make sure you go through the configuration Notebook located at https://github.com/Azure/MachineLearningNotebooks first if you haven't. This sets you up with a working config file that has information on your workspace, subscription id, etc. \n"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Azure Machine Learning Imports\n",
- "\n",
- "In this first code cell, we import key Azure Machine Learning modules that we will use below. "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import azureml.core\n",
- "from azureml.core import Workspace, Run, Experiment, Datastore\n",
- "from azureml.core.compute import AmlCompute\n",
- "from azureml.core.compute import ComputeTarget\n",
- "from azureml.core.compute import DataFactoryCompute\n",
- "from azureml.widgets import RunDetails\n",
- "\n",
- "# Check core SDK version number\n",
- "print(\"SDK version:\", azureml.core.VERSION)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Pipeline-specific SDK imports\n",
- "\n",
- "Here, we import key pipeline modules, whose use will be illustrated in the examples below."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.data.data_reference import DataReference\n",
- "from azureml.pipeline.core import Pipeline, PipelineData, StepSequence\n",
- "from azureml.pipeline.steps import PythonScriptStep\n",
- "from azureml.pipeline.steps import DataTransferStep\n",
- "from azureml.pipeline.core import PublishedPipeline\n",
- "from azureml.pipeline.core.graph import PipelineParameter\n",
- "\n",
- "print(\"Pipeline SDK-specific imports completed\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Initialize Workspace\n",
- "\n",
- "Initialize a [workspace](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.workspace(class%29) object from persisted configuration."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "create workspace"
- ]
- },
- "outputs": [],
- "source": [
- "ws = Workspace.from_config()\n",
- "print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep = '\\n')\n",
- "\n",
- "# Default datastore (Azure file storage)\n",
- "def_file_store = ws.get_default_datastore() \n",
- "# The above call is equivalent to Datastore(ws, \"workspacefilestore\") or simply Datastore(ws)\n",
- "print(\"Default datastore's name: {}\".format(def_file_store.name))\n",
- "\n",
- "# Blob storage associated with the workspace\n",
- "# The following call GETS the Azure Blob Store associated with your workspace.\n",
- "# Note that workspaceblobstore is **the name of this store and CANNOT BE CHANGED and must be used as is** \n",
- "def_blob_store = Datastore(ws, \"workspaceblobstore\")\n",
- "print(\"Blobstore's name: {}\".format(def_blob_store.name))"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# project folder\n",
- "project_folder = '.'\n",
- " \n",
- "print('Sample projects will be created in {}.'.format(project_folder))"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Required data and script files for the the tutorial\n",
- "Sample files required to finish this tutorial are already copied to the project folder specified above. Even though the .py provided in the samples don't have much \"ML work,\" as a data scientist, you will work on this extensively as part of your work. To complete this tutorial, the contents of these files are not very important. The one-line files are for demostration purpose only."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Datastore concepts\n",
- "A [Datastore](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.datastore(class) is a place where data can be stored that is then made accessible to a compute either by means of mounting or copying the data to the compute target. \n",
- "\n",
- "A Datastore can either be backed by an Azure File Storage (default) or by an Azure Blob Storage.\n",
- "\n",
- "In this next step, we will upload the training and test set into the workspace's default storage (File storage), and another piece of data to Azure Blob Storage. When to use [Azure Blobs](https://docs.microsoft.com/en-us/azure/storage/blobs/storage-blobs-introduction), [Azure Files](https://docs.microsoft.com/en-us/azure/storage/files/storage-files-introduction), or [Azure Disks](https://docs.microsoft.com/en-us/azure/virtual-machines/linux/managed-disks-overview) is [detailed here](https://docs.microsoft.com/en-us/azure/storage/common/storage-decide-blobs-files-disks).\n",
- "\n",
- "**Please take good note of the concept of the datastore.**"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Upload data to default datastore\n",
- "Default datastore on workspace is the Azure File storage. The workspace has a Blob storage associated with it as well. Let's upload a file to each of these storages."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# get_default_datastore() gets the default Azure File Store associated with your workspace.\n",
- "# Here we are reusing the def_file_store object we obtained earlier\n",
- "\n",
- "# target_path is the directory at the destination\n",
- "def_file_store.upload_files(['./20news.pkl'], \n",
- " target_path = '20newsgroups', \n",
- " overwrite = True, \n",
- " show_progress = True)\n",
- "\n",
- "# Here we are reusing the def_blob_store we created earlier\n",
- "def_blob_store.upload_files([\"./20news.pkl\"], target_path=\"20newsgroups\", overwrite=True)\n",
- "\n",
- "print(\"Upload calls completed\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### (Optional) See your files using Azure Portal\n",
- "Once you successfully uploaded the files, you can browse to them (or upload more files) using [Azure Portal](https://portal.azure.com). At the portal, make sure you have selected **AzureML Nursery** as your subscription (click *Resource Groups* and then select the subscription). Then look for your **Machine Learning Workspace** (it has your *alias* as the name). It has a link to your storage. Click on the storage link. It will take you to a page where you can see [Blobs](https://docs.microsoft.com/en-us/azure/storage/blobs/storage-blobs-introduction), [Files](https://docs.microsoft.com/en-us/azure/storage/files/storage-files-introduction), [Tables](https://docs.microsoft.com/en-us/azure/storage/tables/table-storage-overview), and [Queues](https://docs.microsoft.com/en-us/azure/storage/queues/storage-queues-introduction). We have just uploaded a file to the Blob storage and another one to the File storage. You should be able to see both of these files in their respective locations. "
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Compute Targets\n",
- "A compute target specifies where to execute your program such as a remote Docker on a VM, or a cluster. A compute target needs to be addressable and accessible by you.\n",
- "\n",
- "**You need at least one compute target to send your payload to. We are planning to use Azure Machine Learning Compute exclusively for this tutorial for all steps. However in some cases you may require multiple compute targets as some steps may run in one compute target like Azure Machine Learning Compute, and some other steps in the same pipeline could run in a different compute target.**\n",
- "\n",
- "*The example belows show creating/retrieving/attaching to an Azure Machine Learning Compute instance.*"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### List of Compute Targets on the workspace"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "cts = ws.compute_targets\n",
- "for ct in cts:\n",
- " print(ct)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Retrieve or create a Azure Machine Learning compute\n",
- "Azure Machine Learning Compute is a service for provisioning and managing clusters of Azure virtual machines for running machine learning workloads. Let's create a new Azure Machine Learning Compute in the current workspace, if it doesn't already exist. We will then run the training script on this compute target.\n",
- "\n",
- "If we could not find the compute with the given name in the previous cell, then we will create a new compute here. We will create an Azure Machine Learning Compute containing **STANDARD_D2_V2 CPU VMs**. This process is broken down into the following steps:\n",
- "\n",
- "1. Create the configuration\n",
- "2. Create the Azure Machine Learning compute\n",
- "\n",
- "**This process will take about 3 minutes and is providing only sparse output in the process. Please make sure to wait until the call returns before moving to the next cell.**"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "\n",
- "aml_compute_target = \"aml-compute\"\n",
- "try:\n",
- " aml_compute = AmlCompute(ws, aml_compute_target)\n",
- " print(\"found existing compute target.\")\n",
- "except:\n",
- " print(\"creating new compute target\")\n",
- " \n",
- " provisioning_config = AmlCompute.provisioning_configuration(vm_size = \"STANDARD_D2_V2\",\n",
- " min_nodes = 1, \n",
- " max_nodes = 4) \n",
- " aml_compute = ComputeTarget.create(ws, aml_compute_target, provisioning_config)\n",
- " aml_compute.wait_for_completion(show_output=True, min_node_count=None, timeout_in_minutes=20)\n",
- " \n",
- "print(\"Azure Machine Learning Compute attached\")\n"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# For a more detailed view of current Azure Machine Learning Compute status, use the 'status' property\n",
- "# example: un-comment the following line.\n",
- "# print(aml_compute.status.serialize())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "**Wait for this call to finish before proceeding (you will see the asterisk turning to a number).**\n",
- "\n",
- "Now that you have created the compute target, let's see what the workspace's compute_targets() function returns. You should now see one entry named 'amlcompute' of type AmlCompute."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "**Now that we have completed learning the basics of Azure Machine Learning (AML), let's go ahead and start understanding the Pipeline concepts.**"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Creating a Step in a Pipeline\n",
- "A Step is a unit of execution. Step typically needs a target of execution (compute target), a script to execute, and may require script arguments and inputs, and can produce outputs. The step also could take a number of other parameters. Azure Machine Learning Pipelines provides the following built-in Steps:\n",
- "\n",
- "- [**PythonScriptStep**](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-steps/azureml.pipeline.steps.python_script_step.pythonscriptstep?view=azure-ml-py): Add a step to run a Python script in a Pipeline.\n",
- "- [**AdlaStep**](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-steps/azureml.pipeline.steps.adla_step.adlastep?view=azure-ml-py): Adds a step to run U-SQL script using Azure Data Lake Analytics.\n",
- "- [**DataTransferStep**](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-steps/azureml.pipeline.steps.data_transfer_step.datatransferstep?view=azure-ml-py): Transfers data between Azure Blob and Data Lake accounts.\n",
- "- [**DatabricksStep**](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-steps/azureml.pipeline.steps.databricks_step.databricksstep?view=azure-ml-py): Adds a DataBricks notebook as a step in a Pipeline.\n",
- "- [**HyperDriveStep**](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-steps/azureml.pipeline.steps.hyper_drive_step.hyperdrivestep?view=azure-ml-py): Creates a Hyper Drive step for Hyper Parameter Tuning in a Pipeline.\n",
- "\n",
- "The following code will create a PythonScriptStep to be executed in the Azure Machine Learning Compute we created above using train.py, one of the files already made available in the project folder.\n",
- "\n",
- "A **PythonScriptStep** is a basic, built-in step to run a Python Script on a compute target. It takes a script name and optionally other parameters like arguments for the script, compute target, inputs and outputs. If no compute target is specified, default compute target for the workspace is used."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Uses default values for PythonScriptStep construct.\n",
- "\n",
- "# Syntax\n",
- "# PythonScriptStep(\n",
- "# script_name, \n",
- "# name=None, \n",
- "# arguments=None, \n",
- "# compute_target=None, \n",
- "# runconfig=None, \n",
- "# inputs=None, \n",
- "# outputs=None, \n",
- "# params=None, \n",
- "# source_directory=None, \n",
- "# allow_reuse=True, \n",
- "# version=None, \n",
- "# hash_paths=None)\n",
- "# This returns a Step\n",
- "step1 = PythonScriptStep(name=\"train_step\",\n",
- " script_name=\"train.py\", \n",
- " compute_target=aml_compute, \n",
- " source_directory=project_folder,\n",
- " allow_reuse=False)\n",
- "print(\"Step1 created\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "**Note:** In the above call to PythonScriptStep(), the flag *allow_reuse* determines whether the step should reuse previous results when run with the same settings/inputs. This flag's default value is *True*; the default is set to *True* because, when inputs and parameters have not changed, we typically do not want to re-run a given pipeline step. \n",
- "\n",
- "If *allow_reuse* is set to *False*, a new run will always be generated for this step during pipeline execution. The *allow_reuse* flag can come in handy in situations where you do *not* want to re-run a pipeline step."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Running a few steps in parallel\n",
- "Here we are looking at a simple scenario where we are running a few steps (all involving PythonScriptStep) in parallel. Running nodes in **parallel** is the default behavior for steps in a pipeline.\n",
- "\n",
- "We already have one step defined earlier. Let's define few more steps."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# All steps use files already available in the project_folder\n",
- "# All steps use the same Azure Machine Learning compute target as well\n",
- "step2 = PythonScriptStep(name=\"compare_step\",\n",
- " script_name=\"compare.py\", \n",
- " compute_target=aml_compute, \n",
- " source_directory=project_folder)\n",
- "\n",
- "step3 = PythonScriptStep(name=\"extract_step\",\n",
- " script_name=\"extract.py\", \n",
- " compute_target=aml_compute, \n",
- " source_directory=project_folder)\n",
- "\n",
- "# list of steps to run\n",
- "steps = [step1, step2, step3]\n",
- "print(\"Step lists created\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Build the pipeline\n",
- "Once we have the steps (or steps collection), we can build the [pipeline](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-core/azureml.pipeline.core.pipeline.pipeline?view=azure-ml-py). By deafult, all these steps will run in **parallel** once we submit the pipeline for run.\n",
- "\n",
- "A pipeline is created with a list of steps and a workspace. Submit a pipeline using [submit](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.experiment%28class%29?view=azure-ml-py#submit). When submit is called, a [PipelineRun](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-core/azureml.pipeline.core.pipelinerun?view=azure-ml-py) is created which in turn creates [StepRun](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-core/azureml.pipeline.core.steprun?view=azure-ml-py) objects for each step in the workflow."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Syntax\n",
- "# Pipeline(workspace, \n",
- "# steps, \n",
- "# description=None, \n",
- "# default_datastore_name=None, \n",
- "# default_source_directory=None, \n",
- "# resolve_closure=True, \n",
- "# _workflow_provider=None, \n",
- "# _service_endpoint=None)\n",
- "\n",
- "pipeline1 = Pipeline(workspace=ws, steps=steps)\n",
- "print (\"Pipeline is built\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Validate the pipeline\n",
- "You have the option to [validate](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-core/azureml.pipeline.core.pipeline.pipeline?view=azure-ml-py#validate) the pipeline prior to submitting for run. The platform runs validation steps such as checking for circular dependencies and parameter checks etc. even if you do not explicitly call validate method."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "pipeline1.validate()\n",
- "print(\"Pipeline validation complete\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Submit the pipeline\n",
- "[Submitting](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-core/azureml.pipeline.core.pipeline.pipeline?view=azure-ml-py#submit) the pipeline involves creating an [Experiment](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.experiment?view=azure-ml-py) object and providing the built pipeline for submission. "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Submit syntax\n",
- "# submit(experiment_name, \n",
- "# pipeline_parameters=None, \n",
- "# continue_on_node_failure=False, \n",
- "# regenerate_outputs=False)\n",
- "\n",
- "pipeline_run1 = Experiment(ws, 'Hello_World1').submit(pipeline1, regenerate_outputs=True)\n",
- "print(\"Pipeline is submitted for execution\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "**Note:** If regenerate_outputs is set to True, a new submit will always force generation of all step outputs, and disallow data reuse for any step of this run. Once this run is complete, however, subsequent runs may reuse the results of this run.\n"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Examine the pipeline run\n",
- "\n",
- "#### Use RunDetails Widget\n",
- "We are going to use the RunDetails widget to examine the run of the pipeline. You can click each row below to get more details on the step runs."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "RunDetails(pipeline_run1).show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Use Pipeline SDK objects\n",
- "You can cycle through the node_run objects and examine job logs, stdout, and stderr of each of the steps."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "step_runs = pipeline_run1.get_children()\n",
- "for step_run in step_runs:\n",
- " status = step_run.get_status()\n",
- " print('Script:', step_run.name, 'status:', status)\n",
- " \n",
- " # Change this if you want to see details even if the Step has succeeded.\n",
- " if status == \"Failed\":\n",
- " joblog = step_run.get_job_log()\n",
- " print('job log:', joblog)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Get additonal run details\n",
- "If you wait until the pipeline_run is finished, you may be able to get additional details on the run. **Since this is a blocking call, the following code is commented out.**"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "#pipeline_run1.wait_for_completion()\n",
- "#for step_run in pipeline_run1.get_children():\n",
- "# print(\"{}: {}\".format(step_run.name, step_run.get_metrics()))"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Running a few steps in sequence\n",
- "Now let's see how we run a few steps in sequence. We already have three steps defined earlier. Let's *reuse* those steps for this part.\n",
- "\n",
- "We will reuse step1, step2, step3, but build the pipeline in such a way that we chain step3 after step2 and step2 after step1. Note that there is no explicit data dependency between these steps, but still steps can be made dependent by using the [run_after](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-core/azureml.pipeline.core.builder.pipelinestep?view=azure-ml-py#run-after) construct."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "step2.run_after(step1)\n",
- "step3.run_after(step2)\n",
- "\n",
- "# Try a loop\n",
- "#step2.run_after(step3)\n",
- "\n",
- "# Now, construct the pipeline using the steps.\n",
- "\n",
- "# We can specify the \"final step\" in the chain, \n",
- "# Pipeline will take care of \"transitive closure\" and \n",
- "# figure out the implicit or explicit dependencies\n",
- "# https://www.geeksforgeeks.org/transitive-closure-of-a-graph/\n",
- "pipeline2 = Pipeline(workspace=ws, steps=[step3])\n",
- "print (\"Pipeline is built\")\n",
- "\n",
- "pipeline2.validate()\n",
- "print(\"Simple validation complete\")"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "pipeline_run2 = Experiment(ws, 'Hello_World2').submit(pipeline2)\n",
- "print(\"Pipeline is submitted for execution\")"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "RunDetails(pipeline_run2).show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Next: Pipelines with data dependency\n",
- "The next [notebook](./aml-pipelines-with-data-dependency-steps.ipynb) demostrates how to construct a pipeline with data dependency."
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "diray"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved. \n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Azure Machine Learning Pipelines: Getting Started\n",
+ "\n",
+ "## Overview\n",
+ "\n",
+ "Read [Azure Machine Learning Pipelines](https://docs.microsoft.com/en-us/azure/machine-learning/service/concept-ml-pipelines) overview, or the [readme article](../README.md) on Azure Machine Learning Pipelines to get more information.\n",
+ " \n",
+ "\n",
+ "This Notebook shows basic construction of a **pipeline** that runs jobs unattended in different compute clusters. "
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Prerequisites and Azure Machine Learning Basics\n",
+ "Make sure you go through the configuration Notebook located at https://github.com/Azure/MachineLearningNotebooks first if you haven't. This sets you up with a working config file that has information on your workspace, subscription id, etc. \n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Azure Machine Learning Imports\n",
+ "\n",
+ "In this first code cell, we import key Azure Machine Learning modules that we will use below. "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import azureml.core\n",
+ "from azureml.core import Workspace, Run, Experiment, Datastore\n",
+ "from azureml.core.compute import AmlCompute\n",
+ "from azureml.core.compute import ComputeTarget\n",
+ "from azureml.core.compute import DataFactoryCompute\n",
+ "from azureml.widgets import RunDetails\n",
+ "\n",
+ "# Check core SDK version number\n",
+ "print(\"SDK version:\", azureml.core.VERSION)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Pipeline-specific SDK imports\n",
+ "\n",
+ "Here, we import key pipeline modules, whose use will be illustrated in the examples below."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.data.data_reference import DataReference\n",
+ "from azureml.pipeline.core import Pipeline, PipelineData, StepSequence\n",
+ "from azureml.pipeline.steps import PythonScriptStep\n",
+ "from azureml.pipeline.steps import DataTransferStep\n",
+ "from azureml.pipeline.core import PublishedPipeline\n",
+ "from azureml.pipeline.core.graph import PipelineParameter\n",
+ "\n",
+ "print(\"Pipeline SDK-specific imports completed\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Initialize Workspace\n",
+ "\n",
+ "Initialize a [workspace](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.workspace(class%29) object from persisted configuration."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "create workspace"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "ws = Workspace.from_config()\n",
+ "print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep = '\\n')\n",
+ "\n",
+ "# Default datastore (Azure file storage)\n",
+ "def_file_store = ws.get_default_datastore() \n",
+ "# The above call is equivalent to Datastore(ws, \"workspacefilestore\") or simply Datastore(ws)\n",
+ "print(\"Default datastore's name: {}\".format(def_file_store.name))\n",
+ "\n",
+ "# Blob storage associated with the workspace\n",
+ "# The following call GETS the Azure Blob Store associated with your workspace.\n",
+ "# Note that workspaceblobstore is **the name of this store and CANNOT BE CHANGED and must be used as is** \n",
+ "def_blob_store = Datastore(ws, \"workspaceblobstore\")\n",
+ "print(\"Blobstore's name: {}\".format(def_blob_store.name))"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# project folder\n",
+ "project_folder = '.'\n",
+ " \n",
+ "print('Sample projects will be created in {}.'.format(project_folder))"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Required data and script files for the the tutorial\n",
+ "Sample files required to finish this tutorial are already copied to the project folder specified above. Even though the .py provided in the samples don't have much \"ML work,\" as a data scientist, you will work on this extensively as part of your work. To complete this tutorial, the contents of these files are not very important. The one-line files are for demostration purpose only."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Datastore concepts\n",
+ "A [Datastore](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.datastore(class) is a place where data can be stored that is then made accessible to a compute either by means of mounting or copying the data to the compute target. \n",
+ "\n",
+ "A Datastore can either be backed by an Azure File Storage (default) or by an Azure Blob Storage.\n",
+ "\n",
+ "In this next step, we will upload the training and test set into the workspace's default storage (File storage), and another piece of data to Azure Blob Storage. When to use [Azure Blobs](https://docs.microsoft.com/en-us/azure/storage/blobs/storage-blobs-introduction), [Azure Files](https://docs.microsoft.com/en-us/azure/storage/files/storage-files-introduction), or [Azure Disks](https://docs.microsoft.com/en-us/azure/virtual-machines/linux/managed-disks-overview) is [detailed here](https://docs.microsoft.com/en-us/azure/storage/common/storage-decide-blobs-files-disks).\n",
+ "\n",
+ "**Please take good note of the concept of the datastore.**"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Upload data to default datastore\n",
+ "Default datastore on workspace is the Azure File storage. The workspace has a Blob storage associated with it as well. Let's upload a file to each of these storages."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# get_default_datastore() gets the default Azure File Store associated with your workspace.\n",
+ "# Here we are reusing the def_file_store object we obtained earlier\n",
+ "\n",
+ "# target_path is the directory at the destination\n",
+ "def_file_store.upload_files(['./20news.pkl'], \n",
+ " target_path = '20newsgroups', \n",
+ " overwrite = True, \n",
+ " show_progress = True)\n",
+ "\n",
+ "# Here we are reusing the def_blob_store we created earlier\n",
+ "def_blob_store.upload_files([\"./20news.pkl\"], target_path=\"20newsgroups\", overwrite=True)\n",
+ "\n",
+ "print(\"Upload calls completed\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### (Optional) See your files using Azure Portal\n",
+ "Once you successfully uploaded the files, you can browse to them (or upload more files) using [Azure Portal](https://portal.azure.com). At the portal, make sure you have selected **AzureML Nursery** as your subscription (click *Resource Groups* and then select the subscription). Then look for your **Machine Learning Workspace** (it has your *alias* as the name). It has a link to your storage. Click on the storage link. It will take you to a page where you can see [Blobs](https://docs.microsoft.com/en-us/azure/storage/blobs/storage-blobs-introduction), [Files](https://docs.microsoft.com/en-us/azure/storage/files/storage-files-introduction), [Tables](https://docs.microsoft.com/en-us/azure/storage/tables/table-storage-overview), and [Queues](https://docs.microsoft.com/en-us/azure/storage/queues/storage-queues-introduction). We have just uploaded a file to the Blob storage and another one to the File storage. You should be able to see both of these files in their respective locations. "
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Compute Targets\n",
+ "A compute target specifies where to execute your program such as a remote Docker on a VM, or a cluster. A compute target needs to be addressable and accessible by you.\n",
+ "\n",
+ "**You need at least one compute target to send your payload to. We are planning to use Azure Machine Learning Compute exclusively for this tutorial for all steps. However in some cases you may require multiple compute targets as some steps may run in one compute target like Azure Machine Learning Compute, and some other steps in the same pipeline could run in a different compute target.**\n",
+ "\n",
+ "*The example belows show creating/retrieving/attaching to an Azure Machine Learning Compute instance.*"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### List of Compute Targets on the workspace"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "cts = ws.compute_targets\n",
+ "for ct in cts:\n",
+ " print(ct)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Retrieve or create a Azure Machine Learning compute\n",
+ "Azure Machine Learning Compute is a service for provisioning and managing clusters of Azure virtual machines for running machine learning workloads. Let's create a new Azure Machine Learning Compute in the current workspace, if it doesn't already exist. We will then run the training script on this compute target.\n",
+ "\n",
+ "If we could not find the compute with the given name in the previous cell, then we will create a new compute here. We will create an Azure Machine Learning Compute containing **STANDARD_D2_V2 CPU VMs**. This process is broken down into the following steps:\n",
+ "\n",
+ "1. Create the configuration\n",
+ "2. Create the Azure Machine Learning compute\n",
+ "\n",
+ "**This process will take about 3 minutes and is providing only sparse output in the process. Please make sure to wait until the call returns before moving to the next cell.**"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "\n",
+ "aml_compute_target = \"aml-compute\"\n",
+ "try:\n",
+ " aml_compute = AmlCompute(ws, aml_compute_target)\n",
+ " print(\"found existing compute target.\")\n",
+ "except:\n",
+ " print(\"creating new compute target\")\n",
+ " \n",
+ " provisioning_config = AmlCompute.provisioning_configuration(vm_size = \"STANDARD_D2_V2\",\n",
+ " min_nodes = 1, \n",
+ " max_nodes = 4) \n",
+ " aml_compute = ComputeTarget.create(ws, aml_compute_target, provisioning_config)\n",
+ " aml_compute.wait_for_completion(show_output=True, min_node_count=None, timeout_in_minutes=20)\n",
+ " \n",
+ "print(\"Azure Machine Learning Compute attached\")\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# For a more detailed view of current Azure Machine Learning Compute status, use get_status()\n",
+ "# example: un-comment the following line.\n",
+ "# print(aml_compute.get_status().serialize())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "**Wait for this call to finish before proceeding (you will see the asterisk turning to a number).**\n",
+ "\n",
+ "Now that you have created the compute target, let's see what the workspace's compute_targets() function returns. You should now see one entry named 'amlcompute' of type AmlCompute."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "**Now that we have completed learning the basics of Azure Machine Learning (AML), let's go ahead and start understanding the Pipeline concepts.**"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Creating a Step in a Pipeline\n",
+ "A Step is a unit of execution. Step typically needs a target of execution (compute target), a script to execute, and may require script arguments and inputs, and can produce outputs. The step also could take a number of other parameters. Azure Machine Learning Pipelines provides the following built-in Steps:\n",
+ "\n",
+ "- [**PythonScriptStep**](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-steps/azureml.pipeline.steps.python_script_step.pythonscriptstep?view=azure-ml-py): Add a step to run a Python script in a Pipeline.\n",
+ "- [**AdlaStep**](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-steps/azureml.pipeline.steps.adla_step.adlastep?view=azure-ml-py): Adds a step to run U-SQL script using Azure Data Lake Analytics.\n",
+ "- [**DataTransferStep**](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-steps/azureml.pipeline.steps.data_transfer_step.datatransferstep?view=azure-ml-py): Transfers data between Azure Blob and Data Lake accounts.\n",
+ "- [**DatabricksStep**](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-steps/azureml.pipeline.steps.databricks_step.databricksstep?view=azure-ml-py): Adds a DataBricks notebook as a step in a Pipeline.\n",
+ "- [**HyperDriveStep**](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-steps/azureml.pipeline.steps.hyper_drive_step.hyperdrivestep?view=azure-ml-py): Creates a Hyper Drive step for Hyper Parameter Tuning in a Pipeline.\n",
+ "\n",
+ "The following code will create a PythonScriptStep to be executed in the Azure Machine Learning Compute we created above using train.py, one of the files already made available in the project folder.\n",
+ "\n",
+ "A **PythonScriptStep** is a basic, built-in step to run a Python Script on a compute target. It takes a script name and optionally other parameters like arguments for the script, compute target, inputs and outputs. If no compute target is specified, default compute target for the workspace is used."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Uses default values for PythonScriptStep construct.\n",
+ "\n",
+ "# Syntax\n",
+ "# PythonScriptStep(\n",
+ "# script_name, \n",
+ "# name=None, \n",
+ "# arguments=None, \n",
+ "# compute_target=None, \n",
+ "# runconfig=None, \n",
+ "# inputs=None, \n",
+ "# outputs=None, \n",
+ "# params=None, \n",
+ "# source_directory=None, \n",
+ "# allow_reuse=True, \n",
+ "# version=None, \n",
+ "# hash_paths=None)\n",
+ "# This returns a Step\n",
+ "step1 = PythonScriptStep(name=\"train_step\",\n",
+ " script_name=\"train.py\", \n",
+ " compute_target=aml_compute, \n",
+ " source_directory=project_folder,\n",
+ " allow_reuse=False)\n",
+ "print(\"Step1 created\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "**Note:** In the above call to PythonScriptStep(), the flag *allow_reuse* determines whether the step should reuse previous results when run with the same settings/inputs. This flag's default value is *True*; the default is set to *True* because, when inputs and parameters have not changed, we typically do not want to re-run a given pipeline step. \n",
+ "\n",
+ "If *allow_reuse* is set to *False*, a new run will always be generated for this step during pipeline execution. The *allow_reuse* flag can come in handy in situations where you do *not* want to re-run a pipeline step."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Running a few steps in parallel\n",
+ "Here we are looking at a simple scenario where we are running a few steps (all involving PythonScriptStep) in parallel. Running nodes in **parallel** is the default behavior for steps in a pipeline.\n",
+ "\n",
+ "We already have one step defined earlier. Let's define few more steps."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# All steps use files already available in the project_folder\n",
+ "# All steps use the same Azure Machine Learning compute target as well\n",
+ "step2 = PythonScriptStep(name=\"compare_step\",\n",
+ " script_name=\"compare.py\", \n",
+ " compute_target=aml_compute, \n",
+ " source_directory=project_folder)\n",
+ "\n",
+ "step3 = PythonScriptStep(name=\"extract_step\",\n",
+ " script_name=\"extract.py\", \n",
+ " compute_target=aml_compute, \n",
+ " source_directory=project_folder)\n",
+ "\n",
+ "# list of steps to run\n",
+ "steps = [step1, step2, step3]\n",
+ "print(\"Step lists created\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Build the pipeline\n",
+ "Once we have the steps (or steps collection), we can build the [pipeline](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-core/azureml.pipeline.core.pipeline.pipeline?view=azure-ml-py). By deafult, all these steps will run in **parallel** once we submit the pipeline for run.\n",
+ "\n",
+ "A pipeline is created with a list of steps and a workspace. Submit a pipeline using [submit](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.experiment%28class%29?view=azure-ml-py#submit). When submit is called, a [PipelineRun](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-core/azureml.pipeline.core.pipelinerun?view=azure-ml-py) is created which in turn creates [StepRun](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-core/azureml.pipeline.core.steprun?view=azure-ml-py) objects for each step in the workflow."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Syntax\n",
+ "# Pipeline(workspace, \n",
+ "# steps, \n",
+ "# description=None, \n",
+ "# default_datastore_name=None, \n",
+ "# default_source_directory=None, \n",
+ "# resolve_closure=True, \n",
+ "# _workflow_provider=None, \n",
+ "# _service_endpoint=None)\n",
+ "\n",
+ "pipeline1 = Pipeline(workspace=ws, steps=steps)\n",
+ "print (\"Pipeline is built\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Validate the pipeline\n",
+ "You have the option to [validate](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-core/azureml.pipeline.core.pipeline.pipeline?view=azure-ml-py#validate) the pipeline prior to submitting for run. The platform runs validation steps such as checking for circular dependencies and parameter checks etc. even if you do not explicitly call validate method."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "pipeline1.validate()\n",
+ "print(\"Pipeline validation complete\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Submit the pipeline\n",
+ "[Submitting](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-core/azureml.pipeline.core.pipeline.pipeline?view=azure-ml-py#submit) the pipeline involves creating an [Experiment](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.experiment?view=azure-ml-py) object and providing the built pipeline for submission. "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Submit syntax\n",
+ "# submit(experiment_name, \n",
+ "# pipeline_parameters=None, \n",
+ "# continue_on_node_failure=False, \n",
+ "# regenerate_outputs=False)\n",
+ "\n",
+ "pipeline_run1 = Experiment(ws, 'Hello_World1').submit(pipeline1, regenerate_outputs=True)\n",
+ "print(\"Pipeline is submitted for execution\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "**Note:** If regenerate_outputs is set to True, a new submit will always force generation of all step outputs, and disallow data reuse for any step of this run. Once this run is complete, however, subsequent runs may reuse the results of this run.\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Examine the pipeline run\n",
+ "\n",
+ "#### Use RunDetails Widget\n",
+ "We are going to use the RunDetails widget to examine the run of the pipeline. You can click each row below to get more details on the step runs."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "RunDetails(pipeline_run1).show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Use Pipeline SDK objects\n",
+ "You can cycle through the node_run objects and examine job logs, stdout, and stderr of each of the steps."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "step_runs = pipeline_run1.get_children()\n",
+ "for step_run in step_runs:\n",
+ " status = step_run.get_status()\n",
+ " print('Script:', step_run.name, 'status:', status)\n",
+ " \n",
+ " # Change this if you want to see details even if the Step has succeeded.\n",
+ " if status == \"Failed\":\n",
+ " joblog = step_run.get_job_log()\n",
+ " print('job log:', joblog)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Get additonal run details\n",
+ "If you wait until the pipeline_run is finished, you may be able to get additional details on the run. **Since this is a blocking call, the following code is commented out.**"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "#pipeline_run1.wait_for_completion()\n",
+ "#for step_run in pipeline_run1.get_children():\n",
+ "# print(\"{}: {}\".format(step_run.name, step_run.get_metrics()))"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Running a few steps in sequence\n",
+ "Now let's see how we run a few steps in sequence. We already have three steps defined earlier. Let's *reuse* those steps for this part.\n",
+ "\n",
+ "We will reuse step1, step2, step3, but build the pipeline in such a way that we chain step3 after step2 and step2 after step1. Note that there is no explicit data dependency between these steps, but still steps can be made dependent by using the [run_after](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-core/azureml.pipeline.core.builder.pipelinestep?view=azure-ml-py#run-after) construct."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "step2.run_after(step1)\n",
+ "step3.run_after(step2)\n",
+ "\n",
+ "# Try a loop\n",
+ "#step2.run_after(step3)\n",
+ "\n",
+ "# Now, construct the pipeline using the steps.\n",
+ "\n",
+ "# We can specify the \"final step\" in the chain, \n",
+ "# Pipeline will take care of \"transitive closure\" and \n",
+ "# figure out the implicit or explicit dependencies\n",
+ "# https://www.geeksforgeeks.org/transitive-closure-of-a-graph/\n",
+ "pipeline2 = Pipeline(workspace=ws, steps=[step3])\n",
+ "print (\"Pipeline is built\")\n",
+ "\n",
+ "pipeline2.validate()\n",
+ "print(\"Simple validation complete\")"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "pipeline_run2 = Experiment(ws, 'Hello_World2').submit(pipeline2)\n",
+ "print(\"Pipeline is submitted for execution\")"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "RunDetails(pipeline_run2).show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Next: Pipelines with data dependency\n",
+ "The next [notebook](./aml-pipelines-with-data-dependency-steps.ipynb) demostrates how to construct a pipeline with data dependency."
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3",
- "language": "python",
- "name": "python3"
+ "metadata": {
+ "authors": [
+ {
+ "name": "diray"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.7"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.7"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/machine-learning-pipelines/intro-to-pipelines/aml-pipelines-publish-and-run-using-rest-endpoint.ipynb b/how-to-use-azureml/machine-learning-pipelines/intro-to-pipelines/aml-pipelines-publish-and-run-using-rest-endpoint.ipynb
index cfadc3b7..b9bba0f7 100644
--- a/how-to-use-azureml/machine-learning-pipelines/intro-to-pipelines/aml-pipelines-publish-and-run-using-rest-endpoint.ipynb
+++ b/how-to-use-azureml/machine-learning-pipelines/intro-to-pipelines/aml-pipelines-publish-and-run-using-rest-endpoint.ipynb
@@ -1,368 +1,368 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved. \n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# How to Publish a Pipeline and Invoke the REST endpoint\n",
- "In this notebook, we will see how we can publish a pipeline and then invoke the REST endpoint."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Prerequisites and Azure Machine Learning Basics\n",
- "Make sure you go through the configuration Notebook located at https://github.com/Azure/MachineLearningNotebooks first if you haven't. This sets you up with a working config file that has information on your workspace, subscription id, etc. \n",
- "\n",
- "### Initialization Steps"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import azureml.core\n",
- "from azureml.core import Workspace, Run, Experiment, Datastore\n",
- "from azureml.core.compute import AmlCompute\n",
- "from azureml.core.compute import ComputeTarget\n",
- "from azureml.core.compute import DataFactoryCompute\n",
- "from azureml.widgets import RunDetails\n",
- "\n",
- "# Check core SDK version number\n",
- "print(\"SDK version:\", azureml.core.VERSION)\n",
- "\n",
- "from azureml.data.data_reference import DataReference\n",
- "from azureml.pipeline.core import Pipeline, PipelineData, StepSequence\n",
- "from azureml.pipeline.steps import PythonScriptStep\n",
- "from azureml.pipeline.steps import DataTransferStep\n",
- "from azureml.pipeline.core import PublishedPipeline\n",
- "from azureml.pipeline.core.graph import PipelineParameter\n",
- "\n",
- "print(\"Pipeline SDK-specific imports completed\")\n",
- "\n",
- "ws = Workspace.from_config()\n",
- "print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep = '\\n')\n",
- "\n",
- "# Default datastore (Azure file storage)\n",
- "def_file_store = ws.get_default_datastore() \n",
- "print(\"Default datastore's name: {}\".format(def_file_store.name))\n",
- "\n",
- "def_blob_store = Datastore(ws, \"workspaceblobstore\")\n",
- "print(\"Blobstore's name: {}\".format(def_blob_store.name))\n",
- "\n",
- "# project folder\n",
- "project_folder = '.'"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Compute Targets\n",
- "#### Retrieve an already attached Azure Machine Learning Compute"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "\n",
- "aml_compute_target = \"aml-compute\"\n",
- "try:\n",
- " aml_compute = AmlCompute(ws, aml_compute_target)\n",
- " print(\"found existing compute target.\")\n",
- "except:\n",
- " print(\"creating new compute target\")\n",
- " \n",
- " provisioning_config = AmlCompute.provisioning_configuration(vm_size = \"STANDARD_D2_V2\",\n",
- " min_nodes = 1, \n",
- " max_nodes = 4) \n",
- " aml_compute = ComputeTarget.create(ws, aml_compute_target, provisioning_config)\n",
- " aml_compute.wait_for_completion(show_output=True, min_node_count=None, timeout_in_minutes=20)\n"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# For a more detailed view of current Azure Machine Learning Compute status, use the 'status' property\n",
- "# example: un-comment the following line.\n",
- "# print(aml_compute.status.serialize())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Building Pipeline Steps with Inputs and Outputs\n",
- "As mentioned earlier, a step in the pipeline can take data as input. This data can be a data source that lives in one of the accessible data locations, or intermediate data produced by a previous step in the pipeline."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Reference the data uploaded to blob storage using DataReference\n",
- "# Assign the datasource to blob_input_data variable\n",
- "blob_input_data = DataReference(\n",
- " datastore=def_blob_store,\n",
- " data_reference_name=\"test_data\",\n",
- " path_on_datastore=\"20newsgroups/20news.pkl\")\n",
- "print(\"DataReference object created\")"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Define intermediate data using PipelineData\n",
- "processed_data1 = PipelineData(\"processed_data1\",datastore=def_blob_store)\n",
- "print(\"PipelineData object created\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Define a Step that consumes a datasource and produces intermediate data.\n",
- "In this step, we define a step that consumes a datasource and produces intermediate data.\n",
- "\n",
- "**Open `train.py` in the local machine and examine the arguments, inputs, and outputs for the script. That will give you a good sense of why the script argument names used below are important.** "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# trainStep consumes the datasource (Datareference) in the previous step\n",
- "# and produces processed_data1\n",
- "trainStep = PythonScriptStep(\n",
- " script_name=\"train.py\", \n",
- " arguments=[\"--input_data\", blob_input_data, \"--output_train\", processed_data1],\n",
- " inputs=[blob_input_data],\n",
- " outputs=[processed_data1],\n",
- " compute_target=aml_compute, \n",
- " source_directory=project_folder\n",
- ")\n",
- "print(\"trainStep created\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Define a Step that consumes intermediate data and produces intermediate data\n",
- "In this step, we define a step that consumes an intermediate data and produces intermediate data.\n",
- "\n",
- "**Open `extract.py` in the local machine and examine the arguments, inputs, and outputs for the script. That will give you a good sense of why the script argument names used below are important.** "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# extractStep to use the intermediate data produced by step4\n",
- "# This step also produces an output processed_data2\n",
- "processed_data2 = PipelineData(\"processed_data2\", datastore=def_blob_store)\n",
- "\n",
- "extractStep = PythonScriptStep(\n",
- " script_name=\"extract.py\",\n",
- " arguments=[\"--input_extract\", processed_data1, \"--output_extract\", processed_data2],\n",
- " inputs=[processed_data1],\n",
- " outputs=[processed_data2],\n",
- " compute_target=aml_compute, \n",
- " source_directory=project_folder)\n",
- "print(\"extractStep created\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Define a Step that consumes multiple intermediate data and produces intermediate data\n",
- "In this step, we define a step that consumes multiple intermediate data and produces intermediate data."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### PipelineParameter"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "This step also has a [PipelineParameter](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-core/azureml.pipeline.core.graph.pipelineparameter?view=azure-ml-py) argument that help with calling the REST endpoint of the published pipeline."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# We will use this later in publishing pipeline\n",
- "pipeline_param = PipelineParameter(name=\"pipeline_arg\", default_value=10)\n",
- "print(\"pipeline parameter created\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "**Open `compare.py` in the local machine and examine the arguments, inputs, and outputs for the script. That will give you a good sense of why the script argument names used below are important.**"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Now define step6 that takes two inputs (both intermediate data), and produce an output\n",
- "processed_data3 = PipelineData(\"processed_data3\", datastore=def_blob_store)\n",
- "\n",
- "\n",
- "\n",
- "compareStep = PythonScriptStep(\n",
- " script_name=\"compare.py\",\n",
- " arguments=[\"--compare_data1\", processed_data1, \"--compare_data2\", processed_data2, \"--output_compare\", processed_data3, \"--pipeline_param\", pipeline_param],\n",
- " inputs=[processed_data1, processed_data2],\n",
- " outputs=[processed_data3], \n",
- " compute_target=aml_compute, \n",
- " source_directory=project_folder)\n",
- "print(\"compareStep created\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Build the pipeline"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "pipeline1 = Pipeline(workspace=ws, steps=[compareStep])\n",
- "print (\"Pipeline is built\")\n",
- "\n",
- "pipeline1.validate()\n",
- "print(\"Simple validation complete\") "
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Publish the pipeline"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "published_pipeline1 = pipeline1.publish(name=\"My_New_Pipeline\", description=\"My Published Pipeline Description\")\n",
- "print(published_pipeline1.id)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Run published pipeline using its REST endpoint"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.authentication import AzureCliAuthentication\n",
- "import requests\n",
- "\n",
- "cli_auth = AzureCliAuthentication()\n",
- "aad_token = cli_auth.get_authentication_header()\n",
- "\n",
- "rest_endpoint1 = published_pipeline1.endpoint\n",
- "\n",
- "print(rest_endpoint1)\n",
- "\n",
- "# specify the param when running the pipeline\n",
- "response = requests.post(rest_endpoint1, \n",
- " headers=aad_token, \n",
- " json={\"ExperimentName\": \"My_Pipeline1\",\n",
- " \"RunSource\": \"SDK\",\n",
- " \"ParameterAssignments\": {\"pipeline_arg\": 45}})\n",
- "run_id = response.json()[\"Id\"]\n",
- "\n",
- "print(run_id)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Next: Data Transfer\n",
- "The next [notebook](./aml-pipelines-data-transfer.ipynb) will showcase data transfer steps between different types of data stores."
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "diray"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved. \n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# How to Publish a Pipeline and Invoke the REST endpoint\n",
+ "In this notebook, we will see how we can publish a pipeline and then invoke the REST endpoint."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Prerequisites and Azure Machine Learning Basics\n",
+ "Make sure you go through the configuration Notebook located at https://github.com/Azure/MachineLearningNotebooks first if you haven't. This sets you up with a working config file that has information on your workspace, subscription id, etc. \n",
+ "\n",
+ "### Initialization Steps"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import azureml.core\n",
+ "from azureml.core import Workspace, Run, Experiment, Datastore\n",
+ "from azureml.core.compute import AmlCompute\n",
+ "from azureml.core.compute import ComputeTarget\n",
+ "from azureml.core.compute import DataFactoryCompute\n",
+ "from azureml.widgets import RunDetails\n",
+ "\n",
+ "# Check core SDK version number\n",
+ "print(\"SDK version:\", azureml.core.VERSION)\n",
+ "\n",
+ "from azureml.data.data_reference import DataReference\n",
+ "from azureml.pipeline.core import Pipeline, PipelineData, StepSequence\n",
+ "from azureml.pipeline.steps import PythonScriptStep\n",
+ "from azureml.pipeline.steps import DataTransferStep\n",
+ "from azureml.pipeline.core import PublishedPipeline\n",
+ "from azureml.pipeline.core.graph import PipelineParameter\n",
+ "\n",
+ "print(\"Pipeline SDK-specific imports completed\")\n",
+ "\n",
+ "ws = Workspace.from_config()\n",
+ "print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep = '\\n')\n",
+ "\n",
+ "# Default datastore (Azure file storage)\n",
+ "def_file_store = ws.get_default_datastore() \n",
+ "print(\"Default datastore's name: {}\".format(def_file_store.name))\n",
+ "\n",
+ "def_blob_store = Datastore(ws, \"workspaceblobstore\")\n",
+ "print(\"Blobstore's name: {}\".format(def_blob_store.name))\n",
+ "\n",
+ "# project folder\n",
+ "project_folder = '.'"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Compute Targets\n",
+ "#### Retrieve an already attached Azure Machine Learning Compute"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "\n",
+ "aml_compute_target = \"aml-compute\"\n",
+ "try:\n",
+ " aml_compute = AmlCompute(ws, aml_compute_target)\n",
+ " print(\"found existing compute target.\")\n",
+ "except:\n",
+ " print(\"creating new compute target\")\n",
+ " \n",
+ " provisioning_config = AmlCompute.provisioning_configuration(vm_size = \"STANDARD_D2_V2\",\n",
+ " min_nodes = 1, \n",
+ " max_nodes = 4) \n",
+ " aml_compute = ComputeTarget.create(ws, aml_compute_target, provisioning_config)\n",
+ " aml_compute.wait_for_completion(show_output=True, min_node_count=None, timeout_in_minutes=20)\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# For a more detailed view of current Azure Machine Learning Compute status, use get_status()\n",
+ "# example: un-comment the following line.\n",
+ "# print(aml_compute.get_status().serialize())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Building Pipeline Steps with Inputs and Outputs\n",
+ "As mentioned earlier, a step in the pipeline can take data as input. This data can be a data source that lives in one of the accessible data locations, or intermediate data produced by a previous step in the pipeline."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Reference the data uploaded to blob storage using DataReference\n",
+ "# Assign the datasource to blob_input_data variable\n",
+ "blob_input_data = DataReference(\n",
+ " datastore=def_blob_store,\n",
+ " data_reference_name=\"test_data\",\n",
+ " path_on_datastore=\"20newsgroups/20news.pkl\")\n",
+ "print(\"DataReference object created\")"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Define intermediate data using PipelineData\n",
+ "processed_data1 = PipelineData(\"processed_data1\",datastore=def_blob_store)\n",
+ "print(\"PipelineData object created\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Define a Step that consumes a datasource and produces intermediate data.\n",
+ "In this step, we define a step that consumes a datasource and produces intermediate data.\n",
+ "\n",
+ "**Open `train.py` in the local machine and examine the arguments, inputs, and outputs for the script. That will give you a good sense of why the script argument names used below are important.** "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# trainStep consumes the datasource (Datareference) in the previous step\n",
+ "# and produces processed_data1\n",
+ "trainStep = PythonScriptStep(\n",
+ " script_name=\"train.py\", \n",
+ " arguments=[\"--input_data\", blob_input_data, \"--output_train\", processed_data1],\n",
+ " inputs=[blob_input_data],\n",
+ " outputs=[processed_data1],\n",
+ " compute_target=aml_compute, \n",
+ " source_directory=project_folder\n",
+ ")\n",
+ "print(\"trainStep created\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Define a Step that consumes intermediate data and produces intermediate data\n",
+ "In this step, we define a step that consumes an intermediate data and produces intermediate data.\n",
+ "\n",
+ "**Open `extract.py` in the local machine and examine the arguments, inputs, and outputs for the script. That will give you a good sense of why the script argument names used below are important.** "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# extractStep to use the intermediate data produced by step4\n",
+ "# This step also produces an output processed_data2\n",
+ "processed_data2 = PipelineData(\"processed_data2\", datastore=def_blob_store)\n",
+ "\n",
+ "extractStep = PythonScriptStep(\n",
+ " script_name=\"extract.py\",\n",
+ " arguments=[\"--input_extract\", processed_data1, \"--output_extract\", processed_data2],\n",
+ " inputs=[processed_data1],\n",
+ " outputs=[processed_data2],\n",
+ " compute_target=aml_compute, \n",
+ " source_directory=project_folder)\n",
+ "print(\"extractStep created\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Define a Step that consumes multiple intermediate data and produces intermediate data\n",
+ "In this step, we define a step that consumes multiple intermediate data and produces intermediate data."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### PipelineParameter"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "This step also has a [PipelineParameter](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-core/azureml.pipeline.core.graph.pipelineparameter?view=azure-ml-py) argument that help with calling the REST endpoint of the published pipeline."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# We will use this later in publishing pipeline\n",
+ "pipeline_param = PipelineParameter(name=\"pipeline_arg\", default_value=10)\n",
+ "print(\"pipeline parameter created\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "**Open `compare.py` in the local machine and examine the arguments, inputs, and outputs for the script. That will give you a good sense of why the script argument names used below are important.**"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Now define step6 that takes two inputs (both intermediate data), and produce an output\n",
+ "processed_data3 = PipelineData(\"processed_data3\", datastore=def_blob_store)\n",
+ "\n",
+ "\n",
+ "\n",
+ "compareStep = PythonScriptStep(\n",
+ " script_name=\"compare.py\",\n",
+ " arguments=[\"--compare_data1\", processed_data1, \"--compare_data2\", processed_data2, \"--output_compare\", processed_data3, \"--pipeline_param\", pipeline_param],\n",
+ " inputs=[processed_data1, processed_data2],\n",
+ " outputs=[processed_data3], \n",
+ " compute_target=aml_compute, \n",
+ " source_directory=project_folder)\n",
+ "print(\"compareStep created\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Build the pipeline"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "pipeline1 = Pipeline(workspace=ws, steps=[compareStep])\n",
+ "print (\"Pipeline is built\")\n",
+ "\n",
+ "pipeline1.validate()\n",
+ "print(\"Simple validation complete\") "
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Publish the pipeline"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "published_pipeline1 = pipeline1.publish(name=\"My_New_Pipeline\", description=\"My Published Pipeline Description\")\n",
+ "print(published_pipeline1.id)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Run published pipeline using its REST endpoint"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.authentication import AzureCliAuthentication\n",
+ "import requests\n",
+ "\n",
+ "cli_auth = AzureCliAuthentication()\n",
+ "aad_token = cli_auth.get_authentication_header()\n",
+ "\n",
+ "rest_endpoint1 = published_pipeline1.endpoint\n",
+ "\n",
+ "print(rest_endpoint1)\n",
+ "\n",
+ "# specify the param when running the pipeline\n",
+ "response = requests.post(rest_endpoint1, \n",
+ " headers=aad_token, \n",
+ " json={\"ExperimentName\": \"My_Pipeline1\",\n",
+ " \"RunSource\": \"SDK\",\n",
+ " \"ParameterAssignments\": {\"pipeline_arg\": 45}})\n",
+ "run_id = response.json()[\"Id\"]\n",
+ "\n",
+ "print(run_id)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Next: Data Transfer\n",
+ "The next [notebook](./aml-pipelines-data-transfer.ipynb) will showcase data transfer steps between different types of data stores."
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3",
- "language": "python",
- "name": "python3"
+ "metadata": {
+ "authors": [
+ {
+ "name": "diray"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.7"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.7"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/machine-learning-pipelines/intro-to-pipelines/aml-pipelines-use-adla-as-compute-target.ipynb b/how-to-use-azureml/machine-learning-pipelines/intro-to-pipelines/aml-pipelines-use-adla-as-compute-target.ipynb
index dcff2125..41066444 100644
--- a/how-to-use-azureml/machine-learning-pipelines/intro-to-pipelines/aml-pipelines-use-adla-as-compute-target.ipynb
+++ b/how-to-use-azureml/machine-learning-pipelines/intro-to-pipelines/aml-pipelines-use-adla-as-compute-target.ipynb
@@ -1,368 +1,368 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved. \n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# AML Pipeline with AdlaStep\n",
- "This notebook is used to demonstrate the use of AdlaStep in AML Pipeline."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## AML and Pipeline SDK-specific imports"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import os\n",
- "import azureml.core\n",
- "from azureml.core.compute import ComputeTarget, DatabricksCompute\n",
- "from azureml.exceptions import ComputeTargetException\n",
- "from azureml.core import Workspace, Run, Experiment\n",
- "from azureml.pipeline.core import Pipeline, PipelineData\n",
- "from azureml.pipeline.steps import AdlaStep\n",
- "from azureml.core.datastore import Datastore\n",
- "from azureml.data.data_reference import DataReference\n",
- "from azureml.core import attach_legacy_compute_target\n",
- "\n",
- "# Check core SDK version number\n",
- "print(\"SDK version:\", azureml.core.VERSION)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Initialize Workspace\n",
- "\n",
- "Initialize a workspace object from persisted configuration. Make sure the config file is present at .\\config.json"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "create workspace"
- ]
- },
- "outputs": [],
- "source": [
- "ws = Workspace.from_config()\n",
- "print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep = '\\n')"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "script_folder = '.'\n",
- "experiment_name = \"adla_101_experiment\"\n",
- "ws._initialize_folder(experiment_name=experiment_name, directory=script_folder)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Register Datastore"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "\n",
- "workspace = ws.name\n",
- "datastore_name='MyAdlsDatastore'\n",
- "subscription_id=os.getenv(\"ADL_SUBSCRIPTION_62\", \"\") # subscription id of ADLS account\n",
- "resource_group=os.getenv(\"ADL_RESOURCE_GROUP_62\", \"\") # resource group of ADLS account\n",
- "store_name=os.getenv(\"ADL_STORENAME_62\", \"\") # ADLS account name\n",
- "tenant_id=os.getenv(\"ADL_TENANT_62\", \"\") # tenant id of service principal\n",
- "client_id=os.getenv(\"ADL_CLIENTID_62\", \"\") # client id of service principal\n",
- "client_secret=os.getenv(\"ADL_CLIENT_62_SECRET\", \"\") # the secret of service principal\n",
- "\n",
- "try:\n",
- " adls_datastore = Datastore.get(ws, datastore_name)\n",
- " print(\"found datastore with name: %s\" % datastore_name)\n",
- "except:\n",
- " adls_datastore = Datastore.register_azure_data_lake(\n",
- " workspace=ws,\n",
- " datastore_name=datastore_name,\n",
- " subscription_id=subscription_id, # subscription id of ADLS account\n",
- " resource_group=resource_group, # resource group of ADLS account\n",
- " store_name=store_name, # ADLS account name\n",
- " tenant_id=tenant_id, # tenant id of service principal\n",
- " client_id=client_id, # client id of service principal\n",
- " client_secret=client_secret) # the secret of service principal\n",
- " print(\"registered datastore with name: %s\" % datastore_name)\n"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Create DataReferences and PipelineData\n",
- "\n",
- "In the code cell below, replace datastorename with your default datastore name. Copy the file `testdata.txt` (located in the pipeline folder that this notebook is in) to the path on the datastore."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "datastorename = \"MyAdlsDatastore\"\n",
- "\n",
- "adls_datastore = Datastore(workspace=ws, name=datastorename)\n",
- "script_input = DataReference(\n",
- " datastore=adls_datastore,\n",
- " data_reference_name=\"script_input\",\n",
- " path_on_datastore=\"testdata/testdata.txt\")\n",
- "\n",
- "script_output = PipelineData(\"script_output\", datastore=adls_datastore)\n",
- "\n",
- "print(\"Created Pipeline Data\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Setup Data Lake Account\n",
- "\n",
- "ADLA can only use data that is located in the default data store associated with that ADLA account. Through Azure portal, check the name of the default data store corresponding to the ADLA account you are using below. Replace the value associated with `adla_compute_name` in the code cell below accordingly."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "adla_compute_name = 'testadl' # Replace this with your default compute\n",
- "\n",
- "from azureml.core.compute import ComputeTarget, AdlaCompute\n",
- "\n",
- "def get_or_create_adla_compute(workspace, compute_name):\n",
- " try:\n",
- " return AdlaCompute(workspace, compute_name)\n",
- " except ComputeTargetException as e:\n",
- " if 'ComputeTargetNotFound' in e.message:\n",
- " print('adla compute not found, creating...')\n",
- " provisioning_config = AdlaCompute.provisioning_configuration()\n",
- " adla_compute = ComputeTarget.create(workspace, compute_name, provisioning_config)\n",
- " adla_compute.wait_for_completion()\n",
- " return adla_compute\n",
- " else:\n",
- " raise e\n",
- " \n",
- "adla_compute = get_or_create_adla_compute(ws, adla_compute_name)\n",
- "\n",
- "# CLI:\n",
- "# Create: az ml computetarget setup adla -n \n",
- "# BYOC: az ml computetarget attach adla -n -i "
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Once the above code cell completes, run the below to check your ADLA compute status:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "print(\"ADLA compute state:{}\".format(adla_compute.provisioning_state))\n",
- "print(\"ADLA compute state:{}\".format(adla_compute.provisioning_errors))\n",
- "print(\"Using ADLA compute:{}\".format(adla_compute.cluster_resource_id))"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Create an AdlaStep"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "**AdlaStep** is used to run U-SQL script using Azure Data Lake Analytics.\n",
- "\n",
- "- **name:** Name of module\n",
- "- **script_name:** name of U-SQL script\n",
- "- **inputs:** List of input port bindings\n",
- "- **outputs:** List of output port bindings\n",
- "- **adla_compute:** the ADLA compute to use for this job\n",
- "- **params:** Dictionary of name-value pairs to pass to U-SQL job *(optional)*\n",
- "- **degree_of_parallelism:** the degree of parallelism to use for this job *(optional)*\n",
- "- **priority:** the priority value to use for the current job *(optional)*\n",
- "- **runtime_version:** the runtime version of the Data Lake Analytics engine *(optional)*\n",
- "- **root_folder:** folder that contains the script, assemblies etc. *(optional)*\n",
- "- **hash_paths:** list of paths to hash to detect a change (script file is always hashed) *(optional)*\n",
- "\n",
- "### Remarks\n",
- "\n",
- "You can use `@@name@@` syntax in your script to refer to inputs, outputs, and params.\n",
- "\n",
- "* if `name` is the name of an input or output port binding, any occurences of `@@name@@` in the script\n",
- "are replaced with actual data path of corresponding port binding.\n",
- "* if `name` matches any key in `params` dict, any occurences of `@@name@@` will be replaced with\n",
- "corresponding value in dict.\n",
- "\n",
- "#### Sample script\n",
- "\n",
- "```\n",
- "@resourcereader =\n",
- " EXTRACT query string\n",
- " FROM \"@@script_input@@\"\n",
- " USING Extractors.Csv();\n",
- "\n",
- "\n",
- "OUTPUT @resourcereader\n",
- "TO \"@@script_output@@\"\n",
- "USING Outputters.Csv();\n",
- "```"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "adla_step = AdlaStep(\n",
- " name='adla_script_step',\n",
- " script_name='test_adla_script.usql',\n",
- " inputs=[script_input],\n",
- " outputs=[script_output],\n",
- " compute_target=adla_compute)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Build and Submit the Experiment"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "pipeline = Pipeline(\n",
- " description=\"adla_102\",\n",
- " workspace=ws, \n",
- " steps=[adla_step],\n",
- " default_source_directory=script_folder)\n",
- "\n",
- "pipeline_run = Experiment(workspace, experiment_name).submit(pipeline)\n",
- "pipeline_run.wait_for_completion()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### View Run Details"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.widgets import RunDetails\n",
- "RunDetails(pipeline_run).show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Examine the run\n",
- "You can cycle through the node_run objects and examine job logs, stdout, and stderr of each of the steps."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "step_runs = pipeline_run.get_children()\n",
- "for step_run in step_runs:\n",
- " status = step_run.get_status()\n",
- " print('node', step_run.name, 'status:', status)\n",
- " if status == \"Failed\":\n",
- " joblog = step_run.get_job_log()\n",
- " print('job log:', joblog)\n",
- " stdout_log = step_run.get_stdout_log()\n",
- " print('stdout log:', stdout_log)\n",
- " stderr_log = step_run.get_stderr_log()\n",
- " print('stderr log:', stderr_log)\n",
- " with open(\"logs-\" + step_run.name + \".txt\", \"w\") as f:\n",
- " f.write(joblog)\n",
- " print(\"Job log written to logs-\"+ step_run.name + \".txt\")\n",
- " if status == \"Finished\":\n",
- " stdout_log = step_run.get_stdout_log()\n",
- " print('stdout log:', stdout_log)"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "diray"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved. \n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# AML Pipeline with AdlaStep\n",
+ "This notebook is used to demonstrate the use of AdlaStep in AML Pipeline."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## AML and Pipeline SDK-specific imports"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import os\n",
+ "import azureml.core\n",
+ "from azureml.core.compute import ComputeTarget, DatabricksCompute\n",
+ "from azureml.exceptions import ComputeTargetException\n",
+ "from azureml.core import Workspace, Run, Experiment\n",
+ "from azureml.pipeline.core import Pipeline, PipelineData\n",
+ "from azureml.pipeline.steps import AdlaStep\n",
+ "from azureml.core.datastore import Datastore\n",
+ "from azureml.data.data_reference import DataReference\n",
+ "from azureml.core import attach_legacy_compute_target\n",
+ "\n",
+ "# Check core SDK version number\n",
+ "print(\"SDK version:\", azureml.core.VERSION)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Initialize Workspace\n",
+ "\n",
+ "Initialize a workspace object from persisted configuration. Make sure the config file is present at .\\config.json"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "create workspace"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "ws = Workspace.from_config()\n",
+ "print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep = '\\n')"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "script_folder = '.'\n",
+ "experiment_name = \"adla_101_experiment\"\n",
+ "ws._initialize_folder(experiment_name=experiment_name, directory=script_folder)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Register Datastore"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "\n",
+ "workspace = ws.name\n",
+ "datastore_name='MyAdlsDatastore'\n",
+ "subscription_id=os.getenv(\"ADL_SUBSCRIPTION_62\", \"\") # subscription id of ADLS account\n",
+ "resource_group=os.getenv(\"ADL_RESOURCE_GROUP_62\", \"\") # resource group of ADLS account\n",
+ "store_name=os.getenv(\"ADL_STORENAME_62\", \"\") # ADLS account name\n",
+ "tenant_id=os.getenv(\"ADL_TENANT_62\", \"\") # tenant id of service principal\n",
+ "client_id=os.getenv(\"ADL_CLIENTID_62\", \"\") # client id of service principal\n",
+ "client_secret=os.getenv(\"ADL_CLIENT_62_SECRET\", \"\") # the secret of service principal\n",
+ "\n",
+ "try:\n",
+ " adls_datastore = Datastore.get(ws, datastore_name)\n",
+ " print(\"found datastore with name: %s\" % datastore_name)\n",
+ "except:\n",
+ " adls_datastore = Datastore.register_azure_data_lake(\n",
+ " workspace=ws,\n",
+ " datastore_name=datastore_name,\n",
+ " subscription_id=subscription_id, # subscription id of ADLS account\n",
+ " resource_group=resource_group, # resource group of ADLS account\n",
+ " store_name=store_name, # ADLS account name\n",
+ " tenant_id=tenant_id, # tenant id of service principal\n",
+ " client_id=client_id, # client id of service principal\n",
+ " client_secret=client_secret) # the secret of service principal\n",
+ " print(\"registered datastore with name: %s\" % datastore_name)\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Create DataReferences and PipelineData\n",
+ "\n",
+ "In the code cell below, replace datastorename with your default datastore name. Copy the file `testdata.txt` (located in the pipeline folder that this notebook is in) to the path on the datastore."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "datastorename = \"MyAdlsDatastore\"\n",
+ "\n",
+ "adls_datastore = Datastore(workspace=ws, name=datastorename)\n",
+ "script_input = DataReference(\n",
+ " datastore=adls_datastore,\n",
+ " data_reference_name=\"script_input\",\n",
+ " path_on_datastore=\"testdata/testdata.txt\")\n",
+ "\n",
+ "script_output = PipelineData(\"script_output\", datastore=adls_datastore)\n",
+ "\n",
+ "print(\"Created Pipeline Data\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Setup Data Lake Account\n",
+ "\n",
+ "ADLA can only use data that is located in the default data store associated with that ADLA account. Through Azure portal, check the name of the default data store corresponding to the ADLA account you are using below. Replace the value associated with `adla_compute_name` in the code cell below accordingly."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "adla_compute_name = 'testadl' # Replace this with your default compute\n",
+ "\n",
+ "from azureml.core.compute import ComputeTarget, AdlaCompute\n",
+ "\n",
+ "def get_or_create_adla_compute(workspace, compute_name):\n",
+ " try:\n",
+ " return AdlaCompute(workspace, compute_name)\n",
+ " except ComputeTargetException as e:\n",
+ " if 'ComputeTargetNotFound' in e.message:\n",
+ " print('adla compute not found, creating...')\n",
+ " provisioning_config = AdlaCompute.provisioning_configuration()\n",
+ " adla_compute = ComputeTarget.create(workspace, compute_name, provisioning_config)\n",
+ " adla_compute.wait_for_completion()\n",
+ " return adla_compute\n",
+ " else:\n",
+ " raise e\n",
+ " \n",
+ "adla_compute = get_or_create_adla_compute(ws, adla_compute_name)\n",
+ "\n",
+ "# CLI:\n",
+ "# Create: az ml computetarget setup adla -n \n",
+ "# BYOC: az ml computetarget attach adla -n -i "
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Once the above code cell completes, run the below to check your ADLA compute status:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "print(\"ADLA compute state:{}\".format(adla_compute.provisioning_state))\n",
+ "print(\"ADLA compute state:{}\".format(adla_compute.provisioning_errors))\n",
+ "print(\"Using ADLA compute:{}\".format(adla_compute.cluster_resource_id))"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Create an AdlaStep"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "**AdlaStep** is used to run U-SQL script using Azure Data Lake Analytics.\n",
+ "\n",
+ "- **name:** Name of module\n",
+ "- **script_name:** name of U-SQL script\n",
+ "- **inputs:** List of input port bindings\n",
+ "- **outputs:** List of output port bindings\n",
+ "- **adla_compute:** the ADLA compute to use for this job\n",
+ "- **params:** Dictionary of name-value pairs to pass to U-SQL job *(optional)*\n",
+ "- **degree_of_parallelism:** the degree of parallelism to use for this job *(optional)*\n",
+ "- **priority:** the priority value to use for the current job *(optional)*\n",
+ "- **runtime_version:** the runtime version of the Data Lake Analytics engine *(optional)*\n",
+ "- **root_folder:** folder that contains the script, assemblies etc. *(optional)*\n",
+ "- **hash_paths:** list of paths to hash to detect a change (script file is always hashed) *(optional)*\n",
+ "\n",
+ "### Remarks\n",
+ "\n",
+ "You can use `@@name@@` syntax in your script to refer to inputs, outputs, and params.\n",
+ "\n",
+ "* if `name` is the name of an input or output port binding, any occurences of `@@name@@` in the script\n",
+ "are replaced with actual data path of corresponding port binding.\n",
+ "* if `name` matches any key in `params` dict, any occurences of `@@name@@` will be replaced with\n",
+ "corresponding value in dict.\n",
+ "\n",
+ "#### Sample script\n",
+ "\n",
+ "```\n",
+ "@resourcereader =\n",
+ " EXTRACT query string\n",
+ " FROM \"@@script_input@@\"\n",
+ " USING Extractors.Csv();\n",
+ "\n",
+ "\n",
+ "OUTPUT @resourcereader\n",
+ "TO \"@@script_output@@\"\n",
+ "USING Outputters.Csv();\n",
+ "```"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "adla_step = AdlaStep(\n",
+ " name='adla_script_step',\n",
+ " script_name='test_adla_script.usql',\n",
+ " inputs=[script_input],\n",
+ " outputs=[script_output],\n",
+ " compute_target=adla_compute)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Build and Submit the Experiment"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "pipeline = Pipeline(\n",
+ " description=\"adla_102\",\n",
+ " workspace=ws, \n",
+ " steps=[adla_step],\n",
+ " default_source_directory=script_folder)\n",
+ "\n",
+ "pipeline_run = Experiment(workspace, experiment_name).submit(pipeline)\n",
+ "pipeline_run.wait_for_completion()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### View Run Details"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.widgets import RunDetails\n",
+ "RunDetails(pipeline_run).show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Examine the run\n",
+ "You can cycle through the node_run objects and examine job logs, stdout, and stderr of each of the steps."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "step_runs = pipeline_run.get_children()\n",
+ "for step_run in step_runs:\n",
+ " status = step_run.get_status()\n",
+ " print('node', step_run.name, 'status:', status)\n",
+ " if status == \"Failed\":\n",
+ " joblog = step_run.get_job_log()\n",
+ " print('job log:', joblog)\n",
+ " stdout_log = step_run.get_stdout_log()\n",
+ " print('stdout log:', stdout_log)\n",
+ " stderr_log = step_run.get_stderr_log()\n",
+ " print('stderr log:', stderr_log)\n",
+ " with open(\"logs-\" + step_run.name + \".txt\", \"w\") as f:\n",
+ " f.write(joblog)\n",
+ " print(\"Job log written to logs-\"+ step_run.name + \".txt\")\n",
+ " if status == \"Finished\":\n",
+ " stdout_log = step_run.get_stdout_log()\n",
+ " print('stdout log:', stdout_log)"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python [default]",
- "language": "python",
- "name": "python3"
+ "metadata": {
+ "authors": [
+ {
+ "name": "diray"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.6"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/machine-learning-pipelines/intro-to-pipelines/aml-pipelines-use-databricks-as-compute-target.ipynb b/how-to-use-azureml/machine-learning-pipelines/intro-to-pipelines/aml-pipelines-use-databricks-as-compute-target.ipynb
index c6c3a98a..b08445a7 100644
--- a/how-to-use-azureml/machine-learning-pipelines/intro-to-pipelines/aml-pipelines-use-databricks-as-compute-target.ipynb
+++ b/how-to-use-azureml/machine-learning-pipelines/intro-to-pipelines/aml-pipelines-use-databricks-as-compute-target.ipynb
@@ -1,703 +1,698 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved. \n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Using Databricks as a Compute Target from Azure Machine Learning Pipeline\n",
- "To use Databricks as a compute target from [Azure Machine Learning Pipeline](https://docs.microsoft.com/en-us/azure/machine-learning/service/concept-ml-pipelines), a [DatabricksStep](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-steps/azureml.pipeline.steps.databricks_step.databricksstep?view=azure-ml-py) is used. This notebook demonstrates the use of DatabricksStep in Azure Machine Learning Pipeline.\n",
- "\n",
- "The notebook will show:\n",
- "1. Running an arbitrary Databricks notebook that the customer has in Databricks workspace\n",
- "2. Running an arbitrary Python script that the customer has in DBFS\n",
- "3. Running an arbitrary Python script that is available on local computer (will upload to DBFS, and then run in Databricks) \n",
- "4. Running a JAR job that the customer has in DBFS.\n",
- "\n",
- "## Before you begin:\n",
- "\n",
- "1. **Create an Azure Databricks workspace** in the same subscription where you have your Azure Machine Learning workspace. You will need details of this workspace later on to define DatabricksStep. [Click here](https://ms.portal.azure.com/#blade/HubsExtension/Resources/resourceType/Microsoft.Databricks%2Fworkspaces) for more information.\n",
- "2. **Create PAT (access token)**: Manually create a Databricks access token at the Azure Databricks portal. See [this](https://docs.databricks.com/api/latest/authentication.html#generate-a-token) for more information.\n",
- "3. **Add demo notebook to ADB**: This notebook has a sample you can use as is. Launch Azure Databricks attached to your Azure Machine Learning workspace and add a new notebook. \n",
- "4. **Create/attach a Blob storage** for use from ADB"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Add demo notebook to ADB Workspace\n",
- "Copy and paste the below code to create a new notebook in your ADB workspace."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "```python\n",
- "# direct access\n",
- "dbutils.widgets.get(\"myparam\")\n",
- "p = getArgument(\"myparam\")\n",
- "print (\"Param -\\'myparam':\")\n",
- "print (p)\n",
- "\n",
- "dbutils.widgets.get(\"input\")\n",
- "i = getArgument(\"input\")\n",
- "print (\"Param -\\'input':\")\n",
- "print (i)\n",
- "\n",
- "dbutils.widgets.get(\"output\")\n",
- "o = getArgument(\"output\")\n",
- "print (\"Param -\\'output':\")\n",
- "print (o)\n",
- "\n",
- "n = i + \"/testdata.txt\"\n",
- "df = spark.read.csv(n)\n",
- "\n",
- "display (df)\n",
- "\n",
- "data = [('value1', 'value2')]\n",
- "df2 = spark.createDataFrame(data)\n",
- "\n",
- "z = o + \"/output.txt\"\n",
- "df2.write.csv(z)\n",
- "```"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Azure Machine Learning and Pipeline SDK-specific imports"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import os\n",
- "import azureml.core\n",
- "from azureml.core.runconfig import JarLibrary\n",
- "from azureml.core.compute import ComputeTarget, DatabricksCompute\n",
- "from azureml.exceptions import ComputeTargetException\n",
- "from azureml.core import Workspace, Run, Experiment\n",
- "from azureml.pipeline.core import Pipeline, PipelineData\n",
- "from azureml.pipeline.steps import DatabricksStep\n",
- "from azureml.core.datastore import Datastore\n",
- "from azureml.data.data_reference import DataReference\n",
- "\n",
- "# Check core SDK version number\n",
- "print(\"SDK version:\", azureml.core.VERSION)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Initialize Workspace\n",
- "\n",
- "Initialize a workspace object from persisted configuration. Make sure the config file is present at .\\config.json"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "ws = Workspace.from_config()\n",
- "print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep = '\\n')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Attach Databricks compute target\n",
- "Next, you need to add your Databricks workspace to Azure Machine Learning as a compute target and give it a name. You will use this name to refer to your Databricks workspace compute target inside Azure Machine Learning.\n",
- "\n",
- "- **Resource Group** - The resource group name of your Azure Machine Learning workspace\n",
- "- **Databricks Workspace Name** - The workspace name of your Azure Databricks workspace\n",
- "- **Databricks Access Token** - The access token you created in ADB\n",
- "\n",
- "**The Databricks workspace need to be present in the same subscription as your AML workspace**"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Replace with your account info before running.\n",
- " \n",
- "db_compute_name=os.getenv(\"DATABRICKS_COMPUTE_NAME\", \"\") # Databricks compute name\n",
- "db_resource_group=os.getenv(\"DATABRICKS_RESOURCE_GROUP\", \"\") # Databricks resource group\n",
- "db_workspace_name=os.getenv(\"DATABRICKS_WORKSPACE_NAME\", \"\") # Databricks workspace name\n",
- "db_access_token=os.getenv(\"DATABRICKS_ACCESS_TOKEN\", \"\") # Databricks access token\n",
- " \n",
- "try:\n",
- " databricks_compute = ComputeTarget(workspace=ws, name=db_compute_name)\n",
- " print('Compute target {} already exists'.format(db_compute_name))\n",
- "except ComputeTargetException:\n",
- " print('Compute not found, will use below parameters to attach new one')\n",
- " print('db_compute_name {}'.format(db_compute_name))\n",
- " print('db_resource_group {}'.format(db_resource_group))\n",
- " print('db_workspace_name {}'.format(db_workspace_name))\n",
- " print('db_access_token {}'.format(db_access_token))\n",
- " \n",
- " config = DatabricksCompute.attach_configuration(\n",
- " resource_group = db_resource_group,\n",
- " workspace_name = db_workspace_name,\n",
- " access_token= db_access_token)\n",
- " databricks_compute=ComputeTarget.attach(ws, db_compute_name, config)\n",
- " databricks_compute.wait_for_completion(True)\n"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Data Connections with Inputs and Outputs\n",
- "The DatabricksStep supports Azure Bloband ADLS for inputs and outputs. You also will need to define a [Secrets](https://docs.azuredatabricks.net/user-guide/secrets/index.html) scope to enable authentication to external data sources such as Blob and ADLS from Databricks.\n",
- "\n",
- "- Databricks documentation on [Azure Blob](https://docs.azuredatabricks.net/spark/latest/data-sources/azure/azure-storage.html)\n",
- "- Databricks documentation on [ADLS](https://docs.databricks.com/spark/latest/data-sources/azure/azure-datalake.html)\n",
- "\n",
- "### Type of Data Access\n",
- "Databricks allows to interact with Azure Blob and ADLS in two ways.\n",
- "- **Direct Access**: Databricks allows you to interact with Azure Blob or ADLS URIs directly. The input or output URIs will be mapped to a Databricks widget param in the Databricks notebook.\n",
- "- **Mouting**: You will be supplied with additional parameters and secrets that will enable you to mount your ADLS or Azure Blob input or output location in your Databricks notebook."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Direct Access: Python sample code\n",
- "If you have a data reference named \"input\" it will represent the URI of the input and you can access it directly in the Databricks python notebook like so:"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "```python\n",
- "dbutils.widgets.get(\"input\")\n",
- "y = getArgument(\"input\")\n",
- "df = spark.read.csv(y)\n",
- "```"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Mounting: Python sample code for Azure Blob\n",
- "Given an Azure Blob data reference named \"input\" the following widget params will be made available in the Databricks notebook:"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "```python\n",
- "# This contains the input URI\n",
- "dbutils.widgets.get(\"input\")\n",
- "myinput_uri = getArgument(\"input\")\n",
- "\n",
- "# How to get the input datastore name inside ADB notebook\n",
- "# This contains the name of a Databricks secret (in the predefined \"amlscope\" secret scope) \n",
- "# that contians an access key or sas for the Azure Blob input (this name is obtained by appending \n",
- "# the name of the input with \"_blob_secretname\". \n",
- "dbutils.widgets.get(\"input_blob_secretname\") \n",
- "myinput_blob_secretname = getArgument(\"input_blob_secretname\")\n",
- "\n",
- "# This contains the required configuration for mounting\n",
- "dbutils.widgets.get(\"input_blob_config\")\n",
- "myinput_blob_config = getArgument(\"input_blob_config\")\n",
- "\n",
- "# Usage\n",
- "dbutils.fs.mount(\n",
- " source = myinput_uri,\n",
- " mount_point = \"/mnt/input\",\n",
- " extra_configs = {myinput_blob_config:dbutils.secrets.get(scope = \"amlscope\", key = myinput_blob_secretname)})\n",
- "```"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Mounting: Python sample code for ADLS\n",
- "Given an ADLS data reference named \"input\" the following widget params will be made available in the Databricks notebook:"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "```python\n",
- "# This contains the input URI\n",
- "dbutils.widgets.get(\"input\") \n",
- "myinput_uri = getArgument(\"input\")\n",
- "\n",
- "# This contains the client id for the service principal \n",
- "# that has access to the adls input\n",
- "dbutils.widgets.get(\"input_adls_clientid\") \n",
- "myinput_adls_clientid = getArgument(\"input_adls_clientid\")\n",
- "\n",
- "# This contains the name of a Databricks secret (in the predefined \"amlscope\" secret scope) \n",
- "# that contains the secret for the above mentioned service principal\n",
- "dbutils.widgets.get(\"input_adls_secretname\") \n",
- "myinput_adls_secretname = getArgument(\"input_adls_secretname\")\n",
- "\n",
- "# This contains the refresh url for the mounting configs\n",
- "dbutils.widgets.get(\"input_adls_refresh_url\") \n",
- "myinput_adls_refresh_url = getArgument(\"input_adls_refresh_url\")\n",
- "\n",
- "# Usage \n",
- "configs = {\"dfs.adls.oauth2.access.token.provider.type\": \"ClientCredential\",\n",
- " \"dfs.adls.oauth2.client.id\": myinput_adls_clientid,\n",
- " \"dfs.adls.oauth2.credential\": dbutils.secrets.get(scope = \"amlscope\", key =myinput_adls_secretname),\n",
- " \"dfs.adls.oauth2.refresh.url\": myinput_adls_refresh_url}\n",
- "\n",
- "dbutils.fs.mount(\n",
- " source = myinput_uri,\n",
- " mount_point = \"/mnt/output\",\n",
- " extra_configs = configs)\n",
- "```"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Use Databricks from Azure Machine Learning Pipeline\n",
- "To use Databricks as a compute target from Azure Machine Learning Pipeline, a DatabricksStep is used. Let's define a datasource (via DataReference) and intermediate data (via PipelineData) to be used in DatabricksStep."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Use the default blob storage\n",
- "def_blob_store = Datastore(ws, \"workspaceblobstore\")\n",
- "print('Datastore {} will be used'.format(def_blob_store.name))\n",
- "\n",
- "# We are uploading a sample file in the local directory to be used as a datasource\n",
- "def_blob_store.upload_files([\"./testdata.txt\"], target_path=\"dbtest\", overwrite=False)\n",
- "\n",
- "step_1_input = DataReference(datastore=def_blob_store, path_on_datastore=\"dbtest\",\n",
- " data_reference_name=\"input\")\n",
- "\n",
- "step_1_output = PipelineData(\"output\", datastore=def_blob_store)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Add a DatabricksStep\n",
- "Adds a Databricks notebook as a step in a Pipeline.\n",
- "- ***name:** Name of the Module\n",
- "- **inputs:** List of input connections for data consumed by this step. Fetch this inside the notebook using dbutils.widgets.get(\"input\")\n",
- "- **outputs:** List of output port definitions for outputs produced by this step. Fetch this inside the notebook using dbutils.widgets.get(\"output\")\n",
- "- **existing_cluster_id:** Cluster ID of an existing Interactive cluster on the Databricks workspace. If you are providing this, do not provide any of the parameters below that are used to create a new cluster such as spark_version, node_type, etc.\n",
- "- **spark_version:** Version of spark for the databricks run cluster. default value: 4.0.x-scala2.11\n",
- "- **node_type:** Azure vm node types for the databricks run cluster. default value: Standard_D3_v2\n",
- "- **num_workers:** Number of workers for the databricks run cluster\n",
- "- **autoscale:** The autoscale configuration for the databricks run cluster\n",
- "- **spark_env_variables:** Spark environment variables for the databricks run cluster (dictionary of {str:str}). default value: {'PYSPARK_PYTHON': '/databricks/python3/bin/python3'}\n",
- "- **notebook_path:** Path to the notebook in the databricks instance. If you are providing this, do not provide python script related paramaters or JAR related parameters.\n",
- "- **notebook_params:** Parameters for the databricks notebook (dictionary of {str:str}). Fetch this inside the notebook using dbutils.widgets.get(\"myparam\")\n",
- "- **python_script_path:** The path to the python script in the DBFS or S3. If you are providing this, do not provide python_script_name which is used for uploading script from local machine.\n",
- "- **python_script_params:** Parameters for the python script (list of str)\n",
- "- **main_class_name:** The name of the entry point in a JAR module. If you are providing this, do not provide any python script or notebook related parameters.\n",
- "- **jar_params:** Parameters for the JAR module (list of str)\n",
- "- **python_script_name:** name of a python script on your local machine (relative to source_directory). If you are providing this do not provide python_script_path which is used to execute a remote python script; or any of the JAR or notebook related parameters.\n",
- "- **source_directory:** folder that contains the script and other files\n",
- "- **hash_paths:** list of paths to hash to detect a change in source_directory (script file is always hashed)\n",
- "- **run_name:** Name in databricks for this run\n",
- "- **timeout_seconds:** Timeout for the databricks run\n",
- "- **runconfig:** Runconfig to use. Either pass runconfig or each library type as a separate parameter but do not mix the two\n",
- "- **maven_libraries:** maven libraries for the databricks run\n",
- "- **pypi_libraries:** pypi libraries for the databricks run\n",
- "- **egg_libraries:** egg libraries for the databricks run\n",
- "- **jar_libraries:** jar libraries for the databricks run\n",
- "- **rcran_libraries:** rcran libraries for the databricks run\n",
- "- **compute_target:** Azure Databricks compute\n",
- "- **allow_reuse:** Whether the step should reuse previous results when run with the same settings/inputs\n",
- "- **version:** Optional version tag to denote a change in functionality for the step\n",
- "\n",
- "\\* *denotes required fields* \n",
- "*You must provide exactly one of num_workers or autoscale paramaters* \n",
- "*You must provide exactly one of databricks_compute or databricks_compute_name parameters*\n",
- "\n",
- "## Use runconfig to specify library dependencies\n",
- "You can use a runconfig to specify the library dependencies for your cluster in Databricks. The runconfig will contain a databricks section as follows:\n",
- "```yaml\n",
- "environment:\n",
- "# Databricks details\n",
- " databricks:\n",
- "# List of maven libraries.\n",
- " mavenLibraries:\n",
- " - coordinates: org.jsoup:jsoup:1.7.1\n",
- " repo: ''\n",
- " exclusions:\n",
- " - slf4j:slf4j\n",
- " - '*:hadoop-client'\n",
- "# List of PyPi libraries\n",
- " pypiLibraries:\n",
- " - package: beautifulsoup4\n",
- " repo: ''\n",
- "# List of RCran libraries\n",
- " rcranLibraries:\n",
- " - package: ada\n",
- " repo: http://cran.us.r-project.org\n",
- "# List of JAR libraries\n",
- " jarLibraries:\n",
- " - library: dbfs:/mnt/libraries/library.jar\n",
- "# List of Egg libraries\n",
- " eggLibraries:\n",
- " - library: dbfs:/mnt/libraries/library.egg\n",
- "```\n",
- "\n",
- "You can then create a RunConfiguration object using this file and pass it as the runconfig parameter to DatabricksStep.\n",
- "```python\n",
- "from azureml.core.runconfig import RunConfiguration\n",
- "\n",
- "runconfig = RunConfiguration()\n",
- "runconfig.load(path='', name='')\n",
- "```"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### 1. Running the demo notebook already added to the Databricks workspace\n",
- "Create a notebook in the Azure Databricks workspace, and provide the path to that notebook as the value associated with the environment variable \"DATABRICKS_NOTEBOOK_PATH\". This will then set the variable notebook_path when you run the code cell below:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "notebook_path=os.getenv(\"DATABRICKS_NOTEBOOK_PATH\", \"\") # Databricks notebook path\n",
- "\n",
- "dbNbStep = DatabricksStep(\n",
- " name=\"DBNotebookInWS\",\n",
- " inputs=[step_1_input],\n",
- " outputs=[step_1_output],\n",
- " num_workers=1,\n",
- " notebook_path=notebook_path,\n",
- " notebook_params={'myparam': 'testparam'},\n",
- " run_name='DB_Notebook_demo',\n",
- " compute_target=databricks_compute,\n",
- " allow_reuse=False\n",
- ")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Build and submit the Experiment"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "#PUBLISHONLY\n",
- "#steps = [dbNbStep]\n",
- "#pipeline = Pipeline(workspace=ws, steps=steps)\n",
- "#pipeline_run = Experiment(ws, 'DB_Notebook_demo').submit(pipeline)\n",
- "#pipeline_run.wait_for_completion()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### View Run Details"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "#PUBLISHONLY\n",
- "#from azureml.widgets import RunDetails\n",
- "#RunDetails(pipeline_run).show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### 2. Running a Python script that is already added in DBFS\n",
- "To run a Python script that is already uploaded to DBFS, follow the instructions below. You will first upload the Python script to DBFS using the [CLI](https://docs.azuredatabricks.net/user-guide/dbfs-databricks-file-system.html).\n",
- "\n",
- "The commented out code in the below cell assumes that you have uploaded `train-db-dbfs.py` to the root folder in DBFS. You can upload `train-db-dbfs.py` to the root folder in DBFS using this commandline so you can use `python_script_path = \"dbfs:/train-db-dbfs.py\"`:\n",
- "\n",
- "```\n",
- "dbfs cp ./train-db-dbfs.py dbfs:/train-db-dbfs.py\n",
- "```"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "python_script_path = \"dbfs:/train-db-dbfs.py\"\n",
- "\n",
- "dbPythonInDbfsStep = DatabricksStep(\n",
- " name=\"DBPythonInDBFS\",\n",
- " inputs=[step_1_input],\n",
- " num_workers=1,\n",
- " python_script_path=python_script_path,\n",
- " python_script_params={'--input_data'},\n",
- " run_name='DB_Python_demo',\n",
- " compute_target=databricks_compute,\n",
- " allow_reuse=False\n",
- ")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Build and submit the Experiment"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "#PUBLISHONLY\n",
- "#steps = [dbPythonInDbfsStep]\n",
- "#pipeline = Pipeline(workspace=ws, steps=steps)\n",
- "#pipeline_run = Experiment(ws, 'DB_Python_demo').submit(pipeline)\n",
- "#pipeline_run.wait_for_completion()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### View Run Details"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "#PUBLISHONLY\n",
- "#from azureml.widgets import RunDetails\n",
- "#RunDetails(pipeline_run).show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### 3. Running a Python script in Databricks that currenlty is in local computer\n",
- "To run a Python script that is currently in your local computer, follow the instructions below. \n",
- "\n",
- "The commented out code below code assumes that you have `train-db-local.py` in the `scripts` subdirectory under the current working directory.\n",
- "\n",
- "In this case, the Python script will be uploaded first to DBFS, and then the script will be run in Databricks."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "python_script_name = \"train-db-local.py\"\n",
- "source_directory = \".\"\n",
- "\n",
- "dbPythonInLocalMachineStep = DatabricksStep(\n",
- " name=\"DBPythonInLocalMachine\",\n",
- " inputs=[step_1_input],\n",
- " num_workers=1,\n",
- " python_script_name=python_script_name,\n",
- " source_directory=source_directory,\n",
- " run_name='DB_Python_Local_demo',\n",
- " compute_target=databricks_compute,\n",
- " allow_reuse=False\n",
- ")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Build and submit the Experiment"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "steps = [dbPythonInLocalMachineStep]\n",
- "pipeline = Pipeline(workspace=ws, steps=steps)\n",
- "pipeline_run = Experiment(ws, 'DB_Python_Local_demo').submit(pipeline)\n",
- "pipeline_run.wait_for_completion()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### View Run Details"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.widgets import RunDetails\n",
- "RunDetails(pipeline_run).show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### 4. Running a JAR job that is alreay added in DBFS\n",
- "To run a JAR job that is already uploaded to DBFS, follow the instructions below. You will first upload the JAR file to DBFS using the [CLI](https://docs.azuredatabricks.net/user-guide/dbfs-databricks-file-system.html).\n",
- "\n",
- "The commented out code in the below cell assumes that you have uploaded `train-db-dbfs.jar` to the root folder in DBFS. You can upload `train-db-dbfs.jar` to the root folder in DBFS using this commandline so you can use `jar_library_dbfs_path = \"dbfs:/train-db-dbfs.jar\"`:\n",
- "\n",
- "```\n",
- "dbfs cp ./train-db-dbfs.jar dbfs:/train-db-dbfs.jar\n",
- "```"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "main_jar_class_name = \"com.microsoft.aeva.Main\"\n",
- "jar_library_dbfs_path = \"dbfs:/train-db-dbfs.jar\"\n",
- "\n",
- "dbJarInDbfsStep = DatabricksStep(\n",
- " name=\"DBJarInDBFS\",\n",
- " inputs=[step_1_input],\n",
- " num_workers=1,\n",
- " main_class_name=main_jar_class_name,\n",
- " jar_params={'arg1', 'arg2'},\n",
- " run_name='DB_JAR_demo',\n",
- " jar_libraries=[JarLibrary(jar_library_dbfs_path)],\n",
- " compute_target=databricks_compute,\n",
- " allow_reuse=False\n",
- ")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Build and submit the Experiment"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "#PUBLISHONLY\n",
- "#steps = [dbJarInDbfsStep]\n",
- "#pipeline = Pipeline(workspace=ws, steps=steps)\n",
- "#pipeline_run = Experiment(ws, 'DB_JAR_demo').submit(pipeline)\n",
- "#pipeline_run.wait_for_completion()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### View Run Details"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "#PUBLISHONLY\n",
- "#from azureml.widgets import RunDetails\n",
- "#RunDetails(pipeline_run).show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Next: ADLA as a Compute Target\n",
- "To use ADLA as a compute target from Azure Machine Learning Pipeline, a AdlaStep is used. This [notebook](./aml-pipelines-use-adla-as-compute-target.ipynb) demonstrates the use of AdlaStep in Azure Machine Learning Pipeline."
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "diray"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved. \n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Using Databricks as a Compute Target from Azure Machine Learning Pipeline\n",
+ "To use Databricks as a compute target from [Azure Machine Learning Pipeline](https://docs.microsoft.com/en-us/azure/machine-learning/service/concept-ml-pipelines), a [DatabricksStep](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-steps/azureml.pipeline.steps.databricks_step.databricksstep?view=azure-ml-py) is used. This notebook demonstrates the use of DatabricksStep in Azure Machine Learning Pipeline.\n",
+ "\n",
+ "The notebook will show:\n",
+ "1. Running an arbitrary Databricks notebook that the customer has in Databricks workspace\n",
+ "2. Running an arbitrary Python script that the customer has in DBFS\n",
+ "3. Running an arbitrary Python script that is available on local computer (will upload to DBFS, and then run in Databricks) \n",
+ "4. Running a JAR job that the customer has in DBFS.\n",
+ "\n",
+ "## Before you begin:\n",
+ "\n",
+ "1. **Create an Azure Databricks workspace** in the same subscription where you have your Azure Machine Learning workspace. You will need details of this workspace later on to define DatabricksStep. [Click here](https://ms.portal.azure.com/#blade/HubsExtension/Resources/resourceType/Microsoft.Databricks%2Fworkspaces) for more information.\n",
+ "2. **Create PAT (access token)**: Manually create a Databricks access token at the Azure Databricks portal. See [this](https://docs.databricks.com/api/latest/authentication.html#generate-a-token) for more information.\n",
+ "3. **Add demo notebook to ADB**: This notebook has a sample you can use as is. Launch Azure Databricks attached to your Azure Machine Learning workspace and add a new notebook. \n",
+ "4. **Create/attach a Blob storage** for use from ADB"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Add demo notebook to ADB Workspace\n",
+ "Copy and paste the below code to create a new notebook in your ADB workspace."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "```python\n",
+ "# direct access\n",
+ "dbutils.widgets.get(\"myparam\")\n",
+ "p = getArgument(\"myparam\")\n",
+ "print (\"Param -\\'myparam':\")\n",
+ "print (p)\n",
+ "\n",
+ "dbutils.widgets.get(\"input\")\n",
+ "i = getArgument(\"input\")\n",
+ "print (\"Param -\\'input':\")\n",
+ "print (i)\n",
+ "\n",
+ "dbutils.widgets.get(\"output\")\n",
+ "o = getArgument(\"output\")\n",
+ "print (\"Param -\\'output':\")\n",
+ "print (o)\n",
+ "\n",
+ "n = i + \"/testdata.txt\"\n",
+ "df = spark.read.csv(n)\n",
+ "\n",
+ "display (df)\n",
+ "\n",
+ "data = [('value1', 'value2')]\n",
+ "df2 = spark.createDataFrame(data)\n",
+ "\n",
+ "z = o + \"/output.txt\"\n",
+ "df2.write.csv(z)\n",
+ "```"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Azure Machine Learning and Pipeline SDK-specific imports"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import os\n",
+ "import azureml.core\n",
+ "from azureml.core.runconfig import JarLibrary\n",
+ "from azureml.core.compute import ComputeTarget, DatabricksCompute\n",
+ "from azureml.exceptions import ComputeTargetException\n",
+ "from azureml.core import Workspace, Run, Experiment\n",
+ "from azureml.pipeline.core import Pipeline, PipelineData\n",
+ "from azureml.pipeline.steps import DatabricksStep\n",
+ "from azureml.core.datastore import Datastore\n",
+ "from azureml.data.data_reference import DataReference\n",
+ "\n",
+ "# Check core SDK version number\n",
+ "print(\"SDK version:\", azureml.core.VERSION)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Initialize Workspace\n",
+ "\n",
+ "Initialize a workspace object from persisted configuration. Make sure the config file is present at .\\config.json"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "ws = Workspace.from_config()\n",
+ "print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep = '\\n')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Attach Databricks compute target\n",
+ "Next, you need to add your Databricks workspace to Azure Machine Learning as a compute target and give it a name. You will use this name to refer to your Databricks workspace compute target inside Azure Machine Learning.\n",
+ "\n",
+ "- **Resource Group** - The resource group name of your Azure Machine Learning workspace\n",
+ "- **Databricks Workspace Name** - The workspace name of your Azure Databricks workspace\n",
+ "- **Databricks Access Token** - The access token you created in ADB\n",
+ "\n",
+ "**The Databricks workspace need to be present in the same subscription as your AML workspace**"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Replace with your account info before running.\n",
+ " \n",
+ "db_compute_name=os.getenv(\"DATABRICKS_COMPUTE_NAME\", \"\") # Databricks compute name\n",
+ "db_resource_group=os.getenv(\"DATABRICKS_RESOURCE_GROUP\", \"\") # Databricks resource group\n",
+ "db_workspace_name=os.getenv(\"DATABRICKS_WORKSPACE_NAME\", \"\") # Databricks workspace name\n",
+ "db_access_token=os.getenv(\"DATABRICKS_ACCESS_TOKEN\", \"\") # Databricks access token\n",
+ " \n",
+ "try:\n",
+ " databricks_compute = ComputeTarget(workspace=ws, name=db_compute_name)\n",
+ " print('Compute target {} already exists'.format(db_compute_name))\n",
+ "except ComputeTargetException:\n",
+ " print('Compute not found, will use below parameters to attach new one')\n",
+ " print('db_compute_name {}'.format(db_compute_name))\n",
+ " print('db_resource_group {}'.format(db_resource_group))\n",
+ " print('db_workspace_name {}'.format(db_workspace_name))\n",
+ " print('db_access_token {}'.format(db_access_token))\n",
+ " \n",
+ " config = DatabricksCompute.attach_configuration(\n",
+ " resource_group = db_resource_group,\n",
+ " workspace_name = db_workspace_name,\n",
+ " access_token= db_access_token)\n",
+ " databricks_compute=ComputeTarget.attach(ws, db_compute_name, config)\n",
+ " databricks_compute.wait_for_completion(True)\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Data Connections with Inputs and Outputs\n",
+ "The DatabricksStep supports Azure Bloband ADLS for inputs and outputs. You also will need to define a [Secrets](https://docs.azuredatabricks.net/user-guide/secrets/index.html) scope to enable authentication to external data sources such as Blob and ADLS from Databricks.\n",
+ "\n",
+ "- Databricks documentation on [Azure Blob](https://docs.azuredatabricks.net/spark/latest/data-sources/azure/azure-storage.html)\n",
+ "- Databricks documentation on [ADLS](https://docs.databricks.com/spark/latest/data-sources/azure/azure-datalake.html)\n",
+ "\n",
+ "### Type of Data Access\n",
+ "Databricks allows to interact with Azure Blob and ADLS in two ways.\n",
+ "- **Direct Access**: Databricks allows you to interact with Azure Blob or ADLS URIs directly. The input or output URIs will be mapped to a Databricks widget param in the Databricks notebook.\n",
+ "- **Mouting**: You will be supplied with additional parameters and secrets that will enable you to mount your ADLS or Azure Blob input or output location in your Databricks notebook."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Direct Access: Python sample code\n",
+ "If you have a data reference named \"input\" it will represent the URI of the input and you can access it directly in the Databricks python notebook like so:"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "```python\n",
+ "dbutils.widgets.get(\"input\")\n",
+ "y = getArgument(\"input\")\n",
+ "df = spark.read.csv(y)\n",
+ "```"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Mounting: Python sample code for Azure Blob\n",
+ "Given an Azure Blob data reference named \"input\" the following widget params will be made available in the Databricks notebook:"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "```python\n",
+ "# This contains the input URI\n",
+ "dbutils.widgets.get(\"input\")\n",
+ "myinput_uri = getArgument(\"input\")\n",
+ "\n",
+ "# How to get the input datastore name inside ADB notebook\n",
+ "# This contains the name of a Databricks secret (in the predefined \"amlscope\" secret scope) \n",
+ "# that contians an access key or sas for the Azure Blob input (this name is obtained by appending \n",
+ "# the name of the input with \"_blob_secretname\". \n",
+ "dbutils.widgets.get(\"input_blob_secretname\") \n",
+ "myinput_blob_secretname = getArgument(\"input_blob_secretname\")\n",
+ "\n",
+ "# This contains the required configuration for mounting\n",
+ "dbutils.widgets.get(\"input_blob_config\")\n",
+ "myinput_blob_config = getArgument(\"input_blob_config\")\n",
+ "\n",
+ "# Usage\n",
+ "dbutils.fs.mount(\n",
+ " source = myinput_uri,\n",
+ " mount_point = \"/mnt/input\",\n",
+ " extra_configs = {myinput_blob_config:dbutils.secrets.get(scope = \"amlscope\", key = myinput_blob_secretname)})\n",
+ "```"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Mounting: Python sample code for ADLS\n",
+ "Given an ADLS data reference named \"input\" the following widget params will be made available in the Databricks notebook:"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "```python\n",
+ "# This contains the input URI\n",
+ "dbutils.widgets.get(\"input\") \n",
+ "myinput_uri = getArgument(\"input\")\n",
+ "\n",
+ "# This contains the client id for the service principal \n",
+ "# that has access to the adls input\n",
+ "dbutils.widgets.get(\"input_adls_clientid\") \n",
+ "myinput_adls_clientid = getArgument(\"input_adls_clientid\")\n",
+ "\n",
+ "# This contains the name of a Databricks secret (in the predefined \"amlscope\" secret scope) \n",
+ "# that contains the secret for the above mentioned service principal\n",
+ "dbutils.widgets.get(\"input_adls_secretname\") \n",
+ "myinput_adls_secretname = getArgument(\"input_adls_secretname\")\n",
+ "\n",
+ "# This contains the refresh url for the mounting configs\n",
+ "dbutils.widgets.get(\"input_adls_refresh_url\") \n",
+ "myinput_adls_refresh_url = getArgument(\"input_adls_refresh_url\")\n",
+ "\n",
+ "# Usage \n",
+ "configs = {\"dfs.adls.oauth2.access.token.provider.type\": \"ClientCredential\",\n",
+ " \"dfs.adls.oauth2.client.id\": myinput_adls_clientid,\n",
+ " \"dfs.adls.oauth2.credential\": dbutils.secrets.get(scope = \"amlscope\", key =myinput_adls_secretname),\n",
+ " \"dfs.adls.oauth2.refresh.url\": myinput_adls_refresh_url}\n",
+ "\n",
+ "dbutils.fs.mount(\n",
+ " source = myinput_uri,\n",
+ " mount_point = \"/mnt/output\",\n",
+ " extra_configs = configs)\n",
+ "```"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Use Databricks from Azure Machine Learning Pipeline\n",
+ "To use Databricks as a compute target from Azure Machine Learning Pipeline, a DatabricksStep is used. Let's define a datasource (via DataReference) and intermediate data (via PipelineData) to be used in DatabricksStep."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Use the default blob storage\n",
+ "def_blob_store = Datastore(ws, \"workspaceblobstore\")\n",
+ "print('Datastore {} will be used'.format(def_blob_store.name))\n",
+ "\n",
+ "# We are uploading a sample file in the local directory to be used as a datasource\n",
+ "def_blob_store.upload_files([\"./testdata.txt\"], target_path=\"dbtest\", overwrite=False)\n",
+ "\n",
+ "step_1_input = DataReference(datastore=def_blob_store, path_on_datastore=\"dbtest\",\n",
+ " data_reference_name=\"input\")\n",
+ "\n",
+ "step_1_output = PipelineData(\"output\", datastore=def_blob_store)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Add a DatabricksStep\n",
+ "Adds a Databricks notebook as a step in a Pipeline.\n",
+ "- ***name:** Name of the Module\n",
+ "- **inputs:** List of input connections for data consumed by this step. Fetch this inside the notebook using dbutils.widgets.get(\"input\")\n",
+ "- **outputs:** List of output port definitions for outputs produced by this step. Fetch this inside the notebook using dbutils.widgets.get(\"output\")\n",
+ "- **existing_cluster_id:** Cluster ID of an existing Interactive cluster on the Databricks workspace. If you are providing this, do not provide any of the parameters below that are used to create a new cluster such as spark_version, node_type, etc.\n",
+ "- **spark_version:** Version of spark for the databricks run cluster. default value: 4.0.x-scala2.11\n",
+ "- **node_type:** Azure vm node types for the databricks run cluster. default value: Standard_D3_v2\n",
+ "- **num_workers:** Specifies a static number of workers for the databricks run cluster\n",
+ "- **min_workers:** Specifies a min number of workers to use for auto-scaling the databricks run cluster\n",
+ "- **max_workers:** Specifies a max number of workers to use for auto-scaling the databricks run cluster\n",
+ "- **spark_env_variables:** Spark environment variables for the databricks run cluster (dictionary of {str:str}). default value: {'PYSPARK_PYTHON': '/databricks/python3/bin/python3'}\n",
+ "- **notebook_path:** Path to the notebook in the databricks instance. If you are providing this, do not provide python script related paramaters or JAR related parameters.\n",
+ "- **notebook_params:** Parameters for the databricks notebook (dictionary of {str:str}). Fetch this inside the notebook using dbutils.widgets.get(\"myparam\")\n",
+ "- **python_script_path:** The path to the python script in the DBFS or S3. If you are providing this, do not provide python_script_name which is used for uploading script from local machine.\n",
+ "- **python_script_params:** Parameters for the python script (list of str)\n",
+ "- **main_class_name:** The name of the entry point in a JAR module. If you are providing this, do not provide any python script or notebook related parameters.\n",
+ "- **jar_params:** Parameters for the JAR module (list of str)\n",
+ "- **python_script_name:** name of a python script on your local machine (relative to source_directory). If you are providing this do not provide python_script_path which is used to execute a remote python script; or any of the JAR or notebook related parameters.\n",
+ "- **source_directory:** folder that contains the script and other files\n",
+ "- **hash_paths:** list of paths to hash to detect a change in source_directory (script file is always hashed)\n",
+ "- **run_name:** Name in databricks for this run\n",
+ "- **timeout_seconds:** Timeout for the databricks run\n",
+ "- **runconfig:** Runconfig to use. Either pass runconfig or each library type as a separate parameter but do not mix the two\n",
+ "- **maven_libraries:** maven libraries for the databricks run\n",
+ "- **pypi_libraries:** pypi libraries for the databricks run\n",
+ "- **egg_libraries:** egg libraries for the databricks run\n",
+ "- **jar_libraries:** jar libraries for the databricks run\n",
+ "- **rcran_libraries:** rcran libraries for the databricks run\n",
+ "- **compute_target:** Azure Databricks compute\n",
+ "- **allow_reuse:** Whether the step should reuse previous results when run with the same settings/inputs\n",
+ "- **version:** Optional version tag to denote a change in functionality for the step\n",
+ "\n",
+ "\\* *denotes required fields* \n",
+ "*You must provide exactly one of num_workers or min_workers and max_workers paramaters* \n",
+ "*You must provide exactly one of databricks_compute or databricks_compute_name parameters*\n",
+ "\n",
+ "## Use runconfig to specify library dependencies\n",
+ "You can use a runconfig to specify the library dependencies for your cluster in Databricks. The runconfig will contain a databricks section as follows:\n",
+ "```yaml\n",
+ "environment:\n",
+ "# Databricks details\n",
+ " databricks:\n",
+ "# List of maven libraries.\n",
+ " mavenLibraries:\n",
+ " - coordinates: org.jsoup:jsoup:1.7.1\n",
+ " repo: ''\n",
+ " exclusions:\n",
+ " - slf4j:slf4j\n",
+ " - '*:hadoop-client'\n",
+ "# List of PyPi libraries\n",
+ " pypiLibraries:\n",
+ " - package: beautifulsoup4\n",
+ " repo: ''\n",
+ "# List of RCran libraries\n",
+ " rcranLibraries:\n",
+ " - package: ada\n",
+ " repo: http://cran.us.r-project.org\n",
+ "# List of JAR libraries\n",
+ " jarLibraries:\n",
+ " - library: dbfs:/mnt/libraries/library.jar\n",
+ "# List of Egg libraries\n",
+ " eggLibraries:\n",
+ " - library: dbfs:/mnt/libraries/library.egg\n",
+ "```\n",
+ "\n",
+ "You can then create a RunConfiguration object using this file and pass it as the runconfig parameter to DatabricksStep.\n",
+ "```python\n",
+ "from azureml.core.runconfig import RunConfiguration\n",
+ "\n",
+ "runconfig = RunConfiguration()\n",
+ "runconfig.load(path='', name='')\n",
+ "```"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### 1. Running the demo notebook already added to the Databricks workspace\n",
+ "Create a notebook in the Azure Databricks workspace, and provide the path to that notebook as the value associated with the environment variable \"DATABRICKS_NOTEBOOK_PATH\". This will then set the variable\u00c2\u00a0notebook_path\u00c2\u00a0when you run the code cell below:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "notebook_path=os.getenv(\"DATABRICKS_NOTEBOOK_PATH\", \"\") # Databricks notebook path\n",
+ "\n",
+ "dbNbStep = DatabricksStep(\n",
+ " name=\"DBNotebookInWS\",\n",
+ " inputs=[step_1_input],\n",
+ " outputs=[step_1_output],\n",
+ " num_workers=1,\n",
+ " notebook_path=notebook_path,\n",
+ " notebook_params={'myparam': 'testparam'},\n",
+ " run_name='DB_Notebook_demo',\n",
+ " compute_target=databricks_compute,\n",
+ " allow_reuse=False\n",
+ ")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Build and submit the Experiment"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "steps = [dbNbStep]\n",
+ "pipeline = Pipeline(workspace=ws, steps=steps)\n",
+ "pipeline_run = Experiment(ws, 'DB_Notebook_demo').submit(pipeline)\n",
+ "pipeline_run.wait_for_completion()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### View Run Details"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.widgets import RunDetails\n",
+ "RunDetails(pipeline_run).show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### 2. Running a Python script that is already added in DBFS\n",
+ "To run a Python script that is already uploaded to DBFS, follow the instructions below. You will first upload the Python script to DBFS using the [CLI](https://docs.azuredatabricks.net/user-guide/dbfs-databricks-file-system.html).\n",
+ "\n",
+ "The commented out code in the below cell assumes that you have uploaded `train-db-dbfs.py` to the root folder in DBFS. You can upload `train-db-dbfs.py` to the root folder in DBFS using this commandline so you can use `python_script_path = \"dbfs:/train-db-dbfs.py\"`:\n",
+ "\n",
+ "```\n",
+ "dbfs cp ./train-db-dbfs.py dbfs:/train-db-dbfs.py\n",
+ "```"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "python_script_path = \"dbfs:/train-db-dbfs.py\"\n",
+ "\n",
+ "dbPythonInDbfsStep = DatabricksStep(\n",
+ " name=\"DBPythonInDBFS\",\n",
+ " inputs=[step_1_input],\n",
+ " num_workers=1,\n",
+ " python_script_path=python_script_path,\n",
+ " python_script_params={'--input_data'},\n",
+ " run_name='DB_Python_demo',\n",
+ " compute_target=databricks_compute,\n",
+ " allow_reuse=False\n",
+ ")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Build and submit the Experiment"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "steps = [dbPythonInDbfsStep]\n",
+ "pipeline = Pipeline(workspace=ws, steps=steps)\n",
+ "pipeline_run = Experiment(ws, 'DB_Python_demo').submit(pipeline)\n",
+ "pipeline_run.wait_for_completion()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### View Run Details"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.widgets import RunDetails\n",
+ "RunDetails(pipeline_run).show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### 3. Running a Python script in Databricks that currenlty is in local computer\n",
+ "To run a Python script that is currently in your local computer, follow the instructions below. \n",
+ "\n",
+ "The commented out code below code assumes that you have `train-db-local.py` in the `scripts` subdirectory under the current working directory.\n",
+ "\n",
+ "In this case, the Python script will be uploaded first to DBFS, and then the script will be run in Databricks."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "python_script_name = \"train-db-local.py\"\n",
+ "source_directory = \".\"\n",
+ "\n",
+ "dbPythonInLocalMachineStep = DatabricksStep(\n",
+ " name=\"DBPythonInLocalMachine\",\n",
+ " inputs=[step_1_input],\n",
+ " num_workers=1,\n",
+ " python_script_name=python_script_name,\n",
+ " source_directory=source_directory,\n",
+ " run_name='DB_Python_Local_demo',\n",
+ " compute_target=databricks_compute,\n",
+ " allow_reuse=False\n",
+ ")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Build and submit the Experiment"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "steps = [dbPythonInLocalMachineStep]\n",
+ "pipeline = Pipeline(workspace=ws, steps=steps)\n",
+ "pipeline_run = Experiment(ws, 'DB_Python_Local_demo').submit(pipeline)\n",
+ "pipeline_run.wait_for_completion()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### View Run Details"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.widgets import RunDetails\n",
+ "RunDetails(pipeline_run).show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### 4. Running a JAR job that is alreay added in DBFS\n",
+ "To run a JAR job that is already uploaded to DBFS, follow the instructions below. You will first upload the JAR file to DBFS using the [CLI](https://docs.azuredatabricks.net/user-guide/dbfs-databricks-file-system.html).\n",
+ "\n",
+ "The commented out code in the below cell assumes that you have uploaded `train-db-dbfs.jar` to the root folder in DBFS. You can upload `train-db-dbfs.jar` to the root folder in DBFS using this commandline so you can use `jar_library_dbfs_path = \"dbfs:/train-db-dbfs.jar\"`:\n",
+ "\n",
+ "```\n",
+ "dbfs cp ./train-db-dbfs.jar dbfs:/train-db-dbfs.jar\n",
+ "```"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "main_jar_class_name = \"com.microsoft.aeva.Main\"\n",
+ "jar_library_dbfs_path = \"dbfs:/train-db-dbfs.jar\"\n",
+ "\n",
+ "dbJarInDbfsStep = DatabricksStep(\n",
+ " name=\"DBJarInDBFS\",\n",
+ " inputs=[step_1_input],\n",
+ " num_workers=1,\n",
+ " main_class_name=main_jar_class_name,\n",
+ " jar_params={'arg1', 'arg2'},\n",
+ " run_name='DB_JAR_demo',\n",
+ " jar_libraries=[JarLibrary(jar_library_dbfs_path)],\n",
+ " compute_target=databricks_compute,\n",
+ " allow_reuse=False\n",
+ ")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Build and submit the Experiment"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "steps = [dbJarInDbfsStep]\n",
+ "pipeline = Pipeline(workspace=ws, steps=steps)\n",
+ "pipeline_run = Experiment(ws, 'DB_JAR_demo').submit(pipeline)\n",
+ "pipeline_run.wait_for_completion()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### View Run Details"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.widgets import RunDetails\n",
+ "RunDetails(pipeline_run).show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Next: ADLA as a Compute Target\n",
+ "To use ADLA as a compute target from Azure Machine Learning Pipeline, a AdlaStep is used. This [notebook](./aml-pipelines-use-adla-as-compute-target.ipynb) demonstrates the use of AdlaStep in Azure Machine Learning Pipeline."
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3",
- "language": "python",
- "name": "python3"
+ "metadata": {
+ "authors": [
+ {
+ "name": "diray"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.2"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.2"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/machine-learning-pipelines/intro-to-pipelines/aml-pipelines-with-data-dependency-steps.ipynb b/how-to-use-azureml/machine-learning-pipelines/intro-to-pipelines/aml-pipelines-with-data-dependency-steps.ipynb
index 642512d2..e2a883b9 100644
--- a/how-to-use-azureml/machine-learning-pipelines/intro-to-pipelines/aml-pipelines-with-data-dependency-steps.ipynb
+++ b/how-to-use-azureml/machine-learning-pipelines/intro-to-pipelines/aml-pipelines-with-data-dependency-steps.ipynb
@@ -1,418 +1,418 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved. \n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Azure Machine Learning Pipelines with Data Dependency\n",
- "In this notebook, we will see how we can build a pipeline with implicit data dependancy."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Prerequisites and Azure Machine Learning Basics\n",
- "Make sure you go through the configuration Notebook located at https://github.com/Azure/MachineLearningNotebooks first if you haven't. This sets you up with a working config file that has information on your workspace, subscription id, etc. \n",
- "\n",
- "### Azure Machine Learning and Pipeline SDK-specific Imports"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import azureml.core\n",
- "from azureml.core import Workspace, Run, Experiment, Datastore\n",
- "from azureml.core.compute import AmlCompute\n",
- "from azureml.core.compute import ComputeTarget\n",
- "from azureml.core.compute import DataFactoryCompute\n",
- "from azureml.widgets import RunDetails\n",
- "\n",
- "# Check core SDK version number\n",
- "print(\"SDK version:\", azureml.core.VERSION)\n",
- "\n",
- "from azureml.data.data_reference import DataReference\n",
- "from azureml.pipeline.core import Pipeline, PipelineData, StepSequence\n",
- "from azureml.pipeline.steps import PythonScriptStep\n",
- "from azureml.pipeline.steps import DataTransferStep\n",
- "from azureml.pipeline.core import PublishedPipeline\n",
- "from azureml.pipeline.core.graph import PipelineParameter\n",
- "\n",
- "print(\"Pipeline SDK-specific imports completed\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Initialize Workspace\n",
- "\n",
- "Initialize a [workspace](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.workspace(class%29) object from persisted configuration."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "create workspace"
- ]
- },
- "outputs": [],
- "source": [
- "ws = Workspace.from_config()\n",
- "print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep = '\\n')\n",
- "\n",
- "# Default datastore (Azure file storage)\n",
- "def_file_store = ws.get_default_datastore() \n",
- "print(\"Default datastore's name: {}\".format(def_file_store.name))\n",
- "\n",
- "def_blob_store = Datastore(ws, \"workspaceblobstore\")\n",
- "print(\"Blobstore's name: {}\".format(def_blob_store.name))"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# project folder\n",
- "project_folder = '.'\n",
- " \n",
- "print('Sample projects will be created in {}.'.format(project_folder))"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Required data and script files for the the tutorial\n",
- "Sample files required to finish this tutorial are already copied to the project folder specified above. Even though the .py provided in the samples don't have much \"ML work,\" as a data scientist, you will work on this extensively as part of your work. To complete this tutorial, the contents of these files are not very important. The one-line files are for demostration purpose only."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Compute Targets\n",
- "See the list of Compute Targets on the workspace."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "cts = ws.compute_targets\n",
- "for ct in cts:\n",
- " print(ct)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Retrieve or create a Aml compute\n",
- "Azure Machine Learning Compute is a service for provisioning and managing clusters of Azure virtual machines for running machine learning workloads. Let's create a new Aml Compute in the current workspace, if it doesn't already exist. We will then run the training script on this compute target."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "\n",
- "aml_compute_target = \"aml-compute\"\n",
- "try:\n",
- " aml_compute = AmlCompute(ws, aml_compute_target)\n",
- " print(\"found existing compute target.\")\n",
- "except:\n",
- " print(\"creating new compute target\")\n",
- " \n",
- " provisioning_config = AmlCompute.provisioning_configuration(vm_size = \"STANDARD_D2_V2\",\n",
- " min_nodes = 1, \n",
- " max_nodes = 4) \n",
- " aml_compute = ComputeTarget.create(ws, aml_compute_target, provisioning_config)\n",
- " aml_compute.wait_for_completion(show_output=True, min_node_count=None, timeout_in_minutes=20)\n",
- " \n",
- "print(\"Aml Compute attached\")\n"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# For a more detailed view of current Azure Machine Learning Compute status, use the 'status' property\n",
- "# example: un-comment the following line.\n",
- "# print(aml_compute.status.serialize())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "**Wait for this call to finish before proceeding (you will see the asterisk turning to a number).**\n",
- "\n",
- "Now that you have created the compute target, let's see what the workspace's compute_targets() function returns. You should now see one entry named 'amlcompute' of type AmlCompute."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Building Pipeline Steps with Inputs and Outputs\n",
- "As mentioned earlier, a step in the pipeline can take data as input. This data can be a data source that lives in one of the accessible data locations, or intermediate data produced by a previous step in the pipeline.\n",
- "\n",
- "### Datasources\n",
- "Datasource is represented by **[DataReference](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.data.data_reference.datareference?view=azure-ml-py)** object and points to data that lives in or is accessible from Datastore. DataReference could be a pointer to a file or a directory."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Reference the data uploaded to blob storage using DataReference\n",
- "# Assign the datasource to blob_input_data variable\n",
- "\n",
- "# DataReference(datastore, \n",
- "# data_reference_name=None, \n",
- "# path_on_datastore=None, \n",
- "# mode='mount', \n",
- "# path_on_compute=None, \n",
- "# overwrite=False)\n",
- "\n",
- "blob_input_data = DataReference(\n",
- " datastore=def_blob_store,\n",
- " data_reference_name=\"test_data\",\n",
- " path_on_datastore=\"20newsgroups/20news.pkl\")\n",
- "print(\"DataReference object created\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Intermediate/Output Data\n",
- "Intermediate data (or output of a Step) is represented by **[PipelineData](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-core/azureml.pipeline.core.pipelinedata?view=azure-ml-py)** object. PipelineData can be produced by one step and consumed in another step by providing the PipelineData object as an output of one step and the input of one or more steps.\n",
- "\n",
- "#### Constructing PipelineData\n",
- "- **name:** [*Required*] Name of the data item within the pipeline graph\n",
- "- **datastore_name:** Name of the Datastore to write this output to\n",
- "- **output_name:** Name of the output\n",
- "- **output_mode:** Specifies \"upload\" or \"mount\" modes for producing output (default: mount)\n",
- "- **output_path_on_compute:** For \"upload\" mode, the path to which the module writes this output during execution\n",
- "- **output_overwrite:** Flag to overwrite pre-existing data"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Define intermediate data using PipelineData\n",
- "# Syntax\n",
- "\n",
- "# PipelineData(name, \n",
- "# datastore=None, \n",
- "# output_name=None, \n",
- "# output_mode='mount', \n",
- "# output_path_on_compute=None, \n",
- "# output_overwrite=None, \n",
- "# data_type=None, \n",
- "# is_directory=None)\n",
- "\n",
- "# Naming the intermediate data as processed_data1 and assigning it to the variable processed_data1.\n",
- "processed_data1 = PipelineData(\"processed_data1\",datastore=def_blob_store)\n",
- "print(\"PipelineData object created\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Pipelines steps using datasources and intermediate data\n",
- "Machine learning pipelines can have many steps and these steps could use or reuse datasources and intermediate data. Here's how we construct such a pipeline:"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Define a Step that consumes a datasource and produces intermediate data.\n",
- "In this step, we define a step that consumes a datasource and produces intermediate data.\n",
- "\n",
- "**Open `train.py` in the local machine and examine the arguments, inputs, and outputs for the script. That will give you a good sense of why the script argument names used below are important.** "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# step4 consumes the datasource (Datareference) in the previous step\n",
- "# and produces processed_data1\n",
- "trainStep = PythonScriptStep(\n",
- " script_name=\"train.py\", \n",
- " arguments=[\"--input_data\", blob_input_data, \"--output_train\", processed_data1],\n",
- " inputs=[blob_input_data],\n",
- " outputs=[processed_data1],\n",
- " compute_target=aml_compute, \n",
- " source_directory=project_folder\n",
- ")\n",
- "print(\"trainStep created\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Define a Step that consumes intermediate data and produces intermediate data\n",
- "In this step, we define a step that consumes an intermediate data and produces intermediate data.\n",
- "\n",
- "**Open `extract.py` in the local machine and examine the arguments, inputs, and outputs for the script. That will give you a good sense of why the script argument names used below are important.** "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# step5 to use the intermediate data produced by step4\n",
- "# This step also produces an output processed_data2\n",
- "processed_data2 = PipelineData(\"processed_data2\", datastore=def_blob_store)\n",
- "\n",
- "extractStep = PythonScriptStep(\n",
- " script_name=\"extract.py\",\n",
- " arguments=[\"--input_extract\", processed_data1, \"--output_extract\", processed_data2],\n",
- " inputs=[processed_data1],\n",
- " outputs=[processed_data2],\n",
- " compute_target=aml_compute, \n",
- " source_directory=project_folder)\n",
- "print(\"extractStep created\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Define a Step that consumes multiple intermediate data and produces intermediate data\n",
- "In this step, we define a step that consumes multiple intermediate data and produces intermediate data.\n",
- "\n",
- "**Open `compare.py` in the local machine and examine the arguments, inputs, and outputs for the script. That will give you a good sense of why the script argument names used below are important.**"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Now define step6 that takes two inputs (both intermediate data), and produce an output\n",
- "processed_data3 = PipelineData(\"processed_data3\", datastore=def_blob_store)\n",
- "\n",
- "compareStep = PythonScriptStep(\n",
- " script_name=\"compare.py\",\n",
- " arguments=[\"--compare_data1\", processed_data1, \"--compare_data2\", processed_data2, \"--output_compare\", processed_data3],\n",
- " inputs=[processed_data1, processed_data2],\n",
- " outputs=[processed_data3], \n",
- " compute_target=aml_compute, \n",
- " source_directory=project_folder)\n",
- "print(\"compareStep created\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Build the pipeline"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "pipeline1 = Pipeline(workspace=ws, steps=[compareStep])\n",
- "print (\"Pipeline is built\")\n",
- "\n",
- "pipeline1.validate()\n",
- "print(\"Simple validation complete\") "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "pipeline_run1 = Experiment(ws, 'Data_dependency').submit(pipeline1)\n",
- "print(\"Pipeline is submitted for execution\")"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "RunDetails(pipeline_run1).show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Next: Publishing the Pipeline and calling it from the REST endpoint\n",
- "See this [notebook](./aml-pipelines-publish-and-run-using-rest-endpoint.ipynb) to understand how the pipeline is published and you can call the REST endpoint to run the pipeline."
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "diray"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved. \n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Azure Machine Learning Pipelines with Data Dependency\n",
+ "In this notebook, we will see how we can build a pipeline with implicit data dependancy."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Prerequisites and Azure Machine Learning Basics\n",
+ "Make sure you go through the configuration Notebook located at https://github.com/Azure/MachineLearningNotebooks first if you haven't. This sets you up with a working config file that has information on your workspace, subscription id, etc. \n",
+ "\n",
+ "### Azure Machine Learning and Pipeline SDK-specific Imports"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import azureml.core\n",
+ "from azureml.core import Workspace, Run, Experiment, Datastore\n",
+ "from azureml.core.compute import AmlCompute\n",
+ "from azureml.core.compute import ComputeTarget\n",
+ "from azureml.core.compute import DataFactoryCompute\n",
+ "from azureml.widgets import RunDetails\n",
+ "\n",
+ "# Check core SDK version number\n",
+ "print(\"SDK version:\", azureml.core.VERSION)\n",
+ "\n",
+ "from azureml.data.data_reference import DataReference\n",
+ "from azureml.pipeline.core import Pipeline, PipelineData, StepSequence\n",
+ "from azureml.pipeline.steps import PythonScriptStep\n",
+ "from azureml.pipeline.steps import DataTransferStep\n",
+ "from azureml.pipeline.core import PublishedPipeline\n",
+ "from azureml.pipeline.core.graph import PipelineParameter\n",
+ "\n",
+ "print(\"Pipeline SDK-specific imports completed\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Initialize Workspace\n",
+ "\n",
+ "Initialize a [workspace](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.workspace(class%29) object from persisted configuration."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "create workspace"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "ws = Workspace.from_config()\n",
+ "print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep = '\\n')\n",
+ "\n",
+ "# Default datastore (Azure file storage)\n",
+ "def_file_store = ws.get_default_datastore() \n",
+ "print(\"Default datastore's name: {}\".format(def_file_store.name))\n",
+ "\n",
+ "def_blob_store = Datastore(ws, \"workspaceblobstore\")\n",
+ "print(\"Blobstore's name: {}\".format(def_blob_store.name))"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# project folder\n",
+ "project_folder = '.'\n",
+ " \n",
+ "print('Sample projects will be created in {}.'.format(project_folder))"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Required data and script files for the the tutorial\n",
+ "Sample files required to finish this tutorial are already copied to the project folder specified above. Even though the .py provided in the samples don't have much \"ML work,\" as a data scientist, you will work on this extensively as part of your work. To complete this tutorial, the contents of these files are not very important. The one-line files are for demostration purpose only."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Compute Targets\n",
+ "See the list of Compute Targets on the workspace."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "cts = ws.compute_targets\n",
+ "for ct in cts:\n",
+ " print(ct)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Retrieve or create a Aml compute\n",
+ "Azure Machine Learning Compute is a service for provisioning and managing clusters of Azure virtual machines for running machine learning workloads. Let's create a new Aml Compute in the current workspace, if it doesn't already exist. We will then run the training script on this compute target."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "\n",
+ "aml_compute_target = \"aml-compute\"\n",
+ "try:\n",
+ " aml_compute = AmlCompute(ws, aml_compute_target)\n",
+ " print(\"found existing compute target.\")\n",
+ "except:\n",
+ " print(\"creating new compute target\")\n",
+ " \n",
+ " provisioning_config = AmlCompute.provisioning_configuration(vm_size = \"STANDARD_D2_V2\",\n",
+ " min_nodes = 1, \n",
+ " max_nodes = 4) \n",
+ " aml_compute = ComputeTarget.create(ws, aml_compute_target, provisioning_config)\n",
+ " aml_compute.wait_for_completion(show_output=True, min_node_count=None, timeout_in_minutes=20)\n",
+ " \n",
+ "print(\"Aml Compute attached\")\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# For a more detailed view of current Azure Machine Learning Compute status, use get_status()\n",
+ "# example: un-comment the following line.\n",
+ "# print(aml_compute.get_status().serialize())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "**Wait for this call to finish before proceeding (you will see the asterisk turning to a number).**\n",
+ "\n",
+ "Now that you have created the compute target, let's see what the workspace's compute_targets() function returns. You should now see one entry named 'amlcompute' of type AmlCompute."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Building Pipeline Steps with Inputs and Outputs\n",
+ "As mentioned earlier, a step in the pipeline can take data as input. This data can be a data source that lives in one of the accessible data locations, or intermediate data produced by a previous step in the pipeline.\n",
+ "\n",
+ "### Datasources\n",
+ "Datasource is represented by **[DataReference](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.data.data_reference.datareference?view=azure-ml-py)** object and points to data that lives in or is accessible from Datastore. DataReference could be a pointer to a file or a directory."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Reference the data uploaded to blob storage using DataReference\n",
+ "# Assign the datasource to blob_input_data variable\n",
+ "\n",
+ "# DataReference(datastore, \n",
+ "# data_reference_name=None, \n",
+ "# path_on_datastore=None, \n",
+ "# mode='mount', \n",
+ "# path_on_compute=None, \n",
+ "# overwrite=False)\n",
+ "\n",
+ "blob_input_data = DataReference(\n",
+ " datastore=def_blob_store,\n",
+ " data_reference_name=\"test_data\",\n",
+ " path_on_datastore=\"20newsgroups/20news.pkl\")\n",
+ "print(\"DataReference object created\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Intermediate/Output Data\n",
+ "Intermediate data (or output of a Step) is represented by **[PipelineData](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-core/azureml.pipeline.core.pipelinedata?view=azure-ml-py)** object. PipelineData can be produced by one step and consumed in another step by providing the PipelineData object as an output of one step and the input of one or more steps.\n",
+ "\n",
+ "#### Constructing PipelineData\n",
+ "- **name:** [*Required*] Name of the data item within the pipeline graph\n",
+ "- **datastore_name:** Name of the Datastore to write this output to\n",
+ "- **output_name:** Name of the output\n",
+ "- **output_mode:** Specifies \"upload\" or \"mount\" modes for producing output (default: mount)\n",
+ "- **output_path_on_compute:** For \"upload\" mode, the path to which the module writes this output during execution\n",
+ "- **output_overwrite:** Flag to overwrite pre-existing data"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Define intermediate data using PipelineData\n",
+ "# Syntax\n",
+ "\n",
+ "# PipelineData(name, \n",
+ "# datastore=None, \n",
+ "# output_name=None, \n",
+ "# output_mode='mount', \n",
+ "# output_path_on_compute=None, \n",
+ "# output_overwrite=None, \n",
+ "# data_type=None, \n",
+ "# is_directory=None)\n",
+ "\n",
+ "# Naming the intermediate data as processed_data1 and assigning it to the variable processed_data1.\n",
+ "processed_data1 = PipelineData(\"processed_data1\",datastore=def_blob_store)\n",
+ "print(\"PipelineData object created\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Pipelines steps using datasources and intermediate data\n",
+ "Machine learning pipelines can have many steps and these steps could use or reuse datasources and intermediate data. Here's how we construct such a pipeline:"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Define a Step that consumes a datasource and produces intermediate data.\n",
+ "In this step, we define a step that consumes a datasource and produces intermediate data.\n",
+ "\n",
+ "**Open `train.py` in the local machine and examine the arguments, inputs, and outputs for the script. That will give you a good sense of why the script argument names used below are important.** "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# step4 consumes the datasource (Datareference) in the previous step\n",
+ "# and produces processed_data1\n",
+ "trainStep = PythonScriptStep(\n",
+ " script_name=\"train.py\", \n",
+ " arguments=[\"--input_data\", blob_input_data, \"--output_train\", processed_data1],\n",
+ " inputs=[blob_input_data],\n",
+ " outputs=[processed_data1],\n",
+ " compute_target=aml_compute, \n",
+ " source_directory=project_folder\n",
+ ")\n",
+ "print(\"trainStep created\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Define a Step that consumes intermediate data and produces intermediate data\n",
+ "In this step, we define a step that consumes an intermediate data and produces intermediate data.\n",
+ "\n",
+ "**Open `extract.py` in the local machine and examine the arguments, inputs, and outputs for the script. That will give you a good sense of why the script argument names used below are important.** "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# step5 to use the intermediate data produced by step4\n",
+ "# This step also produces an output processed_data2\n",
+ "processed_data2 = PipelineData(\"processed_data2\", datastore=def_blob_store)\n",
+ "\n",
+ "extractStep = PythonScriptStep(\n",
+ " script_name=\"extract.py\",\n",
+ " arguments=[\"--input_extract\", processed_data1, \"--output_extract\", processed_data2],\n",
+ " inputs=[processed_data1],\n",
+ " outputs=[processed_data2],\n",
+ " compute_target=aml_compute, \n",
+ " source_directory=project_folder)\n",
+ "print(\"extractStep created\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Define a Step that consumes multiple intermediate data and produces intermediate data\n",
+ "In this step, we define a step that consumes multiple intermediate data and produces intermediate data.\n",
+ "\n",
+ "**Open `compare.py` in the local machine and examine the arguments, inputs, and outputs for the script. That will give you a good sense of why the script argument names used below are important.**"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Now define step6 that takes two inputs (both intermediate data), and produce an output\n",
+ "processed_data3 = PipelineData(\"processed_data3\", datastore=def_blob_store)\n",
+ "\n",
+ "compareStep = PythonScriptStep(\n",
+ " script_name=\"compare.py\",\n",
+ " arguments=[\"--compare_data1\", processed_data1, \"--compare_data2\", processed_data2, \"--output_compare\", processed_data3],\n",
+ " inputs=[processed_data1, processed_data2],\n",
+ " outputs=[processed_data3], \n",
+ " compute_target=aml_compute, \n",
+ " source_directory=project_folder)\n",
+ "print(\"compareStep created\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Build the pipeline"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "pipeline1 = Pipeline(workspace=ws, steps=[compareStep])\n",
+ "print (\"Pipeline is built\")\n",
+ "\n",
+ "pipeline1.validate()\n",
+ "print(\"Simple validation complete\") "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "pipeline_run1 = Experiment(ws, 'Data_dependency').submit(pipeline1)\n",
+ "print(\"Pipeline is submitted for execution\")"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "RunDetails(pipeline_run1).show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Next: Publishing the Pipeline and calling it from the REST endpoint\n",
+ "See this [notebook](./aml-pipelines-publish-and-run-using-rest-endpoint.ipynb) to understand how the pipeline is published and you can call the REST endpoint to run the pipeline."
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3",
- "language": "python",
- "name": "python3"
+ "metadata": {
+ "authors": [
+ {
+ "name": "diray"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.7"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.7"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/machine-learning-pipelines/pipeline-batch-scoring/pipeline-batch-scoring.ipynb b/how-to-use-azureml/machine-learning-pipelines/pipeline-batch-scoring/pipeline-batch-scoring.ipynb
index db73c513..89ec7f32 100644
--- a/how-to-use-azureml/machine-learning-pipelines/pipeline-batch-scoring/pipeline-batch-scoring.ipynb
+++ b/how-to-use-azureml/machine-learning-pipelines/pipeline-batch-scoring/pipeline-batch-scoring.ipynb
@@ -1,573 +1,573 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved. \n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Using Azure Machine Learning Pipelines for batch prediction\n",
- "\n",
- "In this notebook we will demonstrate how to run a batch scoring job using Azure Machine Learning pipelines. Our example job will be to take an already-trained image classification model, and run that model on some unlabeled images. The image classification model that we'll use is the __[Inception-V3 model](https://arxiv.org/abs/1512.00567)__ and we'll run this model on unlabeled images from the __[ImageNet](http://image-net.org/)__ dataset. \n",
- "\n",
- "The outline of this notebook is as follows:\n",
- "\n",
- "- Register the pretrained inception model into the model registry. \n",
- "- Store the dataset images in a blob container.\n",
- "- Use the registered model to do batch scoring on the images in the data blob container."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Prerequisites\n",
- "Make sure you go through the configuration Notebook located at https://github.com/Azure/MachineLearningNotebooks first if you haven't. This sets you up with a working config file that has information on your workspace, subscription id, etc. "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core import Datastore\n",
- "from azureml.core import Experiment\n",
- "from azureml.core.compute import AmlCompute, ComputeTarget\n",
- "from azureml.core.conda_dependencies import CondaDependencies\n",
- "from azureml.core.datastore import Datastore\n",
- "from azureml.core.runconfig import CondaDependencies, RunConfiguration\n",
- "from azureml.data.data_reference import DataReference\n",
- "from azureml.pipeline.core import Pipeline, PipelineData\n",
- "from azureml.pipeline.steps import PythonScriptStep"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import os\n",
- "from azureml.core import Workspace, Run, Experiment\n",
- "\n",
- "ws = Workspace.from_config()\n",
- "print('Workspace name: ' + ws.name, \n",
- " 'Azure region: ' + ws.location, \n",
- " 'Subscription id: ' + ws.subscription_id, \n",
- " 'Resource group: ' + ws.resource_group, sep = '\\n')\n"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Set up machine learning resources"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Set up datastores\n",
- "First, let’s access the datastore that has the model, labels, and images. \n",
- "\n",
- "### Create a datastore that points to a blob container containing sample images\n",
- "\n",
- "We have created a public blob container `sampledata` on an account named `pipelinedata`, containing images from the ImageNet evaluation set. In the next step, we create a datastore with the name `images_datastore`, which points to this container. In the call to `register_azure_blob_container` below, setting the `overwrite` flag to `True` overwrites any datastore that was created previously with that name. \n",
- "\n",
- "This step can be changed to point to your blob container by providing your own `datastore_name`, `container_name`, and `account_name`."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "account_name = \"pipelinedata\"\n",
- "datastore_name=\"images_datastore\"\n",
- "container_name=\"sampledata\"\n",
- "\n",
- "batchscore_blob = Datastore.register_azure_blob_container(ws, \n",
- " datastore_name=datastore_name, \n",
- " container_name= container_name, \n",
- " account_name=account_name, \n",
- " overwrite=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Next, let’s specify the default datastore for the outputs."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "def_data_store = ws.get_default_datastore()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Configure data references\n",
- "Now you need to add references to the data, as inputs to the appropriate pipeline steps in your pipeline. A data source in a pipeline is represented by a DataReference object. The DataReference object points to data that lives in, or is accessible from, a datastore. We need DataReference objects corresponding to the following: the directory containing the input images, the directory in which the pretrained model is stored, the directory containing the labels, and the output directory."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "input_images = DataReference(datastore=batchscore_blob, \n",
- " data_reference_name=\"input_images\",\n",
- " path_on_datastore=\"batchscoring/images\",\n",
- " mode=\"download\"\n",
- " )\n",
- "model_dir = DataReference(datastore=batchscore_blob, \n",
- " data_reference_name=\"input_model\",\n",
- " path_on_datastore=\"batchscoring/models\",\n",
- " mode=\"download\" \n",
- " )\n",
- "label_dir = DataReference(datastore=batchscore_blob, \n",
- " data_reference_name=\"input_labels\",\n",
- " path_on_datastore=\"batchscoring/labels\",\n",
- " mode=\"download\" \n",
- " )\n",
- "output_dir = PipelineData(name=\"scores\", \n",
- " datastore=def_data_store, \n",
- " output_path_on_compute=\"batchscoring/results\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create and attach Compute targets\n",
- "Use the below code to create and attach Compute targets. "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import os\n",
- "\n",
- "# choose a name for your cluster\n",
- "aml_compute_name = os.environ.get(\"AML_COMPUTE_NAME\", \"gpu-cluster\")\n",
- "cluster_min_nodes = os.environ.get(\"AML_COMPUTE_MIN_NODES\", 0)\n",
- "cluster_max_nodes = os.environ.get(\"AML_COMPUTE_MAX_NODES\", 1)\n",
- "vm_size = os.environ.get(\"AML_COMPUTE_SKU\", \"STANDARD_NC6\")\n",
- "\n",
- "\n",
- "if aml_compute_name in ws.compute_targets:\n",
- " compute_target = ws.compute_targets[aml_compute_name]\n",
- " if compute_target and type(compute_target) is AmlCompute:\n",
- " print('found compute target. just use it. ' + aml_compute_name)\n",
- "else:\n",
- " print('creating a new compute target...')\n",
- " provisioning_config = AmlCompute.provisioning_configuration(vm_size = vm_size, # NC6 is GPU-enabled\n",
- " vm_priority = 'lowpriority', # optional\n",
- " min_nodes = cluster_min_nodes, \n",
- " max_nodes = cluster_max_nodes)\n",
- "\n",
- " # create the cluster\n",
- " compute_target = ComputeTarget.create(ws, aml_compute_name, provisioning_config)\n",
- " \n",
- " # can poll for a minimum number of nodes and for a specific timeout. \n",
- " # if no min node count is provided it will use the scale settings for the cluster\n",
- " compute_target.wait_for_completion(show_output=True, min_node_count=None, timeout_in_minutes=20)\n",
- " \n",
- " # For a more detailed view of current Azure Machine Learning Compute status, use the 'status' property \n",
- " print(compute_target.status.serialize())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Prepare the Model"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Download the Model\n",
- "\n",
- "Download and extract the model from http://download.tensorflow.org/models/inception_v3_2016_08_28.tar.gz to `\"models\"`"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# create directory for model\n",
- "model_dir = 'models'\n",
- "if not os.path.isdir(model_dir):\n",
- " os.mkdir(model_dir)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import tarfile\n",
- "import urllib.request\n",
- "\n",
- "url=\"http://download.tensorflow.org/models/inception_v3_2016_08_28.tar.gz\"\n",
- "response = urllib.request.urlretrieve(url, \"model.tar.gz\")\n",
- "tar = tarfile.open(\"model.tar.gz\", \"r:gz\")\n",
- "tar.extractall(model_dir)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Register the model with Workspace"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import shutil\n",
- "from azureml.core.model import Model\n",
- "\n",
- "# register downloaded model \n",
- "model = Model.register(model_path = \"models/inception_v3.ckpt\",\n",
- " model_name = \"inception\", # this is the name the model is registered as\n",
- " tags = {'pretrained': \"inception\"},\n",
- " description = \"Imagenet trained tensorflow inception\",\n",
- " workspace = ws)\n",
- "# remove the downloaded dir after registration if you wish\n",
- "shutil.rmtree(\"models\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Write your scoring script"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "To do the scoring, we use a batch scoring script `batch_scoring.py`, which is located in the same directory that this notebook is in. You can take a look at this script to see how you might modify it for your custom batch scoring task.\n",
- "\n",
- "The python script `batch_scoring.py` takes input images, applies the image classification model to these images, and outputs a classification result to a results file.\n",
- "\n",
- "The script `batch_scoring.py` takes the following parameters:\n",
- "\n",
- "- `--model_name`: the name of the model being used, which is expected to be in the `model_dir` directory\n",
- "- `--label_dir` : the directory holding the `labels.txt` file \n",
- "- `--dataset_path`: the directory containing the input images\n",
- "- `--output_dir` : the script will run the model on the data and output a `results-label.txt` to this directory\n",
- "- `--batch_size` : the batch size used in running the model.\n"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Build and run the batch scoring pipeline\n",
- "You have everything you need to build the pipeline. Let’s put all these together."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Specify the environment to run the script\n",
- "Specify the conda dependencies for your script. You will need this object when you create the pipeline step later on."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.runconfig import DEFAULT_GPU_IMAGE\n",
- "\n",
- "cd = CondaDependencies.create(pip_packages=[\"tensorflow-gpu==1.10.0\", \"azureml-defaults\"])\n",
- "\n",
- "# Runconfig\n",
- "amlcompute_run_config = RunConfiguration(conda_dependencies=cd)\n",
- "amlcompute_run_config.environment.docker.enabled = True\n",
- "amlcompute_run_config.environment.docker.gpu_support = True\n",
- "amlcompute_run_config.environment.docker.base_image = DEFAULT_GPU_IMAGE\n",
- "amlcompute_run_config.environment.spark.precache_packages = False"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Specify the parameters for your pipeline\n",
- "A subset of the parameters to the python script can be given as input when we re-run a `PublishedPipeline`. In the current example, we define `batch_size` taken by the script as such parameter."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.pipeline.core.graph import PipelineParameter\n",
- "batch_size_param = PipelineParameter(name=\"param_batch_size\", default_value=20)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create the pipeline step\n",
- "Create the pipeline step using the script, environment configuration, and parameters. Specify the compute target you already attached to your workspace as the target of execution of the script. We will use PythonScriptStep to create the pipeline step."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "inception_model_name = \"inception_v3.ckpt\"\n",
- "\n",
- "batch_score_step = PythonScriptStep(\n",
- " name=\"batch_scoring\",\n",
- " script_name=\"batch_scoring.py\",\n",
- " arguments=[\"--dataset_path\", input_images, \n",
- " \"--model_name\", \"inception\",\n",
- " \"--label_dir\", label_dir, \n",
- " \"--output_dir\", output_dir, \n",
- " \"--batch_size\", batch_size_param],\n",
- " compute_target=compute_target,\n",
- " inputs=[input_images, label_dir],\n",
- " outputs=[output_dir],\n",
- " runconfig=amlcompute_run_config\n",
- ")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Run the pipeline\n",
- "At this point you can run the pipeline and examine the output it produced. "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "pipeline = Pipeline(workspace=ws, steps=[batch_score_step])\n",
- "pipeline_run = Experiment(ws, 'batch_scoring').submit(pipeline, pipeline_params={\"param_batch_size\": 20})"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Monitor the run"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.widgets import RunDetails\n",
- "RunDetails(pipeline_run).show()"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "pipeline_run.wait_for_completion(show_output=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Download and review output"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "step_run = list(pipeline_run.get_children())[0]\n",
- "step_run.download_file(\"./outputs/result-labels.txt\")"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import pandas as pd\n",
- "df = pd.read_csv(\"result-labels.txt\", delimiter=\":\", header=None)\n",
- "df.columns = [\"Filename\", \"Prediction\"]\n",
- "df.head()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Publish a pipeline and rerun using a REST call"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create a published pipeline\n",
- "Once you are satisfied with the outcome of the run, you can publish the pipeline to run it with different input values later. When you publish a pipeline, you will get a REST endpoint that accepts invoking of the pipeline with the set of parameters you have already incorporated above using PipelineParameter."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "published_pipeline = pipeline_run.publish_pipeline(\n",
- " name=\"Inception_v3_scoring\", description=\"Batch scoring using Inception v3 model\", version=\"1.0\")\n",
- "\n",
- "published_id = published_pipeline.id"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Rerun the pipeline using the REST endpoint"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Get AAD token"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.authentication import AzureCliAuthentication\n",
- "import requests\n",
- "\n",
- "cli_auth = AzureCliAuthentication()\n",
- "aad_token = cli_auth.get_authentication_header()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Run published pipeline"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.pipeline.core import PublishedPipeline\n",
- "\n",
- "rest_endpoint = published_pipeline.endpoint\n",
- "# specify batch size when running the pipeline\n",
- "response = requests.post(rest_endpoint, \n",
- " headers=aad_token, \n",
- " json={\"ExperimentName\": \"batch_scoring\",\n",
- " \"ParameterAssignments\": {\"param_batch_size\": 50}})\n",
- "run_id = response.json()[\"Id\"]"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Monitor the new run"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.pipeline.core.run import PipelineRun\n",
- "published_pipeline_run = PipelineRun(ws.experiments[\"batch_scoring\"], run_id)\n",
- "\n",
- "RunDetails(published_pipeline_run).show()"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "hichando"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved. \n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Using Azure Machine Learning Pipelines for batch prediction\n",
+ "\n",
+ "In this notebook we will demonstrate how to run a batch scoring job using Azure Machine Learning pipelines. Our example job will be to take an already-trained image classification model, and run that model on some unlabeled images. The image classification model that we'll use is the __[Inception-V3 model](https://arxiv.org/abs/1512.00567)__ and we'll run this model on unlabeled images from the __[ImageNet](http://image-net.org/)__ dataset. \n",
+ "\n",
+ "The outline of this notebook is as follows:\n",
+ "\n",
+ "- Register the pretrained inception model into the model registry. \n",
+ "- Store the dataset images in a blob container.\n",
+ "- Use the registered model to do batch scoring on the images in the data blob container."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Prerequisites\n",
+ "Make sure you go through the configuration Notebook located at https://github.com/Azure/MachineLearningNotebooks first if you haven't. This sets you up with a working config file that has information on your workspace, subscription id, etc. "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core import Datastore\n",
+ "from azureml.core import Experiment\n",
+ "from azureml.core.compute import AmlCompute, ComputeTarget\n",
+ "from azureml.core.conda_dependencies import CondaDependencies\n",
+ "from azureml.core.datastore import Datastore\n",
+ "from azureml.core.runconfig import CondaDependencies, RunConfiguration\n",
+ "from azureml.data.data_reference import DataReference\n",
+ "from azureml.pipeline.core import Pipeline, PipelineData\n",
+ "from azureml.pipeline.steps import PythonScriptStep"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import os\n",
+ "from azureml.core import Workspace, Run, Experiment\n",
+ "\n",
+ "ws = Workspace.from_config()\n",
+ "print('Workspace name: ' + ws.name, \n",
+ " 'Azure region: ' + ws.location, \n",
+ " 'Subscription id: ' + ws.subscription_id, \n",
+ " 'Resource group: ' + ws.resource_group, sep = '\\n')\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Set up machine learning resources"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Set up datastores\n",
+ "First, let\u00e2\u20ac\u2122s access the datastore that has the model, labels, and images. \n",
+ "\n",
+ "### Create a datastore that points to a blob container containing sample images\n",
+ "\n",
+ "We have created a public blob container `sampledata` on an account named `pipelinedata`, containing images from the ImageNet evaluation set. In the next step, we create a datastore with the name `images_datastore`, which points to this container. In the call to `register_azure_blob_container` below, setting the `overwrite` flag to `True` overwrites any datastore that was created previously with that name. \n",
+ "\n",
+ "This step can be changed to point to your blob container by providing your own `datastore_name`, `container_name`, and `account_name`."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "account_name = \"pipelinedata\"\n",
+ "datastore_name=\"images_datastore\"\n",
+ "container_name=\"sampledata\"\n",
+ "\n",
+ "batchscore_blob = Datastore.register_azure_blob_container(ws, \n",
+ " datastore_name=datastore_name, \n",
+ " container_name= container_name, \n",
+ " account_name=account_name, \n",
+ " overwrite=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Next, let\u00e2\u20ac\u2122s specify the default datastore for the outputs."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "def_data_store = ws.get_default_datastore()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Configure data references\n",
+ "Now you need to add references to the data, as inputs to the appropriate pipeline steps in your pipeline. A data source in a pipeline is represented by a DataReference object. The DataReference object points to data that lives in, or is accessible from, a datastore. We need DataReference objects corresponding to the following: the directory containing the input images, the directory in which the pretrained model is stored, the directory containing the labels, and the output directory."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "input_images = DataReference(datastore=batchscore_blob, \n",
+ " data_reference_name=\"input_images\",\n",
+ " path_on_datastore=\"batchscoring/images\",\n",
+ " mode=\"download\"\n",
+ " )\n",
+ "model_dir = DataReference(datastore=batchscore_blob, \n",
+ " data_reference_name=\"input_model\",\n",
+ " path_on_datastore=\"batchscoring/models\",\n",
+ " mode=\"download\" \n",
+ " )\n",
+ "label_dir = DataReference(datastore=batchscore_blob, \n",
+ " data_reference_name=\"input_labels\",\n",
+ " path_on_datastore=\"batchscoring/labels\",\n",
+ " mode=\"download\" \n",
+ " )\n",
+ "output_dir = PipelineData(name=\"scores\", \n",
+ " datastore=def_data_store, \n",
+ " output_path_on_compute=\"batchscoring/results\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create and attach Compute targets\n",
+ "Use the below code to create and attach Compute targets. "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import os\n",
+ "\n",
+ "# choose a name for your cluster\n",
+ "aml_compute_name = os.environ.get(\"AML_COMPUTE_NAME\", \"gpu-cluster\")\n",
+ "cluster_min_nodes = os.environ.get(\"AML_COMPUTE_MIN_NODES\", 0)\n",
+ "cluster_max_nodes = os.environ.get(\"AML_COMPUTE_MAX_NODES\", 1)\n",
+ "vm_size = os.environ.get(\"AML_COMPUTE_SKU\", \"STANDARD_NC6\")\n",
+ "\n",
+ "\n",
+ "if aml_compute_name in ws.compute_targets:\n",
+ " compute_target = ws.compute_targets[aml_compute_name]\n",
+ " if compute_target and type(compute_target) is AmlCompute:\n",
+ " print('found compute target. just use it. ' + aml_compute_name)\n",
+ "else:\n",
+ " print('creating a new compute target...')\n",
+ " provisioning_config = AmlCompute.provisioning_configuration(vm_size = vm_size, # NC6 is GPU-enabled\n",
+ " vm_priority = 'lowpriority', # optional\n",
+ " min_nodes = cluster_min_nodes, \n",
+ " max_nodes = cluster_max_nodes)\n",
+ "\n",
+ " # create the cluster\n",
+ " compute_target = ComputeTarget.create(ws, aml_compute_name, provisioning_config)\n",
+ " \n",
+ " # can poll for a minimum number of nodes and for a specific timeout. \n",
+ " # if no min node count is provided it will use the scale settings for the cluster\n",
+ " compute_target.wait_for_completion(show_output=True, min_node_count=None, timeout_in_minutes=20)\n",
+ " \n",
+ " # For a more detailed view of current Azure Machine Learning Compute status, use get_status()\n",
+ " print(compute_target.get_status().serialize())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Prepare the Model"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Download the Model\n",
+ "\n",
+ "Download and extract the model from http://download.tensorflow.org/models/inception_v3_2016_08_28.tar.gz to `\"models\"`"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# create directory for model\n",
+ "model_dir = 'models'\n",
+ "if not os.path.isdir(model_dir):\n",
+ " os.mkdir(model_dir)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import tarfile\n",
+ "import urllib.request\n",
+ "\n",
+ "url=\"http://download.tensorflow.org/models/inception_v3_2016_08_28.tar.gz\"\n",
+ "response = urllib.request.urlretrieve(url, \"model.tar.gz\")\n",
+ "tar = tarfile.open(\"model.tar.gz\", \"r:gz\")\n",
+ "tar.extractall(model_dir)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Register the model with Workspace"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import shutil\n",
+ "from azureml.core.model import Model\n",
+ "\n",
+ "# register downloaded model \n",
+ "model = Model.register(model_path = \"models/inception_v3.ckpt\",\n",
+ " model_name = \"inception\", # this is the name the model is registered as\n",
+ " tags = {'pretrained': \"inception\"},\n",
+ " description = \"Imagenet trained tensorflow inception\",\n",
+ " workspace = ws)\n",
+ "# remove the downloaded dir after registration if you wish\n",
+ "shutil.rmtree(\"models\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Write your scoring script"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "To do the scoring, we use a batch scoring script `batch_scoring.py`, which is located in the same directory that this notebook is in. You can take a look at this script to see how you might modify it for your custom batch scoring task.\n",
+ "\n",
+ "The python script `batch_scoring.py` takes input images, applies the image classification model to these images, and outputs a classification result to a results file.\n",
+ "\n",
+ "The script `batch_scoring.py` takes the following parameters:\n",
+ "\n",
+ "- `--model_name`: the name of the model being used, which is expected to be in the `model_dir` directory\n",
+ "- `--label_dir` : the directory holding the `labels.txt` file \n",
+ "- `--dataset_path`: the directory containing the input images\n",
+ "- `--output_dir` : the script will run the model on the data and output a `results-label.txt` to this directory\n",
+ "- `--batch_size` : the batch size used in running the model.\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Build and run the batch scoring pipeline\n",
+ "You have everything you need to build the pipeline. Let\u00e2\u20ac\u2122s put all these together."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Specify the environment to run the script\n",
+ "Specify the conda dependencies for your script. You will need this object when you create the pipeline step later on."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.runconfig import DEFAULT_GPU_IMAGE\n",
+ "\n",
+ "cd = CondaDependencies.create(pip_packages=[\"tensorflow-gpu==1.10.0\", \"azureml-defaults\"])\n",
+ "\n",
+ "# Runconfig\n",
+ "amlcompute_run_config = RunConfiguration(conda_dependencies=cd)\n",
+ "amlcompute_run_config.environment.docker.enabled = True\n",
+ "amlcompute_run_config.environment.docker.gpu_support = True\n",
+ "amlcompute_run_config.environment.docker.base_image = DEFAULT_GPU_IMAGE\n",
+ "amlcompute_run_config.environment.spark.precache_packages = False"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Specify the parameters for your pipeline\n",
+ "A subset of the parameters to the python script can be given as input when we re-run a `PublishedPipeline`. In the current example, we define `batch_size` taken by the script as such parameter."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.pipeline.core.graph import PipelineParameter\n",
+ "batch_size_param = PipelineParameter(name=\"param_batch_size\", default_value=20)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create the pipeline step\n",
+ "Create the pipeline step using the script, environment configuration, and parameters. Specify the compute target you already attached to your workspace as the target of execution of the script. We will use PythonScriptStep to create the pipeline step."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "inception_model_name = \"inception_v3.ckpt\"\n",
+ "\n",
+ "batch_score_step = PythonScriptStep(\n",
+ " name=\"batch_scoring\",\n",
+ " script_name=\"batch_scoring.py\",\n",
+ " arguments=[\"--dataset_path\", input_images, \n",
+ " \"--model_name\", \"inception\",\n",
+ " \"--label_dir\", label_dir, \n",
+ " \"--output_dir\", output_dir, \n",
+ " \"--batch_size\", batch_size_param],\n",
+ " compute_target=compute_target,\n",
+ " inputs=[input_images, label_dir],\n",
+ " outputs=[output_dir],\n",
+ " runconfig=amlcompute_run_config\n",
+ ")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Run the pipeline\n",
+ "At this point you can run the pipeline and examine the output it produced. "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "pipeline = Pipeline(workspace=ws, steps=[batch_score_step])\n",
+ "pipeline_run = Experiment(ws, 'batch_scoring').submit(pipeline, pipeline_params={\"param_batch_size\": 20})"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Monitor the run"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.widgets import RunDetails\n",
+ "RunDetails(pipeline_run).show()"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "pipeline_run.wait_for_completion(show_output=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Download and review output"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "step_run = list(pipeline_run.get_children())[0]\n",
+ "step_run.download_file(\"./outputs/result-labels.txt\")"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import pandas as pd\n",
+ "df = pd.read_csv(\"result-labels.txt\", delimiter=\":\", header=None)\n",
+ "df.columns = [\"Filename\", \"Prediction\"]\n",
+ "df.head()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Publish a pipeline and rerun using a REST call"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create a published pipeline\n",
+ "Once you are satisfied with the outcome of the run, you can publish the pipeline to run it with different input values later. When you publish a pipeline, you will get a REST endpoint that accepts invoking of the pipeline with the set of parameters you have already incorporated above using PipelineParameter."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "published_pipeline = pipeline_run.publish_pipeline(\n",
+ " name=\"Inception_v3_scoring\", description=\"Batch scoring using Inception v3 model\", version=\"1.0\")\n",
+ "\n",
+ "published_id = published_pipeline.id"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Rerun the pipeline using the REST endpoint"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Get AAD token"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.authentication import AzureCliAuthentication\n",
+ "import requests\n",
+ "\n",
+ "cli_auth = AzureCliAuthentication()\n",
+ "aad_token = cli_auth.get_authentication_header()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Run published pipeline"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.pipeline.core import PublishedPipeline\n",
+ "\n",
+ "rest_endpoint = published_pipeline.endpoint\n",
+ "# specify batch size when running the pipeline\n",
+ "response = requests.post(rest_endpoint, \n",
+ " headers=aad_token, \n",
+ " json={\"ExperimentName\": \"batch_scoring\",\n",
+ " \"ParameterAssignments\": {\"param_batch_size\": 50}})\n",
+ "run_id = response.json()[\"Id\"]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Monitor the new run"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.pipeline.core.run import PipelineRun\n",
+ "published_pipeline_run = PipelineRun(ws.experiments[\"batch_scoring\"], run_id)\n",
+ "\n",
+ "RunDetails(published_pipeline_run).show()"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3",
- "language": "python",
- "name": "python3"
+ "metadata": {
+ "authors": [
+ {
+ "name": "hichando"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.7"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.7"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/machine-learning-pipelines/pipeline-style-transfer/pipeline-style-transfer.ipynb b/how-to-use-azureml/machine-learning-pipelines/pipeline-style-transfer/pipeline-style-transfer.ipynb
index b1b6674c..8aa6966e 100644
--- a/how-to-use-azureml/machine-learning-pipelines/pipeline-style-transfer/pipeline-style-transfer.ipynb
+++ b/how-to-use-azureml/machine-learning-pipelines/pipeline-style-transfer/pipeline-style-transfer.ipynb
@@ -1,610 +1,610 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Neural style transfer on video\n",
- "Using modified code from `pytorch`'s neural style [example](https://pytorch.org/tutorials/advanced/neural_style_tutorial.html), we show how to setup a pipeline for doing style transfer on video. The pipeline has following steps:\n",
- "1. Split a video into images\n",
- "2. Run neural style on each image using one of the provided models (from `pytorch` pretrained models for this example).\n",
- "3. Stitch the image back into a video."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Prerequisites\n",
- "Make sure you go through the configuration Notebook located at https://github.com/Azure/MachineLearningNotebooks first if you haven't. This sets you up with a working config file that has information on your workspace, subscription id, etc. "
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Initialize Workspace\n",
- "\n",
- "Initialize a workspace object from persisted configuration."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import os\n",
- "from azureml.core import Workspace, Run, Experiment\n",
- "\n",
- "ws = Workspace.from_config()\n",
- "print('Workspace name: ' + ws.name, \n",
- " 'Azure region: ' + ws.location, \n",
- " 'Subscription id: ' + ws.subscription_id, \n",
- " 'Resource group: ' + ws.resource_group, sep = '\\n')\n",
- "\n",
- "scripts_folder = \"scripts_folder\"\n",
- "\n",
- "if not os.path.isdir(scripts_folder):\n",
- " os.mkdir(scripts_folder)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.compute import AmlCompute, ComputeTarget\n",
- "from azureml.core.datastore import Datastore\n",
- "from azureml.data.data_reference import DataReference\n",
- "from azureml.pipeline.core import Pipeline, PipelineData\n",
- "from azureml.pipeline.steps import PythonScriptStep, MpiStep\n",
- "from azureml.core.runconfig import CondaDependencies, RunConfiguration"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Create or use existing compute"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# AmlCompute\n",
- "cpu_cluster_name = \"cpucluster\"\n",
- "try:\n",
- " cpu_cluster = AmlCompute(ws, cpu_cluster_name)\n",
- " print(\"found existing cluster.\")\n",
- "except:\n",
- " print(\"creating new cluster\")\n",
- " provisioning_config = AmlCompute.provisioning_configuration(vm_size = \"STANDARD_D2_v2\",\n",
- " max_nodes = 1)\n",
- "\n",
- " # create the cluster\n",
- " cpu_cluster = ComputeTarget.create(ws, cpu_cluster_name, provisioning_config)\n",
- " cpu_cluster.wait_for_completion(show_output=True)\n",
- " \n",
- "# AmlCompute\n",
- "gpu_cluster_name = \"gpucluster\"\n",
- "try:\n",
- " gpu_cluster = AmlCompute(ws, gpu_cluster_name)\n",
- " print(\"found existing cluster.\")\n",
- "except:\n",
- " print(\"creating new cluster\")\n",
- " provisioning_config = AmlCompute.provisioning_configuration(vm_size = \"STANDARD_NC6\",\n",
- " max_nodes = 3)\n",
- "\n",
- " # create the cluster\n",
- " gpu_cluster = ComputeTarget.create(ws, gpu_cluster_name, provisioning_config)\n",
- " gpu_cluster.wait_for_completion(show_output=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Python Scripts\n",
- "We use an edited version of `neural_style_mpi.py` (original is [here](https://github.com/pytorch/examples/blob/master/fast_neural_style/neural_style/neural_style_mpi.py)). Scripts to split and stitch the video are thin wrappers to calls to `ffmpeg`. \n",
- "\n",
- "We install `ffmpeg` through conda dependencies."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import shutil\n",
- "shutil.copy(\"neural_style_mpi.py\", scripts_folder)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%writefile $scripts_folder/process_video.py\n",
- "import argparse\n",
- "import glob\n",
- "import os\n",
- "import subprocess\n",
- "\n",
- "parser = argparse.ArgumentParser(description=\"Process input video\")\n",
- "parser.add_argument('--input_video', required=True)\n",
- "parser.add_argument('--output_audio', required=True)\n",
- "parser.add_argument('--output_images', required=True)\n",
- "\n",
- "args = parser.parse_args()\n",
- "\n",
- "os.makedirs(args.output_audio, exist_ok=True)\n",
- "os.makedirs(args.output_images, exist_ok=True)\n",
- "\n",
- "subprocess.run(\"ffmpeg -i {} {}/video.aac\"\n",
- " .format(args.input_video, args.output_audio),\n",
- " shell=True, check=True\n",
- " )\n",
- "\n",
- "subprocess.run(\"ffmpeg -i {} {}/%05d_video.jpg -hide_banner\"\n",
- " .format(args.input_video, args.output_images),\n",
- " shell=True, check=True\n",
- " )"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%writefile $scripts_folder/stitch_video.py\n",
- "import argparse\n",
- "import os\n",
- "import subprocess\n",
- "\n",
- "parser = argparse.ArgumentParser(description=\"Process input video\")\n",
- "parser.add_argument('--images_dir', required=True)\n",
- "parser.add_argument('--input_audio', required=True)\n",
- "parser.add_argument('--output_dir', required=True)\n",
- "\n",
- "args = parser.parse_args()\n",
- "\n",
- "os.makedirs(args.output_dir, exist_ok=True)\n",
- "\n",
- "subprocess.run(\"ffmpeg -framerate 30 -i {}/%05d_video.jpg -c:v libx264 -profile:v high -crf 20 -pix_fmt yuv420p \"\n",
- " \"-y {}/video_without_audio.mp4\"\n",
- " .format(args.images_dir, args.output_dir),\n",
- " shell=True, check=True\n",
- " )\n",
- "\n",
- "subprocess.run(\"ffmpeg -i {}/video_without_audio.mp4 -i {}/video.aac -map 0:0 -map 1:0 -vcodec \"\n",
- " \"copy -acodec copy -y {}/video_with_audio.mp4\"\n",
- " .format(args.output_dir, args.input_audio, args.output_dir),\n",
- " shell=True, check=True\n",
- " )"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# datastore for input video\n",
- "account_name = \"happypathspublic\"\n",
- "video_ds = Datastore.register_azure_blob_container(ws, \"videos\", \"videos\",\n",
- " account_name=account_name, overwrite=True)\n",
- "\n",
- "# datastore for models\n",
- "models_ds = Datastore.register_azure_blob_container(ws, \"models\", \"styletransfer\", \n",
- " account_name=\"pipelinedata\", \n",
- " overwrite=True)\n",
- " \n",
- "# downloaded models from https://pytorch.org/tutorials/advanced/neural_style_tutorial.html are kept here\n",
- "models_dir = DataReference(data_reference_name=\"models\", datastore=models_ds, \n",
- " path_on_datastore=\"saved_models\", mode=\"download\")\n",
- "\n",
- "# the default blob store attached to a workspace\n",
- "default_datastore = ws.get_default_datastore()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Sample video"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "orangutan_video = DataReference(datastore=video_ds,\n",
- " data_reference_name=\"video\",\n",
- " path_on_datastore=\"orangutan.mp4\", mode=\"download\")"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "cd = CondaDependencies()\n",
- "\n",
- "cd.add_channel(\"conda-forge\")\n",
- "cd.add_conda_package(\"ffmpeg\")\n",
- "\n",
- "cd.add_channel(\"pytorch\")\n",
- "cd.add_conda_package(\"pytorch\")\n",
- "cd.add_conda_package(\"torchvision\")\n",
- "\n",
- "# Runconfig\n",
- "amlcompute_run_config = RunConfiguration(conda_dependencies=cd)\n",
- "amlcompute_run_config.environment.docker.enabled = True\n",
- "amlcompute_run_config.environment.docker.gpu_support = True\n",
- "amlcompute_run_config.environment.docker.base_image = \"pytorch/pytorch\"\n",
- "amlcompute_run_config.environment.spark.precache_packages = False"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "ffmpeg_audio = PipelineData(name=\"ffmpeg_audio\", datastore=default_datastore)\n",
- "ffmpeg_images = PipelineData(name=\"ffmpeg_images\", datastore=default_datastore)\n",
- "processed_images = PipelineData(name=\"processed_images\", datastore=default_datastore)\n",
- "output_video = PipelineData(name=\"output_video\", datastore=default_datastore)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Define tweakable parameters to pipeline\n",
- "These parameters can be changed when the pipeline is published and rerun from a REST call"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.pipeline.core.graph import PipelineParameter\n",
- "# create a parameter for style (one of \"candy\", \"mosaic\", \"rain_princess\", \"udnie\") to transfer the images to\n",
- "style_param = PipelineParameter(name=\"style\", default_value=\"mosaic\")\n",
- "# create a parameter for the number of nodes to use in step no. 2 (style transfer)\n",
- "nodecount_param = PipelineParameter(name=\"nodecount\", default_value=1)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "split_video_step = PythonScriptStep(\n",
- " name=\"split video\",\n",
- " script_name=\"process_video.py\",\n",
- " arguments=[\"--input_video\", orangutan_video,\n",
- " \"--output_audio\", ffmpeg_audio,\n",
- " \"--output_images\", ffmpeg_images,\n",
- " ],\n",
- " compute_target=cpu_cluster,\n",
- " inputs=[orangutan_video],\n",
- " outputs=[ffmpeg_images, ffmpeg_audio],\n",
- " runconfig=amlcompute_run_config,\n",
- " source_directory=scripts_folder\n",
- ")\n",
- "\n",
- "# create a MPI step for distributing style transfer step across multiple nodes in AmlCompute \n",
- "# using 'nodecount_param' PipelineParameter\n",
- "distributed_style_transfer_step = MpiStep(\n",
- " name=\"mpi style transfer\",\n",
- " script_name=\"neural_style_mpi.py\",\n",
- " arguments=[\"--content-dir\", ffmpeg_images,\n",
- " \"--output-dir\", processed_images,\n",
- " \"--model-dir\", models_dir,\n",
- " \"--style\", style_param,\n",
- " \"--cuda\", 1\n",
- " ],\n",
- " compute_target=gpu_cluster,\n",
- " node_count=nodecount_param, \n",
- " process_count_per_node=1,\n",
- " inputs=[models_dir, ffmpeg_images],\n",
- " outputs=[processed_images],\n",
- " pip_packages=[\"mpi4py\", \"torch\", \"torchvision\"],\n",
- " runconfig=amlcompute_run_config,\n",
- " use_gpu=True,\n",
- " source_directory=scripts_folder\n",
- ")\n",
- "\n",
- "stitch_video_step = PythonScriptStep(\n",
- " name=\"stitch\",\n",
- " script_name=\"stitch_video.py\",\n",
- " arguments=[\"--images_dir\", processed_images, \n",
- " \"--input_audio\", ffmpeg_audio, \n",
- " \"--output_dir\", output_video],\n",
- " compute_target=cpu_cluster,\n",
- " inputs=[processed_images, ffmpeg_audio],\n",
- " outputs=[output_video],\n",
- " runconfig=amlcompute_run_config,\n",
- " source_directory=scripts_folder\n",
- ")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Run the pipeline"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "pipeline = Pipeline(workspace=ws, steps=[stitch_video_step])\n",
- "# submit the pipeline and provide values for the PipelineParameters used in the pipeline\n",
- "pipeline_run = Experiment(ws, 'style_transfer').submit(pipeline, pipeline_params={\"style\": \"mosaic\", \"nodecount\": 3})"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Monitor using widget"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.widgets import RunDetails\n",
- "RunDetails(pipeline_run).show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Downloads the video in `output_video` folder"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Download output video"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "def download_video(run, target_dir=None):\n",
- " stitch_run = run.find_step_run(\"stitch\")[0]\n",
- " port_data = stitch_run.get_output_data(\"output_video\")\n",
- " port_data.download(target_dir, show_progress=True)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "pipeline_run.wait_for_completion()\n",
- "download_video(pipeline_run, \"output_video_mosaic\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Publish pipeline"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "published_pipeline = pipeline_run.publish_pipeline(\n",
- " name=\"batch score style transfer\", description=\"style transfer\", version=\"1.0\")\n",
- "\n",
- "published_id = published_pipeline.id"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Re-run pipeline through REST calls for other styles"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Get AAD token"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.authentication import AzureCliAuthentication\n",
- "import requests\n",
- "\n",
- "cli_auth = AzureCliAuthentication()\n",
- "aad_token = cli_auth.get_authentication_header()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Get endpoint URL"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "rest_endpoint = published_pipeline.endpoint"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Send request and monitor"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# run the pipeline using PipelineParameter values style='candy' and nodecount=2\n",
- "response = requests.post(rest_endpoint, \n",
- " headers=aad_token,\n",
- " json={\"ExperimentName\": \"style_transfer\",\n",
- " \"ParameterAssignments\": {\"style\": \"candy\", \"nodecount\": 2}}) \n",
- "run_id = response.json()[\"Id\"]\n",
- "\n",
- "from azureml.pipeline.core.run import PipelineRun\n",
- "published_pipeline_run_candy = PipelineRun(ws.experiments[\"style_transfer\"], run_id)\n",
- "\n",
- "RunDetails(published_pipeline_run_candy).show()"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# run the pipeline using PipelineParameter values style='rain_princess' and nodecount=3\n",
- "response = requests.post(rest_endpoint, \n",
- " headers=aad_token,\n",
- " json={\"ExperimentName\": \"style_transfer\",\n",
- " \"ParameterAssignments\": {\"style\": \"rain_princess\", \"nodecount\": 3}}) \n",
- "run_id = response.json()[\"Id\"]\n",
- "\n",
- "from azureml.pipeline.core.run import PipelineRun\n",
- "published_pipeline_run_rain = PipelineRun(ws.experiments[\"style_transfer\"], run_id)\n",
- "\n",
- "RunDetails(published_pipeline_run_rain).show()"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# run the pipeline using PipelineParameter values style='udnie' and nodecount=4\n",
- "response = requests.post(rest_endpoint, \n",
- " headers=aad_token,\n",
- " json={\"ExperimentName\": \"style_transfer\",\n",
- " \"ParameterAssignments\": {\"style\": \"udnie\", \"nodecount\": 4}}) \n",
- "run_id = response.json()[\"Id\"]\n",
- "\n",
- "from azureml.pipeline.core.run import PipelineRun\n",
- "published_pipeline_run_udnie = PipelineRun(ws.experiments[\"style_transfer\"], run_id)\n",
- "\n",
- "RunDetails(published_pipeline_run_udnie).show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Download output from re-run"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "published_pipeline_run_candy.wait_for_completion()\n",
- "published_pipeline_run_rain.wait_for_completion()\n",
- "published_pipeline_run_udnie.wait_for_completion()"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "download_video(published_pipeline_run_candy, target_dir=\"output_video_candy\")\n",
- "download_video(published_pipeline_run_rain, target_dir=\"output_video_rain_princess\")\n",
- "download_video(published_pipeline_run_udnie, target_dir=\"output_video_udnie\")"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "hichando"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Neural style transfer on video\n",
+ "Using modified code from `pytorch`'s neural style [example](https://pytorch.org/tutorials/advanced/neural_style_tutorial.html), we show how to setup a pipeline for doing style transfer on video. The pipeline has following steps:\n",
+ "1. Split a video into images\n",
+ "2. Run neural style on each image using one of the provided models (from `pytorch` pretrained models for this example).\n",
+ "3. Stitch the image back into a video."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Prerequisites\n",
+ "Make sure you go through the configuration Notebook located at https://github.com/Azure/MachineLearningNotebooks first if you haven't. This sets you up with a working config file that has information on your workspace, subscription id, etc. "
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Initialize Workspace\n",
+ "\n",
+ "Initialize a workspace object from persisted configuration."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import os\n",
+ "from azureml.core import Workspace, Run, Experiment\n",
+ "\n",
+ "ws = Workspace.from_config()\n",
+ "print('Workspace name: ' + ws.name, \n",
+ " 'Azure region: ' + ws.location, \n",
+ " 'Subscription id: ' + ws.subscription_id, \n",
+ " 'Resource group: ' + ws.resource_group, sep = '\\n')\n",
+ "\n",
+ "scripts_folder = \"scripts_folder\"\n",
+ "\n",
+ "if not os.path.isdir(scripts_folder):\n",
+ " os.mkdir(scripts_folder)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.compute import AmlCompute, ComputeTarget\n",
+ "from azureml.core.datastore import Datastore\n",
+ "from azureml.data.data_reference import DataReference\n",
+ "from azureml.pipeline.core import Pipeline, PipelineData\n",
+ "from azureml.pipeline.steps import PythonScriptStep, MpiStep\n",
+ "from azureml.core.runconfig import CondaDependencies, RunConfiguration"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Create or use existing compute"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# AmlCompute\n",
+ "cpu_cluster_name = \"cpucluster\"\n",
+ "try:\n",
+ " cpu_cluster = AmlCompute(ws, cpu_cluster_name)\n",
+ " print(\"found existing cluster.\")\n",
+ "except:\n",
+ " print(\"creating new cluster\")\n",
+ " provisioning_config = AmlCompute.provisioning_configuration(vm_size = \"STANDARD_D2_v2\",\n",
+ " max_nodes = 1)\n",
+ "\n",
+ " # create the cluster\n",
+ " cpu_cluster = ComputeTarget.create(ws, cpu_cluster_name, provisioning_config)\n",
+ " cpu_cluster.wait_for_completion(show_output=True)\n",
+ " \n",
+ "# AmlCompute\n",
+ "gpu_cluster_name = \"gpucluster\"\n",
+ "try:\n",
+ " gpu_cluster = AmlCompute(ws, gpu_cluster_name)\n",
+ " print(\"found existing cluster.\")\n",
+ "except:\n",
+ " print(\"creating new cluster\")\n",
+ " provisioning_config = AmlCompute.provisioning_configuration(vm_size = \"STANDARD_NC6\",\n",
+ " max_nodes = 3)\n",
+ "\n",
+ " # create the cluster\n",
+ " gpu_cluster = ComputeTarget.create(ws, gpu_cluster_name, provisioning_config)\n",
+ " gpu_cluster.wait_for_completion(show_output=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Python Scripts\n",
+ "We use an edited version of `neural_style_mpi.py` (original is [here](https://github.com/pytorch/examples/blob/master/fast_neural_style/neural_style/neural_style_mpi.py)). Scripts to split and stitch the video are thin wrappers to calls to `ffmpeg`. \n",
+ "\n",
+ "We install `ffmpeg` through conda dependencies."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import shutil\n",
+ "shutil.copy(\"neural_style_mpi.py\", scripts_folder)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%writefile $scripts_folder/process_video.py\n",
+ "import argparse\n",
+ "import glob\n",
+ "import os\n",
+ "import subprocess\n",
+ "\n",
+ "parser = argparse.ArgumentParser(description=\"Process input video\")\n",
+ "parser.add_argument('--input_video', required=True)\n",
+ "parser.add_argument('--output_audio', required=True)\n",
+ "parser.add_argument('--output_images', required=True)\n",
+ "\n",
+ "args = parser.parse_args()\n",
+ "\n",
+ "os.makedirs(args.output_audio, exist_ok=True)\n",
+ "os.makedirs(args.output_images, exist_ok=True)\n",
+ "\n",
+ "subprocess.run(\"ffmpeg -i {} {}/video.aac\"\n",
+ " .format(args.input_video, args.output_audio),\n",
+ " shell=True, check=True\n",
+ " )\n",
+ "\n",
+ "subprocess.run(\"ffmpeg -i {} {}/%05d_video.jpg -hide_banner\"\n",
+ " .format(args.input_video, args.output_images),\n",
+ " shell=True, check=True\n",
+ " )"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%writefile $scripts_folder/stitch_video.py\n",
+ "import argparse\n",
+ "import os\n",
+ "import subprocess\n",
+ "\n",
+ "parser = argparse.ArgumentParser(description=\"Process input video\")\n",
+ "parser.add_argument('--images_dir', required=True)\n",
+ "parser.add_argument('--input_audio', required=True)\n",
+ "parser.add_argument('--output_dir', required=True)\n",
+ "\n",
+ "args = parser.parse_args()\n",
+ "\n",
+ "os.makedirs(args.output_dir, exist_ok=True)\n",
+ "\n",
+ "subprocess.run(\"ffmpeg -framerate 30 -i {}/%05d_video.jpg -c:v libx264 -profile:v high -crf 20 -pix_fmt yuv420p \"\n",
+ " \"-y {}/video_without_audio.mp4\"\n",
+ " .format(args.images_dir, args.output_dir),\n",
+ " shell=True, check=True\n",
+ " )\n",
+ "\n",
+ "subprocess.run(\"ffmpeg -i {}/video_without_audio.mp4 -i {}/video.aac -map 0:0 -map 1:0 -vcodec \"\n",
+ " \"copy -acodec copy -y {}/video_with_audio.mp4\"\n",
+ " .format(args.output_dir, args.input_audio, args.output_dir),\n",
+ " shell=True, check=True\n",
+ " )"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# datastore for input video\n",
+ "account_name = \"happypathspublic\"\n",
+ "video_ds = Datastore.register_azure_blob_container(ws, \"videos\", \"videos\",\n",
+ " account_name=account_name, overwrite=True)\n",
+ "\n",
+ "# datastore for models\n",
+ "models_ds = Datastore.register_azure_blob_container(ws, \"models\", \"styletransfer\", \n",
+ " account_name=\"pipelinedata\", \n",
+ " overwrite=True)\n",
+ " \n",
+ "# downloaded models from https://pytorch.org/tutorials/advanced/neural_style_tutorial.html are kept here\n",
+ "models_dir = DataReference(data_reference_name=\"models\", datastore=models_ds, \n",
+ " path_on_datastore=\"saved_models\", mode=\"download\")\n",
+ "\n",
+ "# the default blob store attached to a workspace\n",
+ "default_datastore = ws.get_default_datastore()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Sample video"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "orangutan_video = DataReference(datastore=video_ds,\n",
+ " data_reference_name=\"video\",\n",
+ " path_on_datastore=\"orangutan.mp4\", mode=\"download\")"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "cd = CondaDependencies()\n",
+ "\n",
+ "cd.add_channel(\"conda-forge\")\n",
+ "cd.add_conda_package(\"ffmpeg\")\n",
+ "\n",
+ "cd.add_channel(\"pytorch\")\n",
+ "cd.add_conda_package(\"pytorch\")\n",
+ "cd.add_conda_package(\"torchvision\")\n",
+ "\n",
+ "# Runconfig\n",
+ "amlcompute_run_config = RunConfiguration(conda_dependencies=cd)\n",
+ "amlcompute_run_config.environment.docker.enabled = True\n",
+ "amlcompute_run_config.environment.docker.gpu_support = True\n",
+ "amlcompute_run_config.environment.docker.base_image = \"pytorch/pytorch\"\n",
+ "amlcompute_run_config.environment.spark.precache_packages = False"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "ffmpeg_audio = PipelineData(name=\"ffmpeg_audio\", datastore=default_datastore)\n",
+ "ffmpeg_images = PipelineData(name=\"ffmpeg_images\", datastore=default_datastore)\n",
+ "processed_images = PipelineData(name=\"processed_images\", datastore=default_datastore)\n",
+ "output_video = PipelineData(name=\"output_video\", datastore=default_datastore)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Define tweakable parameters to pipeline\n",
+ "These parameters can be changed when the pipeline is published and rerun from a REST call"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.pipeline.core.graph import PipelineParameter\n",
+ "# create a parameter for style (one of \"candy\", \"mosaic\", \"rain_princess\", \"udnie\") to transfer the images to\n",
+ "style_param = PipelineParameter(name=\"style\", default_value=\"mosaic\")\n",
+ "# create a parameter for the number of nodes to use in step no. 2 (style transfer)\n",
+ "nodecount_param = PipelineParameter(name=\"nodecount\", default_value=1)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "split_video_step = PythonScriptStep(\n",
+ " name=\"split video\",\n",
+ " script_name=\"process_video.py\",\n",
+ " arguments=[\"--input_video\", orangutan_video,\n",
+ " \"--output_audio\", ffmpeg_audio,\n",
+ " \"--output_images\", ffmpeg_images,\n",
+ " ],\n",
+ " compute_target=cpu_cluster,\n",
+ " inputs=[orangutan_video],\n",
+ " outputs=[ffmpeg_images, ffmpeg_audio],\n",
+ " runconfig=amlcompute_run_config,\n",
+ " source_directory=scripts_folder\n",
+ ")\n",
+ "\n",
+ "# create a MPI step for distributing style transfer step across multiple nodes in AmlCompute \n",
+ "# using 'nodecount_param' PipelineParameter\n",
+ "distributed_style_transfer_step = MpiStep(\n",
+ " name=\"mpi style transfer\",\n",
+ " script_name=\"neural_style_mpi.py\",\n",
+ " arguments=[\"--content-dir\", ffmpeg_images,\n",
+ " \"--output-dir\", processed_images,\n",
+ " \"--model-dir\", models_dir,\n",
+ " \"--style\", style_param,\n",
+ " \"--cuda\", 1\n",
+ " ],\n",
+ " compute_target=gpu_cluster,\n",
+ " node_count=nodecount_param, \n",
+ " process_count_per_node=1,\n",
+ " inputs=[models_dir, ffmpeg_images],\n",
+ " outputs=[processed_images],\n",
+ " pip_packages=[\"mpi4py\", \"torch\", \"torchvision\"],\n",
+ " runconfig=amlcompute_run_config,\n",
+ " use_gpu=True,\n",
+ " source_directory=scripts_folder\n",
+ ")\n",
+ "\n",
+ "stitch_video_step = PythonScriptStep(\n",
+ " name=\"stitch\",\n",
+ " script_name=\"stitch_video.py\",\n",
+ " arguments=[\"--images_dir\", processed_images, \n",
+ " \"--input_audio\", ffmpeg_audio, \n",
+ " \"--output_dir\", output_video],\n",
+ " compute_target=cpu_cluster,\n",
+ " inputs=[processed_images, ffmpeg_audio],\n",
+ " outputs=[output_video],\n",
+ " runconfig=amlcompute_run_config,\n",
+ " source_directory=scripts_folder\n",
+ ")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Run the pipeline"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "pipeline = Pipeline(workspace=ws, steps=[stitch_video_step])\n",
+ "# submit the pipeline and provide values for the PipelineParameters used in the pipeline\n",
+ "pipeline_run = Experiment(ws, 'style_transfer').submit(pipeline, pipeline_params={\"style\": \"mosaic\", \"nodecount\": 3})"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Monitor using widget"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.widgets import RunDetails\n",
+ "RunDetails(pipeline_run).show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Downloads the video in `output_video` folder"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Download output video"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "def download_video(run, target_dir=None):\n",
+ " stitch_run = run.find_step_run(\"stitch\")[0]\n",
+ " port_data = stitch_run.get_output_data(\"output_video\")\n",
+ " port_data.download(target_dir, show_progress=True)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "pipeline_run.wait_for_completion()\n",
+ "download_video(pipeline_run, \"output_video_mosaic\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Publish pipeline"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "published_pipeline = pipeline_run.publish_pipeline(\n",
+ " name=\"batch score style transfer\", description=\"style transfer\", version=\"1.0\")\n",
+ "\n",
+ "published_id = published_pipeline.id"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Re-run pipeline through REST calls for other styles"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Get AAD token"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.authentication import AzureCliAuthentication\n",
+ "import requests\n",
+ "\n",
+ "cli_auth = AzureCliAuthentication()\n",
+ "aad_token = cli_auth.get_authentication_header()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Get endpoint URL"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "rest_endpoint = published_pipeline.endpoint"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Send request and monitor"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# run the pipeline using PipelineParameter values style='candy' and nodecount=2\n",
+ "response = requests.post(rest_endpoint, \n",
+ " headers=aad_token,\n",
+ " json={\"ExperimentName\": \"style_transfer\",\n",
+ " \"ParameterAssignments\": {\"style\": \"candy\", \"nodecount\": 2}}) \n",
+ "run_id = response.json()[\"Id\"]\n",
+ "\n",
+ "from azureml.pipeline.core.run import PipelineRun\n",
+ "published_pipeline_run_candy = PipelineRun(ws.experiments[\"style_transfer\"], run_id)\n",
+ "\n",
+ "RunDetails(published_pipeline_run_candy).show()"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# run the pipeline using PipelineParameter values style='rain_princess' and nodecount=3\n",
+ "response = requests.post(rest_endpoint, \n",
+ " headers=aad_token,\n",
+ " json={\"ExperimentName\": \"style_transfer\",\n",
+ " \"ParameterAssignments\": {\"style\": \"rain_princess\", \"nodecount\": 3}}) \n",
+ "run_id = response.json()[\"Id\"]\n",
+ "\n",
+ "from azureml.pipeline.core.run import PipelineRun\n",
+ "published_pipeline_run_rain = PipelineRun(ws.experiments[\"style_transfer\"], run_id)\n",
+ "\n",
+ "RunDetails(published_pipeline_run_rain).show()"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# run the pipeline using PipelineParameter values style='udnie' and nodecount=4\n",
+ "response = requests.post(rest_endpoint, \n",
+ " headers=aad_token,\n",
+ " json={\"ExperimentName\": \"style_transfer\",\n",
+ " \"ParameterAssignments\": {\"style\": \"udnie\", \"nodecount\": 4}}) \n",
+ "run_id = response.json()[\"Id\"]\n",
+ "\n",
+ "from azureml.pipeline.core.run import PipelineRun\n",
+ "published_pipeline_run_udnie = PipelineRun(ws.experiments[\"style_transfer\"], run_id)\n",
+ "\n",
+ "RunDetails(published_pipeline_run_udnie).show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Download output from re-run"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "published_pipeline_run_candy.wait_for_completion()\n",
+ "published_pipeline_run_rain.wait_for_completion()\n",
+ "published_pipeline_run_udnie.wait_for_completion()"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "download_video(published_pipeline_run_candy, target_dir=\"output_video_candy\")\n",
+ "download_video(published_pipeline_run_rain, target_dir=\"output_video_rain_princess\")\n",
+ "download_video(published_pipeline_run_udnie, target_dir=\"output_video_udnie\")"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3",
- "language": "python",
- "name": "python3"
+ "metadata": {
+ "authors": [
+ {
+ "name": "hichando"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.7"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.7"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/training-with-deep-learning/distributed-cntk-with-custom-docker/distributed-cntk-with-custom-docker.ipynb b/how-to-use-azureml/training-with-deep-learning/distributed-cntk-with-custom-docker/distributed-cntk-with-custom-docker.ipynb
index f70b19d0..1d8eb645 100644
--- a/how-to-use-azureml/training-with-deep-learning/distributed-cntk-with-custom-docker/distributed-cntk-with-custom-docker.ipynb
+++ b/how-to-use-azureml/training-with-deep-learning/distributed-cntk-with-custom-docker/distributed-cntk-with-custom-docker.ipynb
@@ -1,394 +1,394 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Distributed CNTK using custom docker images\n",
- "In this tutorial, you will train a CNTK model on the [MNIST](http://yann.lecun.com/exdb/mnist/) dataset using a custom docker image and distributed training."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Prerequisites\n",
- "* Understand the [architecture and terms](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture) introduced by Azure Machine Learning\n",
- "* Go through the [00.configuration.ipynb]() notebook to:\n",
- " * install the AML SDK\n",
- " * create a workspace and its configuration file (`config.json`)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Check core SDK version number\n",
- "import azureml.core\n",
- "\n",
- "print(\"SDK version:\", azureml.core.VERSION)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Diagnostics\n",
- "Opt-in diagnostics for better experience, quality, and security of future releases."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "Diagnostics"
- ]
- },
- "outputs": [],
- "source": [
- "from azureml.telemetry import set_diagnostics_collection\n",
- "\n",
- "set_diagnostics_collection(send_diagnostics=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Initialize workspace\n",
- "\n",
- "Initialize a [Workspace](https://review.docs.microsoft.com/en-us/azure/machine-learning/service/concept-azure-machine-learning-architecture?branch=release-ignite-aml#workspace) object from the existing workspace you created in the Prerequisites step. `Workspace.from_config()` creates a workspace object from the details stored in `config.json`."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.workspace import Workspace\n",
- "\n",
- "ws = Workspace.from_config()\n",
- "print('Workspace name: ' + ws.name, \n",
- " 'Azure region: ' + ws.location, \n",
- " 'Subscription id: ' + ws.subscription_id, \n",
- " 'Resource group: ' + ws.resource_group, sep = '\\n')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Create or Attach existing AmlCompute\n",
- "You will need to create a [compute target](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#compute-target) for training your model. In this tutorial, you create `AmlCompute` as your training compute resource.\n",
- "\n",
- "**Creation of AmlCompute takes approximately 5 minutes.** If the AmlCompute with that name is already in your workspace this code will skip the creation process.\n",
- "\n",
- "As with other Azure services, there are limits on certain resources (e.g. AmlCompute) associated with the Azure Machine Learning service. Please read [this article](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-manage-quotas) on the default limits and how to request more quota."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.compute import ComputeTarget, AmlCompute\n",
- "from azureml.core.compute_target import ComputeTargetException\n",
- "\n",
- "# choose a name for your cluster\n",
- "cluster_name = \"gpucluster\"\n",
- "\n",
- "try:\n",
- " compute_target = ComputeTarget(workspace=ws, name=cluster_name)\n",
- " print('Found existing compute target.')\n",
- "except ComputeTargetException:\n",
- " print('Creating a new compute target...')\n",
- " compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_NC6',\n",
- " max_nodes=4)\n",
- "\n",
- " # create the cluster\n",
- " compute_target = ComputeTarget.create(ws, cluster_name, compute_config)\n",
- "\n",
- " compute_target.wait_for_completion(show_output=True)\n",
- "\n",
- "# Use the 'status' property to get a detailed status for the current AmlCompute. \n",
- "print(compute_target.status.serialize())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Upload training data\n",
- "For this tutorial, we will be using the MNIST dataset.\n",
- "\n",
- "First, let's download the dataset. We've included the `install_mnist.py` script to download the data and convert it to a CNTK-supported format. Our data files will get written to a directory named `'mnist'`."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import install_mnist\n",
- "\n",
- "install_mnist.main('mnist')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "To make the data accessible for remote training, you will need to upload the data from your local machine to the cloud. AML provides a convenient way to do so via a [Datastore](https://docs.microsoft.com/azure/machine-learning/service/how-to-access-data). The datastore provides a mechanism for you to upload/download data, and interact with it from your remote compute targets. \n",
- "\n",
- "Each workspace is associated with a default datastore. In this tutorial, we will upload the training data to this default datastore, which we will then mount on the remote compute for training in the next section."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "ds = ws.get_default_datastore()\n",
- "print(ds.datastore_type, ds.account_name, ds.container_name)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "The following code will upload the training data to the path `./mnist` on the default datastore."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "ds.upload(src_dir='./mnist', target_path='./mnist')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Now let's get a reference to the path on the datastore with the training data. We can do so using the `path` method. In the next section, we can then pass this reference to our training script's `--data_dir` argument. "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "path_on_datastore = 'mnist'\n",
- "ds_data = ds.path(path_on_datastore)\n",
- "print(ds_data)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Train model on the remote compute\n",
- "Now that we have the cluster ready to go, let's run our distributed training job."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create a project directory\n",
- "Create a directory that will contain all the necessary code from your local machine that you will need access to on the remote resource. This includes the training script, and any additional files your training script depends on."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import os\n",
- "\n",
- "project_folder = './cntk-distr'\n",
- "os.makedirs(project_folder, exist_ok=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copy the training script `cntk_distr_mnist.py` into this project directory."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import shutil\n",
- "\n",
- "shutil.copy('cntk_distr_mnist.py', project_folder)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create an experiment\n",
- "Create an [experiment](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#experiment) to track all the runs in your workspace for this distributed CNTK tutorial. "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core import Experiment\n",
- "\n",
- "experiment_name = 'cntk-distr'\n",
- "experiment = Experiment(ws, name=experiment_name)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create an Estimator\n",
- "The AML SDK's base Estimator enables you to easily submit custom scripts for both single-node and distributed runs. You should this generic estimator for training code using frameworks such as sklearn or CNTK that don't have corresponding custom estimators. For more information on using the generic estimator, refer [here](https://docs.microsoft.com/azure/machine-learning/service/how-to-train-ml-models)."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.train.estimator import *\n",
- "\n",
- "script_params = {\n",
- " '--num_epochs': 20,\n",
- " '--data_dir': ds_data.as_mount(),\n",
- " '--output_dir': './outputs'\n",
- "}\n",
- "\n",
- "estimator = Estimator(source_directory=project_folder,\n",
- " compute_target=compute_target,\n",
- " entry_script='cntk_distr_mnist.py',\n",
- " script_params=script_params,\n",
- " node_count=2,\n",
- " process_count_per_node=1,\n",
- " distributed_backend='mpi', \n",
- " pip_packages=['cntk-gpu==2.6'],\n",
- " custom_docker_base_image='microsoft/mmlspark:gpu-0.12',\n",
- " use_gpu=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "We would like to train our model using a [pre-built Docker container](https://hub.docker.com/r/microsoft/mmlspark/). To do so, specify the name of the docker image to the argument `custom_docker_base_image`. You can only provide images available in public docker repositories such as Docker Hub using this argument. To use an image from a private docker repository, use the constructor's `environment_definition` parameter instead. Finally, we provide the `cntk` package to `pip_packages` to install CNTK 2.6 on our custom image.\n",
- "\n",
- "The above code specifies that we will run our training script on `2` nodes, with one worker per node. In order to run distributed CNTK, which uses MPI, you must provide the argument `distributed_backend='mpi'`."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Submit job\n",
- "Run your experiment by submitting your estimator object. Note that this call is asynchronous."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run = experiment.submit(estimator)\n",
- "print(run)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Monitor your run\n",
- "You can monitor the progress of the run with a Jupyter widget. Like the run submission, the widget is asynchronous and provides live updates every 10-15 seconds until the job completes."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.widgets import RunDetails\n",
- "\n",
- "RunDetails(run).show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Alternatively, you can block until the script has completed training before running more code."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run.wait_for_completion(show_output=True)"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "minxia"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Distributed CNTK using custom docker images\n",
+ "In this tutorial, you will train a CNTK model on the [MNIST](http://yann.lecun.com/exdb/mnist/) dataset using a custom docker image and distributed training."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Prerequisites\n",
+ "* Understand the [architecture and terms](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture) introduced by Azure Machine Learning\n",
+ "* Go through the [configuration notebook](../../../configuration.ipynb) to:\n",
+ " * install the AML SDK\n",
+ " * create a workspace and its configuration file (`config.json`)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Check core SDK version number\n",
+ "import azureml.core\n",
+ "\n",
+ "print(\"SDK version:\", azureml.core.VERSION)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Diagnostics\n",
+ "Opt-in diagnostics for better experience, quality, and security of future releases."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "Diagnostics"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "from azureml.telemetry import set_diagnostics_collection\n",
+ "\n",
+ "set_diagnostics_collection(send_diagnostics=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Initialize workspace\n",
+ "\n",
+ "Initialize a [Workspace](https://review.docs.microsoft.com/en-us/azure/machine-learning/service/concept-azure-machine-learning-architecture?branch=release-ignite-aml#workspace) object from the existing workspace you created in the Prerequisites step. `Workspace.from_config()` creates a workspace object from the details stored in `config.json`."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.workspace import Workspace\n",
+ "\n",
+ "ws = Workspace.from_config()\n",
+ "print('Workspace name: ' + ws.name, \n",
+ " 'Azure region: ' + ws.location, \n",
+ " 'Subscription id: ' + ws.subscription_id, \n",
+ " 'Resource group: ' + ws.resource_group, sep = '\\n')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Create or Attach existing AmlCompute\n",
+ "You will need to create a [compute target](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#compute-target) for training your model. In this tutorial, you create `AmlCompute` as your training compute resource.\n",
+ "\n",
+ "**Creation of AmlCompute takes approximately 5 minutes.** If the AmlCompute with that name is already in your workspace this code will skip the creation process.\n",
+ "\n",
+ "As with other Azure services, there are limits on certain resources (e.g. AmlCompute) associated with the Azure Machine Learning service. Please read [this article](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-manage-quotas) on the default limits and how to request more quota."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.compute import ComputeTarget, AmlCompute\n",
+ "from azureml.core.compute_target import ComputeTargetException\n",
+ "\n",
+ "# choose a name for your cluster\n",
+ "cluster_name = \"gpucluster\"\n",
+ "\n",
+ "try:\n",
+ " compute_target = ComputeTarget(workspace=ws, name=cluster_name)\n",
+ " print('Found existing compute target.')\n",
+ "except ComputeTargetException:\n",
+ " print('Creating a new compute target...')\n",
+ " compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_NC6',\n",
+ " max_nodes=4)\n",
+ "\n",
+ " # create the cluster\n",
+ " compute_target = ComputeTarget.create(ws, cluster_name, compute_config)\n",
+ "\n",
+ " compute_target.wait_for_completion(show_output=True)\n",
+ "\n",
+ "# use get_status() to get a detailed status for the current AmlCompute. \n",
+ "print(compute_target.get_status().serialize())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Upload training data\n",
+ "For this tutorial, we will be using the MNIST dataset.\n",
+ "\n",
+ "First, let's download the dataset. We've included the `install_mnist.py` script to download the data and convert it to a CNTK-supported format. Our data files will get written to a directory named `'mnist'`."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import install_mnist\n",
+ "\n",
+ "install_mnist.main('mnist')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "To make the data accessible for remote training, you will need to upload the data from your local machine to the cloud. AML provides a convenient way to do so via a [Datastore](https://docs.microsoft.com/azure/machine-learning/service/how-to-access-data). The datastore provides a mechanism for you to upload/download data, and interact with it from your remote compute targets. \n",
+ "\n",
+ "Each workspace is associated with a default datastore. In this tutorial, we will upload the training data to this default datastore, which we will then mount on the remote compute for training in the next section."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "ds = ws.get_default_datastore()\n",
+ "print(ds.datastore_type, ds.account_name, ds.container_name)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "The following code will upload the training data to the path `./mnist` on the default datastore."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "ds.upload(src_dir='./mnist', target_path='./mnist')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Now let's get a reference to the path on the datastore with the training data. We can do so using the `path` method. In the next section, we can then pass this reference to our training script's `--data_dir` argument. "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "path_on_datastore = 'mnist'\n",
+ "ds_data = ds.path(path_on_datastore)\n",
+ "print(ds_data)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Train model on the remote compute\n",
+ "Now that we have the cluster ready to go, let's run our distributed training job."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create a project directory\n",
+ "Create a directory that will contain all the necessary code from your local machine that you will need access to on the remote resource. This includes the training script, and any additional files your training script depends on."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import os\n",
+ "\n",
+ "project_folder = './cntk-distr'\n",
+ "os.makedirs(project_folder, exist_ok=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copy the training script `cntk_distr_mnist.py` into this project directory."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import shutil\n",
+ "\n",
+ "shutil.copy('cntk_distr_mnist.py', project_folder)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create an experiment\n",
+ "Create an [experiment](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#experiment) to track all the runs in your workspace for this distributed CNTK tutorial. "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core import Experiment\n",
+ "\n",
+ "experiment_name = 'cntk-distr'\n",
+ "experiment = Experiment(ws, name=experiment_name)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create an Estimator\n",
+ "The AML SDK's base Estimator enables you to easily submit custom scripts for both single-node and distributed runs. You should this generic estimator for training code using frameworks such as sklearn or CNTK that don't have corresponding custom estimators. For more information on using the generic estimator, refer [here](https://docs.microsoft.com/azure/machine-learning/service/how-to-train-ml-models)."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.train.estimator import *\n",
+ "\n",
+ "script_params = {\n",
+ " '--num_epochs': 20,\n",
+ " '--data_dir': ds_data.as_mount(),\n",
+ " '--output_dir': './outputs'\n",
+ "}\n",
+ "\n",
+ "estimator = Estimator(source_directory=project_folder,\n",
+ " compute_target=compute_target,\n",
+ " entry_script='cntk_distr_mnist.py',\n",
+ " script_params=script_params,\n",
+ " node_count=2,\n",
+ " process_count_per_node=1,\n",
+ " distributed_backend='mpi', \n",
+ " pip_packages=['cntk-gpu==2.6'],\n",
+ " custom_docker_base_image='microsoft/mmlspark:gpu-0.12',\n",
+ " use_gpu=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "We would like to train our model using a [pre-built Docker container](https://hub.docker.com/r/microsoft/mmlspark/). To do so, specify the name of the docker image to the argument `custom_docker_base_image`. You can only provide images available in public docker repositories such as Docker Hub using this argument. To use an image from a private docker repository, use the constructor's `environment_definition` parameter instead. Finally, we provide the `cntk` package to `pip_packages` to install CNTK 2.6 on our custom image.\n",
+ "\n",
+ "The above code specifies that we will run our training script on `2` nodes, with one worker per node. In order to run distributed CNTK, which uses MPI, you must provide the argument `distributed_backend='mpi'`."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Submit job\n",
+ "Run your experiment by submitting your estimator object. Note that this call is asynchronous."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run = experiment.submit(estimator)\n",
+ "print(run)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Monitor your run\n",
+ "You can monitor the progress of the run with a Jupyter widget. Like the run submission, the widget is asynchronous and provides live updates every 10-15 seconds until the job completes."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.widgets import RunDetails\n",
+ "\n",
+ "RunDetails(run).show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Alternatively, you can block until the script has completed training before running more code."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run.wait_for_completion(show_output=True)"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "minxia"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.6"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/training-with-deep-learning/distributed-pytorch-with-horovod/distributed-pytorch-with-horovod.ipynb b/how-to-use-azureml/training-with-deep-learning/distributed-pytorch-with-horovod/distributed-pytorch-with-horovod.ipynb
index 53122acc..5581ff44 100644
--- a/how-to-use-azureml/training-with-deep-learning/distributed-pytorch-with-horovod/distributed-pytorch-with-horovod.ipynb
+++ b/how-to-use-azureml/training-with-deep-learning/distributed-pytorch-with-horovod/distributed-pytorch-with-horovod.ipynb
@@ -1,335 +1,335 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Distributed PyTorch with Horovod\n",
- "In this tutorial, you will train a PyTorch model on the [MNIST](http://yann.lecun.com/exdb/mnist/) dataset using distributed training via [Horovod](https://github.com/uber/horovod) across a GPU cluster."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Prerequisites\n",
- "* Go through the [Configuration](https://github.com/Azure/MachineLearningNotebooks/blob/master/configuration.ipynb) notebook to install the Azure Machine Learning Python SDK and create an Azure ML `Workspace`\n",
- "* Review the [tutorial](https://github.com/Azure/MachineLearningNotebooks/blob/master/how-to-use-azureml/training-with-deep-learning/train-hyperparameter-tune-deploy-with-pytorch/train-hyperparameter-tune-deploy-with-pytorch.ipynb) on single-node PyTorch training using Azure Machine Learning"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Check core SDK version number\n",
- "import azureml.core\n",
- "\n",
- "print(\"SDK version:\", azureml.core.VERSION)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Diagnostics\n",
- "Opt-in diagnostics for better experience, quality, and security of future releases."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "Diagnostics"
- ]
- },
- "outputs": [],
- "source": [
- "from azureml.telemetry import set_diagnostics_collection\n",
- "\n",
- "set_diagnostics_collection(send_diagnostics=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Initialize workspace\n",
- "\n",
- "Initialize a [Workspace](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#workspace) object from the existing workspace you created in the Prerequisites step. `Workspace.from_config()` creates a workspace object from the details stored in `config.json`."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.workspace import Workspace\n",
- "\n",
- "ws = Workspace.from_config()\n",
- "print('Workspace name: ' + ws.name, \n",
- " 'Azure region: ' + ws.location, \n",
- " 'Subscription id: ' + ws.subscription_id, \n",
- " 'Resource group: ' + ws.resource_group, sep = '\\n')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Create or attach existing AmlCompute\n",
- "You will need to create a [compute target](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#compute-target) for training your model. In this tutorial, we use Azure ML managed compute ([AmlCompute](https://docs.microsoft.com/azure/machine-learning/service/how-to-set-up-training-targets#amlcompute)) for our remote training compute resource. Specifically, the below code creates an `STANDARD_NC6` GPU cluster that autoscales from `0` to `4` nodes.\n",
- "\n",
- "**Creation of AmlCompute takes approximately 5 minutes.** If the AmlCompute with that name is already in your workspace, this code will skip the creation process.\n",
- "\n",
- "As with other Azure services, there are limits on certain resources (e.g. AmlCompute) associated with the Azure Machine Learning service. Please read [this article](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-manage-quotas) on the default limits and how to request more quota."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.compute import ComputeTarget, AmlCompute\n",
- "from azureml.core.compute_target import ComputeTargetException\n",
- "\n",
- "# choose a name for your cluster\n",
- "cluster_name = \"gpucluster\"\n",
- "\n",
- "try:\n",
- " compute_target = ComputeTarget(workspace=ws, name=cluster_name)\n",
- " print('Found existing compute target.')\n",
- "except ComputeTargetException:\n",
- " print('Creating a new compute target...')\n",
- " compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_NC6',\n",
- " max_nodes=4)\n",
- "\n",
- " # create the cluster\n",
- " compute_target = ComputeTarget.create(ws, cluster_name, compute_config)\n",
- "\n",
- " compute_target.wait_for_completion(show_output=True)\n",
- "\n",
- "# Use the 'status' property to get a detailed status for the current AmlCompute. \n",
- "print(compute_target.status.serialize())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "The above code creates GPU compute. If you instead want to create CPU compute, provide a different VM size to the `vm_size` parameter, such as `STANDARD_D2_V2`."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Train model on the remote compute\n",
- "Now that we have the AmlCompute ready to go, let's run our distributed training job."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create a project directory\n",
- "Create a directory that will contain all the necessary code from your local machine that you will need access to on the remote resource. This includes the training script and any additional files your training script depends on."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import os\n",
- "\n",
- "project_folder = './pytorch-distr-hvd'\n",
- "os.makedirs(project_folder, exist_ok=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Prepare training script\n",
- "Now you will need to create your training script. In this tutorial, the script for distributed training of MNIST is already provided for you at `pytorch_horovod_mnist.py`. In practice, you should be able to take any custom PyTorch training script as is and run it with Azure ML without having to modify your code.\n",
- "\n",
- "However, if you would like to use Azure ML's [metric logging](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#logging) capabilities, you will have to add a small amount of Azure ML logic inside your training script. In this example, at each logging interval, we will log the loss for that minibatch to our Azure ML run.\n",
- "\n",
- "To do so, in `pytorch_horovod_mnist.py`, we will first access the Azure ML `Run` object within the script:\n",
- "```Python\n",
- "from azureml.core.run import Run\n",
- "run = Run.get_context()\n",
- "```\n",
- "Later within the script, we log the loss metric to our run:\n",
- "```Python\n",
- "run.log('loss', loss.item())\n",
- "```"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Once your script is ready, copy the training script `pytorch_horovod_mnist.py` into the project directory."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import shutil\n",
- "\n",
- "shutil.copy('pytorch_horovod_mnist.py', project_folder)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create an experiment\n",
- "Create an [Experiment](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#experiment) to track all the runs in your workspace for this distributed PyTorch tutorial. "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core import Experiment\n",
- "\n",
- "experiment_name = 'pytorch-distr-hvd'\n",
- "experiment = Experiment(ws, name=experiment_name)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create a PyTorch estimator\n",
- "The Azure ML SDK's PyTorch estimator enables you to easily submit PyTorch training jobs for both single-node and distributed runs. For more information on the PyTorch estimator, refer [here](https://docs.microsoft.com/azure/machine-learning/service/how-to-train-pytorch)."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.train.dnn import PyTorch\n",
- "\n",
- "estimator = PyTorch(source_directory=project_folder,\n",
- " compute_target=compute_target,\n",
- " entry_script='pytorch_horovod_mnist.py',\n",
- " node_count=2,\n",
- " process_count_per_node=1,\n",
- " distributed_backend='mpi',\n",
- " use_gpu=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "The above code specifies that we will run our training script on `2` nodes, with one worker per node. In order to execute a distributed run using MPI/Horovod, you must provide the argument `distributed_backend='mpi'`. Using this estimator with these settings, PyTorch, Horovod and their dependencies will be installed for you. However, if your script also uses other packages, make sure to install them via the `PyTorch` constructor's `pip_packages` or `conda_packages` parameters."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Submit job\n",
- "Run your experiment by submitting your estimator object. Note that this call is asynchronous."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run = experiment.submit(estimator)\n",
- "print(run)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Monitor your run\n",
- "You can monitor the progress of the run with a Jupyter widget. Like the run submission, the widget is asynchronous and provides live updates every 10-15 seconds until the job completes. You can see that the widget automatically plots and visualizes the loss metric that we logged to the Azure ML run."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.widgets import RunDetails\n",
- "\n",
- "RunDetails(run).show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Alternatively, you can block until the script has completed training before running more code."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run.wait_for_completion(show_output=True) # this provides a verbose log"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "minxia"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Distributed PyTorch with Horovod\n",
+ "In this tutorial, you will train a PyTorch model on the [MNIST](http://yann.lecun.com/exdb/mnist/) dataset using distributed training via [Horovod](https://github.com/uber/horovod) across a GPU cluster."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Prerequisites\n",
+ "* Go through the [Configuration](../../../configuration.ipynb) notebook to install the Azure Machine Learning Python SDK and create an Azure ML `Workspace`\n",
+ "* Review the [tutorial](https://github.com/Azure/MachineLearningNotebooks/blob/master/how-to-use-azureml/training-with-deep-learning/train-hyperparameter-tune-deploy-with-pytorch/train-hyperparameter-tune-deploy-with-pytorch.ipynb) on single-node PyTorch training using Azure Machine Learning"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Check core SDK version number\n",
+ "import azureml.core\n",
+ "\n",
+ "print(\"SDK version:\", azureml.core.VERSION)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Diagnostics\n",
+ "Opt-in diagnostics for better experience, quality, and security of future releases."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "Diagnostics"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "from azureml.telemetry import set_diagnostics_collection\n",
+ "\n",
+ "set_diagnostics_collection(send_diagnostics=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Initialize workspace\n",
+ "\n",
+ "Initialize a [Workspace](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#workspace) object from the existing workspace you created in the Prerequisites step. `Workspace.from_config()` creates a workspace object from the details stored in `config.json`."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.workspace import Workspace\n",
+ "\n",
+ "ws = Workspace.from_config()\n",
+ "print('Workspace name: ' + ws.name, \n",
+ " 'Azure region: ' + ws.location, \n",
+ " 'Subscription id: ' + ws.subscription_id, \n",
+ " 'Resource group: ' + ws.resource_group, sep = '\\n')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Create or attach existing AmlCompute\n",
+ "You will need to create a [compute target](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#compute-target) for training your model. In this tutorial, we use Azure ML managed compute ([AmlCompute](https://docs.microsoft.com/azure/machine-learning/service/how-to-set-up-training-targets#amlcompute)) for our remote training compute resource. Specifically, the below code creates an `STANDARD_NC6` GPU cluster that autoscales from `0` to `4` nodes.\n",
+ "\n",
+ "**Creation of AmlCompute takes approximately 5 minutes.** If the AmlCompute with that name is already in your workspace, this code will skip the creation process.\n",
+ "\n",
+ "As with other Azure services, there are limits on certain resources (e.g. AmlCompute) associated with the Azure Machine Learning service. Please read [this article](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-manage-quotas) on the default limits and how to request more quota."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.compute import ComputeTarget, AmlCompute\n",
+ "from azureml.core.compute_target import ComputeTargetException\n",
+ "\n",
+ "# choose a name for your cluster\n",
+ "cluster_name = \"gpucluster\"\n",
+ "\n",
+ "try:\n",
+ " compute_target = ComputeTarget(workspace=ws, name=cluster_name)\n",
+ " print('Found existing compute target.')\n",
+ "except ComputeTargetException:\n",
+ " print('Creating a new compute target...')\n",
+ " compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_NC6',\n",
+ " max_nodes=4)\n",
+ "\n",
+ " # create the cluster\n",
+ " compute_target = ComputeTarget.create(ws, cluster_name, compute_config)\n",
+ "\n",
+ " compute_target.wait_for_completion(show_output=True)\n",
+ "\n",
+ "# use get_status() to get a detailed status for the current AmlCompute. \n",
+ "print(compute_target.get_status().serialize())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "The above code creates GPU compute. If you instead want to create CPU compute, provide a different VM size to the `vm_size` parameter, such as `STANDARD_D2_V2`."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Train model on the remote compute\n",
+ "Now that we have the AmlCompute ready to go, let's run our distributed training job."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create a project directory\n",
+ "Create a directory that will contain all the necessary code from your local machine that you will need access to on the remote resource. This includes the training script and any additional files your training script depends on."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import os\n",
+ "\n",
+ "project_folder = './pytorch-distr-hvd'\n",
+ "os.makedirs(project_folder, exist_ok=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Prepare training script\n",
+ "Now you will need to create your training script. In this tutorial, the script for distributed training of MNIST is already provided for you at `pytorch_horovod_mnist.py`. In practice, you should be able to take any custom PyTorch training script as is and run it with Azure ML without having to modify your code.\n",
+ "\n",
+ "However, if you would like to use Azure ML's [metric logging](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#logging) capabilities, you will have to add a small amount of Azure ML logic inside your training script. In this example, at each logging interval, we will log the loss for that minibatch to our Azure ML run.\n",
+ "\n",
+ "To do so, in `pytorch_horovod_mnist.py`, we will first access the Azure ML `Run` object within the script:\n",
+ "```Python\n",
+ "from azureml.core.run import Run\n",
+ "run = Run.get_context()\n",
+ "```\n",
+ "Later within the script, we log the loss metric to our run:\n",
+ "```Python\n",
+ "run.log('loss', loss.item())\n",
+ "```"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Once your script is ready, copy the training script `pytorch_horovod_mnist.py` into the project directory."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import shutil\n",
+ "\n",
+ "shutil.copy('pytorch_horovod_mnist.py', project_folder)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create an experiment\n",
+ "Create an [Experiment](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#experiment) to track all the runs in your workspace for this distributed PyTorch tutorial. "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core import Experiment\n",
+ "\n",
+ "experiment_name = 'pytorch-distr-hvd'\n",
+ "experiment = Experiment(ws, name=experiment_name)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create a PyTorch estimator\n",
+ "The Azure ML SDK's PyTorch estimator enables you to easily submit PyTorch training jobs for both single-node and distributed runs. For more information on the PyTorch estimator, refer [here](https://docs.microsoft.com/azure/machine-learning/service/how-to-train-pytorch)."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.train.dnn import PyTorch\n",
+ "\n",
+ "estimator = PyTorch(source_directory=project_folder,\n",
+ " compute_target=compute_target,\n",
+ " entry_script='pytorch_horovod_mnist.py',\n",
+ " node_count=2,\n",
+ " process_count_per_node=1,\n",
+ " distributed_backend='mpi',\n",
+ " use_gpu=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "The above code specifies that we will run our training script on `2` nodes, with one worker per node. In order to execute a distributed run using MPI/Horovod, you must provide the argument `distributed_backend='mpi'`. Using this estimator with these settings, PyTorch, Horovod and their dependencies will be installed for you. However, if your script also uses other packages, make sure to install them via the `PyTorch` constructor's `pip_packages` or `conda_packages` parameters."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Submit job\n",
+ "Run your experiment by submitting your estimator object. Note that this call is asynchronous."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run = experiment.submit(estimator)\n",
+ "print(run)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Monitor your run\n",
+ "You can monitor the progress of the run with a Jupyter widget. Like the run submission, the widget is asynchronous and provides live updates every 10-15 seconds until the job completes. You can see that the widget automatically plots and visualizes the loss metric that we logged to the Azure ML run."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.widgets import RunDetails\n",
+ "\n",
+ "RunDetails(run).show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Alternatively, you can block until the script has completed training before running more code."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run.wait_for_completion(show_output=True) # this provides a verbose log"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "minxia"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.6"
+ },
+ "msauthor": "minxia"
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- },
- "msauthor": "minxia"
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/training-with-deep-learning/distributed-tensorflow-with-horovod/distributed-tensorflow-with-horovod.ipynb b/how-to-use-azureml/training-with-deep-learning/distributed-tensorflow-with-horovod/distributed-tensorflow-with-horovod.ipynb
index ebdd57d7..8ba590f8 100644
--- a/how-to-use-azureml/training-with-deep-learning/distributed-tensorflow-with-horovod/distributed-tensorflow-with-horovod.ipynb
+++ b/how-to-use-azureml/training-with-deep-learning/distributed-tensorflow-with-horovod/distributed-tensorflow-with-horovod.ipynb
@@ -1,404 +1,404 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Distributed Tensorflow with Horovod\n",
- "In this tutorial, you will train a word2vec model in TensorFlow using distributed training via [Horovod](https://github.com/uber/horovod)."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Prerequisites\n",
- "* Understand the [architecture and terms](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture) introduced by Azure Machine Learning (AML)\n",
- "* Go through the [00.configuration.ipynb](https://github.com/Azure/MachineLearningNotebooks/blob/master/00.configuration.ipynb) notebook to:\n",
- " * install the AML SDK\n",
- " * create a workspace and its configuration file (`config.json`)\n",
- "* Review the [tutorial](https://aka.ms/aml-notebook-hyperdrive) on single-node TensorFlow training using the SDK"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Check core SDK version number\n",
- "import azureml.core\n",
- "\n",
- "print(\"SDK version:\", azureml.core.VERSION)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Diagnostics\n",
- "Opt-in diagnostics for better experience, quality, and security of future releases."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "Diagnostics"
- ]
- },
- "outputs": [],
- "source": [
- "from azureml.telemetry import set_diagnostics_collection\n",
- "\n",
- "set_diagnostics_collection(send_diagnostics=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Initialize workspace\n",
- "Initialize a [Workspace](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#workspace) object from the existing workspace you created in the Prerequisites step. `Workspace.from_config()` creates a workspace object from the details stored in `config.json`."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.workspace import Workspace\n",
- "\n",
- "ws = Workspace.from_config()\n",
- "print('Workspace name: ' + ws.name, \n",
- " 'Azure region: ' + ws.location, \n",
- " 'Subscription id: ' + ws.subscription_id, \n",
- " 'Resource group: ' + ws.resource_group, sep = '\\n')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Create or Attach existing AmlCompute\n",
- "You will need to create a [compute target](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#compute-target) for training your model. In this tutorial, you create `AmlCompute` as your training compute resource.\n",
- "\n",
- "**Creation of AmlCompute takes approximately 5 minutes.** If the AmlCompute with that name is already in your workspace this code will skip the creation process.\n",
- "\n",
- "As with other Azure services, there are limits on certain resources (e.g. AmlCompute) associated with the Azure Machine Learning service. Please read [this article](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-manage-quotas) on the default limits and how to request more quota."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.compute import ComputeTarget, AmlCompute\n",
- "from azureml.core.compute_target import ComputeTargetException\n",
- "\n",
- "# choose a name for your cluster\n",
- "cluster_name = \"gpucluster\"\n",
- "\n",
- "try:\n",
- " compute_target = ComputeTarget(workspace=ws, name=cluster_name)\n",
- " print('Found existing compute target')\n",
- "except ComputeTargetException:\n",
- " print('Creating a new compute target...')\n",
- " compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_NC6', \n",
- " max_nodes=4)\n",
- "\n",
- " # create the cluster\n",
- " compute_target = ComputeTarget.create(ws, cluster_name, compute_config)\n",
- "\n",
- " compute_target.wait_for_completion(show_output=True)\n",
- "\n",
- "# Use the 'status' property to get a detailed status for the current cluster. \n",
- "print(compute_target.status.serialize())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "The above code creates a GPU cluster. If you instead want to create a CPU cluster, provide a different VM size to the `vm_size` parameter, such as `STANDARD_D2_V2`."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Upload data to datastore\n",
- "To make data accessible for remote training, AML provides a convenient way to do so via a [Datastore](https://docs.microsoft.com/azure/machine-learning/service/how-to-access-data). The datastore provides a mechanism for you to upload/download data to Azure Storage, and interact with it from your remote compute targets. \n",
- "\n",
- "If your data is already stored in Azure, or you download the data as part of your training script, you will not need to do this step. For this tutorial, although you can download the data in your training script, we will demonstrate how to upload the training data to a datastore and access it during training to illustrate the datastore functionality."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "First, download the training data from [here](http://mattmahoney.net/dc/text8.zip) to your local machine:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import os\n",
- "import urllib\n",
- "\n",
- "os.makedirs('./data', exist_ok=True)\n",
- "download_url = 'http://mattmahoney.net/dc/text8.zip'\n",
- "urllib.request.urlretrieve(download_url, filename='./data/text8.zip')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Each workspace is associated with a default datastore. In this tutorial, we will upload the training data to this default datastore."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "ds = ws.get_default_datastore()\n",
- "print(ds.datastore_type, ds.account_name, ds.container_name)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Upload the contents of the data directory to the path `./data` on the default datastore."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "ds.upload(src_dir='data', target_path='data', overwrite=True, show_progress=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "For convenience, let's get a reference to the path on the datastore with the zip file of training data. We can do so using the `path` method. In the next section, we can then pass this reference to our training script's `--input_data` argument. "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "path_on_datastore = 'data/text8.zip'\n",
- "ds_data = ds.path(path_on_datastore)\n",
- "print(ds_data)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Train model on the remote compute"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create a project directory\n",
- "Create a directory that will contain all the necessary code from your local machine that you will need access to on the remote resource. This includes the training script, and any additional files your training script depends on."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import os\n",
- "\n",
- "project_folder = './tf-distr-hvd'\n",
- "os.makedirs(project_folder, exist_ok=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copy the training script `tf_horovod_word2vec.py` into this project directory."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import shutil\n",
- "\n",
- "shutil.copy('tf_horovod_word2vec.py', project_folder)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create an experiment\n",
- "Create an [Experiment](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#experiment) to track all the runs in your workspace for this distributed TensorFlow tutorial. "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core import Experiment\n",
- "\n",
- "experiment_name = 'tf-distr-hvd'\n",
- "experiment = Experiment(ws, name=experiment_name)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create a TensorFlow estimator\n",
- "The AML SDK's TensorFlow estimator enables you to easily submit TensorFlow training jobs for both single-node and distributed runs. For more information on the TensorFlow estimator, refer [here](https://docs.microsoft.com/azure/machine-learning/service/how-to-train-tensorflow)."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.train.dnn import TensorFlow\n",
- "\n",
- "script_params={\n",
- " '--input_data': ds_data\n",
- "}\n",
- "\n",
- "estimator= TensorFlow(source_directory=project_folder,\n",
- " compute_target=compute_target,\n",
- " script_params=script_params,\n",
- " entry_script='tf_horovod_word2vec.py',\n",
- " node_count=2,\n",
- " process_count_per_node=1,\n",
- " distributed_backend='mpi',\n",
- " use_gpu=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "The above code specifies that we will run our training script on `2` nodes, with one worker per node. In order to execute a distributed run using MPI/Horovod, you must provide the argument `distributed_backend='mpi'`. Using this estimator with these settings, TensorFlow, Horovod and their dependencies will be installed for you. However, if your script also uses other packages, make sure to install them via the `TensorFlow` constructor's `pip_packages` or `conda_packages` parameters.\n",
- "\n",
- "Note that we passed our training data reference `ds_data` to our script's `--input_data` argument. This will 1) mount our datastore on the remote compute and 2) provide the path to the data zip file on our datastore."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Submit job\n",
- "Run your experiment by submitting your estimator object. Note that this call is asynchronous."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run = experiment.submit(estimator)\n",
- "print(run)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Monitor your run\n",
- "You can monitor the progress of the run with a Jupyter widget. Like the run submission, the widget is asynchronous and provides live updates every 10-15 seconds until the job completes."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.widgets import RunDetails\n",
- "RunDetails(run).show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Alternatively, you can block until the script has completed training before running more code."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run.wait_for_completion(show_output=True)"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "roastala"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Distributed Tensorflow with Horovod\n",
+ "In this tutorial, you will train a word2vec model in TensorFlow using distributed training via [Horovod](https://github.com/uber/horovod)."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Prerequisites\n",
+ "* Understand the [architecture and terms](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture) introduced by Azure Machine Learning (AML)\n",
+ "* Go through the [configuration notebook](../../../configuration.ipynb) to:\n",
+ " * install the AML SDK\n",
+ " * create a workspace and its configuration file (`config.json`)\n",
+ "* Review the [tutorial](https://aka.ms/aml-notebook-hyperdrive) on single-node TensorFlow training using the SDK"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Check core SDK version number\n",
+ "import azureml.core\n",
+ "\n",
+ "print(\"SDK version:\", azureml.core.VERSION)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Diagnostics\n",
+ "Opt-in diagnostics for better experience, quality, and security of future releases."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "Diagnostics"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "from azureml.telemetry import set_diagnostics_collection\n",
+ "\n",
+ "set_diagnostics_collection(send_diagnostics=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Initialize workspace\n",
+ "Initialize a [Workspace](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#workspace) object from the existing workspace you created in the Prerequisites step. `Workspace.from_config()` creates a workspace object from the details stored in `config.json`."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.workspace import Workspace\n",
+ "\n",
+ "ws = Workspace.from_config()\n",
+ "print('Workspace name: ' + ws.name, \n",
+ " 'Azure region: ' + ws.location, \n",
+ " 'Subscription id: ' + ws.subscription_id, \n",
+ " 'Resource group: ' + ws.resource_group, sep = '\\n')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Create or Attach existing AmlCompute\n",
+ "You will need to create a [compute target](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#compute-target) for training your model. In this tutorial, you create `AmlCompute` as your training compute resource.\n",
+ "\n",
+ "**Creation of AmlCompute takes approximately 5 minutes.** If the AmlCompute with that name is already in your workspace this code will skip the creation process.\n",
+ "\n",
+ "As with other Azure services, there are limits on certain resources (e.g. AmlCompute) associated with the Azure Machine Learning service. Please read [this article](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-manage-quotas) on the default limits and how to request more quota."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.compute import ComputeTarget, AmlCompute\n",
+ "from azureml.core.compute_target import ComputeTargetException\n",
+ "\n",
+ "# choose a name for your cluster\n",
+ "cluster_name = \"gpucluster\"\n",
+ "\n",
+ "try:\n",
+ " compute_target = ComputeTarget(workspace=ws, name=cluster_name)\n",
+ " print('Found existing compute target')\n",
+ "except ComputeTargetException:\n",
+ " print('Creating a new compute target...')\n",
+ " compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_NC6', \n",
+ " max_nodes=4)\n",
+ "\n",
+ " # create the cluster\n",
+ " compute_target = ComputeTarget.create(ws, cluster_name, compute_config)\n",
+ "\n",
+ " compute_target.wait_for_completion(show_output=True)\n",
+ "\n",
+ "# use get_status() to get a detailed status for the current cluster. \n",
+ "print(compute_target.get_status().serialize())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "The above code creates a GPU cluster. If you instead want to create a CPU cluster, provide a different VM size to the `vm_size` parameter, such as `STANDARD_D2_V2`."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Upload data to datastore\n",
+ "To make data accessible for remote training, AML provides a convenient way to do so via a [Datastore](https://docs.microsoft.com/azure/machine-learning/service/how-to-access-data). The datastore provides a mechanism for you to upload/download data to Azure Storage, and interact with it from your remote compute targets. \n",
+ "\n",
+ "If your data is already stored in Azure, or you download the data as part of your training script, you will not need to do this step. For this tutorial, although you can download the data in your training script, we will demonstrate how to upload the training data to a datastore and access it during training to illustrate the datastore functionality."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "First, download the training data from [here](http://mattmahoney.net/dc/text8.zip) to your local machine:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import os\n",
+ "import urllib\n",
+ "\n",
+ "os.makedirs('./data', exist_ok=True)\n",
+ "download_url = 'http://mattmahoney.net/dc/text8.zip'\n",
+ "urllib.request.urlretrieve(download_url, filename='./data/text8.zip')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Each workspace is associated with a default datastore. In this tutorial, we will upload the training data to this default datastore."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "ds = ws.get_default_datastore()\n",
+ "print(ds.datastore_type, ds.account_name, ds.container_name)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Upload the contents of the data directory to the path `./data` on the default datastore."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "ds.upload(src_dir='data', target_path='data', overwrite=True, show_progress=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "For convenience, let's get a reference to the path on the datastore with the zip file of training data. We can do so using the `path` method. In the next section, we can then pass this reference to our training script's `--input_data` argument. "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "path_on_datastore = 'data/text8.zip'\n",
+ "ds_data = ds.path(path_on_datastore)\n",
+ "print(ds_data)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Train model on the remote compute"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create a project directory\n",
+ "Create a directory that will contain all the necessary code from your local machine that you will need access to on the remote resource. This includes the training script, and any additional files your training script depends on."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import os\n",
+ "\n",
+ "project_folder = './tf-distr-hvd'\n",
+ "os.makedirs(project_folder, exist_ok=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copy the training script `tf_horovod_word2vec.py` into this project directory."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import shutil\n",
+ "\n",
+ "shutil.copy('tf_horovod_word2vec.py', project_folder)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create an experiment\n",
+ "Create an [Experiment](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#experiment) to track all the runs in your workspace for this distributed TensorFlow tutorial. "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core import Experiment\n",
+ "\n",
+ "experiment_name = 'tf-distr-hvd'\n",
+ "experiment = Experiment(ws, name=experiment_name)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create a TensorFlow estimator\n",
+ "The AML SDK's TensorFlow estimator enables you to easily submit TensorFlow training jobs for both single-node and distributed runs. For more information on the TensorFlow estimator, refer [here](https://docs.microsoft.com/azure/machine-learning/service/how-to-train-tensorflow)."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.train.dnn import TensorFlow\n",
+ "\n",
+ "script_params={\n",
+ " '--input_data': ds_data\n",
+ "}\n",
+ "\n",
+ "estimator= TensorFlow(source_directory=project_folder,\n",
+ " compute_target=compute_target,\n",
+ " script_params=script_params,\n",
+ " entry_script='tf_horovod_word2vec.py',\n",
+ " node_count=2,\n",
+ " process_count_per_node=1,\n",
+ " distributed_backend='mpi',\n",
+ " use_gpu=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "The above code specifies that we will run our training script on `2` nodes, with one worker per node. In order to execute a distributed run using MPI/Horovod, you must provide the argument `distributed_backend='mpi'`. Using this estimator with these settings, TensorFlow, Horovod and their dependencies will be installed for you. However, if your script also uses other packages, make sure to install them via the `TensorFlow` constructor's `pip_packages` or `conda_packages` parameters.\n",
+ "\n",
+ "Note that we passed our training data reference `ds_data` to our script's `--input_data` argument. This will 1) mount our datastore on the remote compute and 2) provide the path to the data zip file on our datastore."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Submit job\n",
+ "Run your experiment by submitting your estimator object. Note that this call is asynchronous."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run = experiment.submit(estimator)\n",
+ "print(run)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Monitor your run\n",
+ "You can monitor the progress of the run with a Jupyter widget. Like the run submission, the widget is asynchronous and provides live updates every 10-15 seconds until the job completes."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.widgets import RunDetails\n",
+ "RunDetails(run).show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Alternatively, you can block until the script has completed training before running more code."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run.wait_for_completion(show_output=True)"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "roastala"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.6"
+ },
+ "msauthor": "minxia"
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- },
- "msauthor": "minxia"
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/training-with-deep-learning/distributed-tensorflow-with-parameter-server/distributed-tensorflow-with-parameter-server.ipynb b/how-to-use-azureml/training-with-deep-learning/distributed-tensorflow-with-parameter-server/distributed-tensorflow-with-parameter-server.ipynb
index 9765f084..4f9df419 100644
--- a/how-to-use-azureml/training-with-deep-learning/distributed-tensorflow-with-parameter-server/distributed-tensorflow-with-parameter-server.ipynb
+++ b/how-to-use-azureml/training-with-deep-learning/distributed-tensorflow-with-parameter-server/distributed-tensorflow-with-parameter-server.ipynb
@@ -1,317 +1,317 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Distributed TensorFlow with parameter server\n",
- "In this tutorial, you will train a TensorFlow model on the [MNIST](http://yann.lecun.com/exdb/mnist/) dataset using native [distributed TensorFlow](https://www.tensorflow.org/deploy/distributed)."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Prerequisites\n",
- "* Understand the [architecture and terms](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture) introduced by Azure Machine Learning (AML)\n",
- "* Go through the [00.configuration.ipynb](https://github.com/Azure/MachineLearningNotebooks/blob/master/00.configuration.ipynb) notebook to:\n",
- " * install the AML SDK\n",
- " * create a workspace and its configuration file (`config.json`)\n",
- "* Review the [tutorial](https://aka.ms/aml-notebook-hyperdrive) on single-node TensorFlow training using the SDK"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Check core SDK version number\n",
- "import azureml.core\n",
- "\n",
- "print(\"SDK version:\", azureml.core.VERSION)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Diagnostics\n",
- "Opt-in diagnostics for better experience, quality, and security of future releases."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "Diagnostics"
- ]
- },
- "outputs": [],
- "source": [
- "from azureml.telemetry import set_diagnostics_collection\n",
- "\n",
- "set_diagnostics_collection(send_diagnostics=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Initialize workspace\n",
- "Initialize a [Workspace](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#workspace) object from the existing workspace you created in the Prerequisites step. `Workspace.from_config()` creates a workspace object from the details stored in `config.json`."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.workspace import Workspace\n",
- "\n",
- "ws = Workspace.from_config()\n",
- "print('Workspace name: ' + ws.name, \n",
- " 'Azure region: ' + ws.location, \n",
- " 'Subscription id: ' + ws.subscription_id, \n",
- " 'Resource group: ' + ws.resource_group, sep = '\\n')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Create or Attach existing AmlCompute\n",
- "You will need to create a [compute target](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#compute-target) for training your model. In this tutorial, you create `AmlCompute` as your training compute resource.\n",
- "\n",
- "**Creation of AmlCompute takes approximately 5 minutes.** If the AmlCompute with that name is already in your workspace this code will skip the creation process.\n",
- "\n",
- "As with other Azure services, there are limits on certain resources (e.g. AmlCompute) associated with the Azure Machine Learning service. Please read [this article](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-manage-quotas) on the default limits and how to request more quota."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.compute import ComputeTarget, AmlCompute\n",
- "from azureml.core.compute_target import ComputeTargetException\n",
- "\n",
- "# choose a name for your cluster\n",
- "cluster_name = \"gpucluster\"\n",
- "\n",
- "try:\n",
- " compute_target = ComputeTarget(workspace=ws, name=cluster_name)\n",
- " print('Found existing compute target.')\n",
- "except ComputeTargetException:\n",
- " print('Creating a new compute target...')\n",
- " compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_NC6', \n",
- " max_nodes=4)\n",
- "\n",
- " # create the cluster\n",
- " compute_target = ComputeTarget.create(ws, cluster_name, compute_config)\n",
- "\n",
- " compute_target.wait_for_completion(show_output=True)\n",
- "\n",
- "# Use the 'status' property to get a detailed status for the current cluster. \n",
- "print(compute_target.status.serialize())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Train model on the remote compute\n",
- "Now that we have the cluster ready to go, let's run our distributed training job."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create a project directory\n",
- "Create a directory that will contain all the necessary code from your local machine that you will need access to on the remote resource. This includes the training script, and any additional files your training script depends on."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import os\n",
- "\n",
- "project_folder = './tf-distr-ps'\n",
- "os.makedirs(project_folder, exist_ok=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copy the training script `tf_mnist_replica.py` into this project directory."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import shutil\n",
- "\n",
- "shutil.copy('tf_mnist_replica.py', project_folder)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create an experiment\n",
- "Create an [Experiment](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#experiment) to track all the runs in your workspace for this distributed TensorFlow tutorial. "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core import Experiment\n",
- "\n",
- "experiment_name = 'tf-distr-ps'\n",
- "experiment = Experiment(ws, name=experiment_name)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create a TensorFlow estimator\n",
- "The AML SDK's TensorFlow estimator enables you to easily submit TensorFlow training jobs for both single-node and distributed runs. For more information on the TensorFlow estimator, refer [here](https://docs.microsoft.com/azure/machine-learning/service/how-to-train-tensorflow)."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.train.dnn import TensorFlow\n",
- "\n",
- "script_params={\n",
- " '--num_gpus': 1,\n",
- " '--train_steps': 500\n",
- "}\n",
- "\n",
- "estimator = TensorFlow(source_directory=project_folder,\n",
- " compute_target=compute_target,\n",
- " script_params=script_params,\n",
- " entry_script='tf_mnist_replica.py',\n",
- " node_count=2,\n",
- " worker_count=2,\n",
- " parameter_server_count=1, \n",
- " distributed_backend='ps',\n",
- " use_gpu=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "The above code specifies that we will run our training script on `2` nodes, with two workers and one parameter server. In order to execute a native distributed TensorFlow run, you must provide the argument `distributed_backend='ps'`. Using this estimator with these settings, TensorFlow and its dependencies will be installed for you. However, if your script also uses other packages, make sure to install them via the `TensorFlow` constructor's `pip_packages` or `conda_packages` parameters."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Submit job\n",
- "Run your experiment by submitting your estimator object. Note that this call is asynchronous."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run = experiment.submit(estimator)\n",
- "print(run)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Monitor your run\n",
- "You can monitor the progress of the run with a Jupyter widget. Like the run submission, the widget is asynchronous and provides live updates every 10-15 seconds until the job completes."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.widgets import RunDetails\n",
- "\n",
- "RunDetails(run).show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Alternatively, you can block until the script has completed training before running more code."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run.wait_for_completion(show_output=True) # this provides a verbose log"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "minxia"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Distributed TensorFlow with parameter server\n",
+ "In this tutorial, you will train a TensorFlow model on the [MNIST](http://yann.lecun.com/exdb/mnist/) dataset using native [distributed TensorFlow](https://www.tensorflow.org/deploy/distributed)."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Prerequisites\n",
+ "* Understand the [architecture and terms](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture) introduced by Azure Machine Learning (AML)\n",
+ "* Go through the [configuration notebook](../../../configuration.ipynb) to:\n",
+ " * install the AML SDK\n",
+ " * create a workspace and its configuration file (`config.json`)\n",
+ "* Review the [tutorial](https://aka.ms/aml-notebook-hyperdrive) on single-node TensorFlow training using the SDK"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Check core SDK version number\n",
+ "import azureml.core\n",
+ "\n",
+ "print(\"SDK version:\", azureml.core.VERSION)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Diagnostics\n",
+ "Opt-in diagnostics for better experience, quality, and security of future releases."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "Diagnostics"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "from azureml.telemetry import set_diagnostics_collection\n",
+ "\n",
+ "set_diagnostics_collection(send_diagnostics=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Initialize workspace\n",
+ "Initialize a [Workspace](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#workspace) object from the existing workspace you created in the Prerequisites step. `Workspace.from_config()` creates a workspace object from the details stored in `config.json`."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.workspace import Workspace\n",
+ "\n",
+ "ws = Workspace.from_config()\n",
+ "print('Workspace name: ' + ws.name, \n",
+ " 'Azure region: ' + ws.location, \n",
+ " 'Subscription id: ' + ws.subscription_id, \n",
+ " 'Resource group: ' + ws.resource_group, sep = '\\n')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Create or Attach existing AmlCompute\n",
+ "You will need to create a [compute target](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#compute-target) for training your model. In this tutorial, you create `AmlCompute` as your training compute resource.\n",
+ "\n",
+ "**Creation of AmlCompute takes approximately 5 minutes.** If the AmlCompute with that name is already in your workspace this code will skip the creation process.\n",
+ "\n",
+ "As with other Azure services, there are limits on certain resources (e.g. AmlCompute) associated with the Azure Machine Learning service. Please read [this article](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-manage-quotas) on the default limits and how to request more quota."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.compute import ComputeTarget, AmlCompute\n",
+ "from azureml.core.compute_target import ComputeTargetException\n",
+ "\n",
+ "# choose a name for your cluster\n",
+ "cluster_name = \"gpucluster\"\n",
+ "\n",
+ "try:\n",
+ " compute_target = ComputeTarget(workspace=ws, name=cluster_name)\n",
+ " print('Found existing compute target.')\n",
+ "except ComputeTargetException:\n",
+ " print('Creating a new compute target...')\n",
+ " compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_NC6', \n",
+ " max_nodes=4)\n",
+ "\n",
+ " # create the cluster\n",
+ " compute_target = ComputeTarget.create(ws, cluster_name, compute_config)\n",
+ "\n",
+ " compute_target.wait_for_completion(show_output=True)\n",
+ "\n",
+ "# use get_status() to get a detailed status for the current cluster. \n",
+ "print(compute_target.get_status().serialize())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Train model on the remote compute\n",
+ "Now that we have the cluster ready to go, let's run our distributed training job."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create a project directory\n",
+ "Create a directory that will contain all the necessary code from your local machine that you will need access to on the remote resource. This includes the training script, and any additional files your training script depends on."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import os\n",
+ "\n",
+ "project_folder = './tf-distr-ps'\n",
+ "os.makedirs(project_folder, exist_ok=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copy the training script `tf_mnist_replica.py` into this project directory."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import shutil\n",
+ "\n",
+ "shutil.copy('tf_mnist_replica.py', project_folder)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create an experiment\n",
+ "Create an [Experiment](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#experiment) to track all the runs in your workspace for this distributed TensorFlow tutorial. "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core import Experiment\n",
+ "\n",
+ "experiment_name = 'tf-distr-ps'\n",
+ "experiment = Experiment(ws, name=experiment_name)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create a TensorFlow estimator\n",
+ "The AML SDK's TensorFlow estimator enables you to easily submit TensorFlow training jobs for both single-node and distributed runs. For more information on the TensorFlow estimator, refer [here](https://docs.microsoft.com/azure/machine-learning/service/how-to-train-tensorflow)."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.train.dnn import TensorFlow\n",
+ "\n",
+ "script_params={\n",
+ " '--num_gpus': 1,\n",
+ " '--train_steps': 500\n",
+ "}\n",
+ "\n",
+ "estimator = TensorFlow(source_directory=project_folder,\n",
+ " compute_target=compute_target,\n",
+ " script_params=script_params,\n",
+ " entry_script='tf_mnist_replica.py',\n",
+ " node_count=2,\n",
+ " worker_count=2,\n",
+ " parameter_server_count=1, \n",
+ " distributed_backend='ps',\n",
+ " use_gpu=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "The above code specifies that we will run our training script on `2` nodes, with two workers and one parameter server. In order to execute a native distributed TensorFlow run, you must provide the argument `distributed_backend='ps'`. Using this estimator with these settings, TensorFlow and its dependencies will be installed for you. However, if your script also uses other packages, make sure to install them via the `TensorFlow` constructor's `pip_packages` or `conda_packages` parameters."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Submit job\n",
+ "Run your experiment by submitting your estimator object. Note that this call is asynchronous."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run = experiment.submit(estimator)\n",
+ "print(run)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Monitor your run\n",
+ "You can monitor the progress of the run with a Jupyter widget. Like the run submission, the widget is asynchronous and provides live updates every 10-15 seconds until the job completes."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.widgets import RunDetails\n",
+ "\n",
+ "RunDetails(run).show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Alternatively, you can block until the script has completed training before running more code."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run.wait_for_completion(show_output=True) # this provides a verbose log"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "minxia"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.6"
+ },
+ "msauthor": "minxia"
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- },
- "msauthor": "minxia"
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/training-with-deep-learning/export-run-history-to-tensorboard/export-run-history-to-tensorboard.ipynb b/how-to-use-azureml/training-with-deep-learning/export-run-history-to-tensorboard/export-run-history-to-tensorboard.ipynb
index bd005204..7a1cdf1a 100644
--- a/how-to-use-azureml/training-with-deep-learning/export-run-history-to-tensorboard/export-run-history-to-tensorboard.ipynb
+++ b/how-to-use-azureml/training-with-deep-learning/export-run-history-to-tensorboard/export-run-history-to-tensorboard.ipynb
@@ -1,267 +1,267 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Export Run History as Tensorboard logs\n",
- "\n",
- "1. Run some training and log some metrics into Run History\n",
- "2. Export the run history to some directory as Tensorboard logs\n",
- "3. Launch a local Tensorboard to view the run history"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Prerequisites\n",
- "* Understand the [architecture and terms](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture) introduced by Azure Machine Learning\n",
- "* Go through the [00.configuration.ipynb](https://github.com/Azure/MachineLearningNotebooks/blob/master/00.configuration.ipynb) notebook to:\n",
- " * install the AML SDK\n",
- " * create a workspace and its configuration file (`config.json`)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Check core SDK version number\n",
- "import azureml.core\n",
- "\n",
- "print(\"SDK version:\", azureml.core.VERSION)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Install the Azure ML TensorBoard integration package if you haven't already."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "!pip install azureml-contrib-tensorboard"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Initialize Workspace\n",
- "\n",
- "Initialize a workspace object from persisted configuration."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core import Workspace, Run, Experiment\n",
- "\n",
- "\n",
- "ws = Workspace.from_config()\n",
- "print('Workspace name: ' + ws.name, \n",
- " 'Azure region: ' + ws.location, \n",
- " 'Subscription id: ' + ws.subscription_id, \n",
- " 'Resource group: ' + ws.resource_group, sep = '\\n')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Set experiment name and start the run"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "experiment_name = 'export-to-tensorboard'\n",
- "exp = Experiment(ws, experiment_name)\n",
- "root_run = exp.start_logging()"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# load diabetes dataset, a well-known built-in small dataset that comes with scikit-learn\n",
- "from sklearn.datasets import load_diabetes\n",
- "from sklearn.linear_model import Ridge\n",
- "from sklearn.metrics import mean_squared_error\n",
- "from sklearn.model_selection import train_test_split\n",
- "\n",
- "X, y = load_diabetes(return_X_y=True)\n",
- "\n",
- "columns = ['age', 'gender', 'bmi', 'bp', 's1', 's2', 's3', 's4', 's5', 's6']\n",
- "\n",
- "x_train, x_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=0)\n",
- "data = {\n",
- " \"train\":{\"x\":x_train, \"y\":y_train}, \n",
- " \"test\":{\"x\":x_test, \"y\":y_test}\n",
- "}"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Example experiment\n",
- "from tqdm import tqdm\n",
- "\n",
- "alphas = [.1, .2, .3, .4, .5, .6 , .7]\n",
- "\n",
- "# try a bunch of alpha values in a Linear Regression (Ridge) model\n",
- "for alpha in tqdm(alphas):\n",
- " # create a bunch of child runs\n",
- " with root_run.child_run(\"alpha\" + str(alpha)) as run:\n",
- " # More data science stuff\n",
- " reg = Ridge(alpha=alpha)\n",
- " reg.fit(data[\"train\"][\"x\"], data[\"train\"][\"y\"])\n",
- " # TODO save model\n",
- " preds = reg.predict(data[\"test\"][\"x\"])\n",
- " mse = mean_squared_error(preds, data[\"test\"][\"y\"])\n",
- " # End train and eval\n",
- "\n",
- " # log alpha, mean_squared_error and feature names in run history\n",
- " root_run.log(\"alpha\", alpha)\n",
- " root_run.log(\"mse\", mse)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Export Run History to Tensorboard logs"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Export Run History to Tensorboard logs\n",
- "from azureml.contrib.tensorboard.export import export_to_tensorboard\n",
- "import os\n",
- "import tensorflow as tf\n",
- "\n",
- "logdir = 'exportedTBlogs'\n",
- "log_path = os.path.join(os.getcwd(), logdir)\n",
- "try:\n",
- " os.stat(log_path)\n",
- "except os.error:\n",
- " os.mkdir(log_path)\n",
- "print(logdir)\n",
- "\n",
- "# export run history for the project\n",
- "export_to_tensorboard(root_run, logdir)\n",
- "\n",
- "# or export a particular run\n",
- "# export_to_tensorboard(run, logdir)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "root_run.complete()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Start Tensorboard\n",
- "\n",
- "Or you can start the Tensorboard outside this notebook to view the result"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.contrib.tensorboard import Tensorboard\n",
- "\n",
- "# The Tensorboard constructor takes an array of runs, so be sure and pass it in as a single-element array here\n",
- "tb = Tensorboard([], local_root=logdir, port=6006)\n",
- "\n",
- "# If successful, start() returns a string with the URI of the instance.\n",
- "tb.start()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Stop Tensorboard\n",
- "\n",
- "When you're done, make sure to call the `stop()` method of the Tensorboard object."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "tb.stop()"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "roastala"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Export Run History as Tensorboard logs\n",
+ "\n",
+ "1. Run some training and log some metrics into Run History\n",
+ "2. Export the run history to some directory as Tensorboard logs\n",
+ "3. Launch a local Tensorboard to view the run history"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Prerequisites\n",
+ "* Understand the [architecture and terms](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture) introduced by Azure Machine Learning\n",
+ "* Go through the [configuration notebook](../../../configuration.ipynb) notebook to:\n",
+ " * install the AML SDK\n",
+ " * create a workspace and its configuration file (`config.json`)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Check core SDK version number\n",
+ "import azureml.core\n",
+ "\n",
+ "print(\"SDK version:\", azureml.core.VERSION)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Install the Azure ML TensorBoard integration package if you haven't already."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "!pip install azureml-contrib-tensorboard"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Initialize Workspace\n",
+ "\n",
+ "Initialize a workspace object from persisted configuration."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core import Workspace, Run, Experiment\n",
+ "\n",
+ "\n",
+ "ws = Workspace.from_config()\n",
+ "print('Workspace name: ' + ws.name, \n",
+ " 'Azure region: ' + ws.location, \n",
+ " 'Subscription id: ' + ws.subscription_id, \n",
+ " 'Resource group: ' + ws.resource_group, sep = '\\n')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Set experiment name and start the run"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "experiment_name = 'export-to-tensorboard'\n",
+ "exp = Experiment(ws, experiment_name)\n",
+ "root_run = exp.start_logging()"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# load diabetes dataset, a well-known built-in small dataset that comes with scikit-learn\n",
+ "from sklearn.datasets import load_diabetes\n",
+ "from sklearn.linear_model import Ridge\n",
+ "from sklearn.metrics import mean_squared_error\n",
+ "from sklearn.model_selection import train_test_split\n",
+ "\n",
+ "X, y = load_diabetes(return_X_y=True)\n",
+ "\n",
+ "columns = ['age', 'gender', 'bmi', 'bp', 's1', 's2', 's3', 's4', 's5', 's6']\n",
+ "\n",
+ "x_train, x_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=0)\n",
+ "data = {\n",
+ " \"train\":{\"x\":x_train, \"y\":y_train}, \n",
+ " \"test\":{\"x\":x_test, \"y\":y_test}\n",
+ "}"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Example experiment\n",
+ "from tqdm import tqdm\n",
+ "\n",
+ "alphas = [.1, .2, .3, .4, .5, .6 , .7]\n",
+ "\n",
+ "# try a bunch of alpha values in a Linear Regression (Ridge) model\n",
+ "for alpha in tqdm(alphas):\n",
+ " # create a bunch of child runs\n",
+ " with root_run.child_run(\"alpha\" + str(alpha)) as run:\n",
+ " # More data science stuff\n",
+ " reg = Ridge(alpha=alpha)\n",
+ " reg.fit(data[\"train\"][\"x\"], data[\"train\"][\"y\"])\n",
+ " # TODO save model\n",
+ " preds = reg.predict(data[\"test\"][\"x\"])\n",
+ " mse = mean_squared_error(preds, data[\"test\"][\"y\"])\n",
+ " # End train and eval\n",
+ "\n",
+ " # log alpha, mean_squared_error and feature names in run history\n",
+ " root_run.log(\"alpha\", alpha)\n",
+ " root_run.log(\"mse\", mse)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Export Run History to Tensorboard logs"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Export Run History to Tensorboard logs\n",
+ "from azureml.contrib.tensorboard.export import export_to_tensorboard\n",
+ "import os\n",
+ "import tensorflow as tf\n",
+ "\n",
+ "logdir = 'exportedTBlogs'\n",
+ "log_path = os.path.join(os.getcwd(), logdir)\n",
+ "try:\n",
+ " os.stat(log_path)\n",
+ "except os.error:\n",
+ " os.mkdir(log_path)\n",
+ "print(logdir)\n",
+ "\n",
+ "# export run history for the project\n",
+ "export_to_tensorboard(root_run, logdir)\n",
+ "\n",
+ "# or export a particular run\n",
+ "# export_to_tensorboard(run, logdir)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "root_run.complete()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Start Tensorboard\n",
+ "\n",
+ "Or you can start the Tensorboard outside this notebook to view the result"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.contrib.tensorboard import Tensorboard\n",
+ "\n",
+ "# The Tensorboard constructor takes an array of runs, so be sure and pass it in as a single-element array here\n",
+ "tb = Tensorboard([], local_root=logdir, port=6006)\n",
+ "\n",
+ "# If successful, start() returns a string with the URI of the instance.\n",
+ "tb.start()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Stop Tensorboard\n",
+ "\n",
+ "When you're done, make sure to call the `stop()` method of the Tensorboard object."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "tb.stop()"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "roastala"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.5"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.5"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/training-with-deep-learning/tensorboard/tensorboard.ipynb b/how-to-use-azureml/training-with-deep-learning/tensorboard/tensorboard.ipynb
index dfff67c7..44c71353 100644
--- a/how-to-use-azureml/training-with-deep-learning/tensorboard/tensorboard.ipynb
+++ b/how-to-use-azureml/training-with-deep-learning/tensorboard/tensorboard.ipynb
@@ -1,568 +1,568 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Tensorboard Integration with Run History\n",
- "\n",
- "1. Run a Tensorflow job locally and view its TB output live.\n",
- "2. The same, for a DSVM.\n",
- "3. And once more, with an AmlCompute cluster.\n",
- "4. Finally, we'll collect all of these historical runs together into a single Tensorboard graph."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Prerequisites\n",
- "* Understand the [architecture and terms](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture) introduced by Azure Machine Learning\n",
- "* Go through the [00.configuration.ipynb](https://github.com/Azure/MachineLearningNotebooks/blob/master/00.configuration.ipynb) notebook to:\n",
- " * install the AML SDK\n",
- " * create a workspace and its configuration file (`config.json`)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Check core SDK version number\n",
- "import azureml.core\n",
- "\n",
- "print(\"SDK version:\", azureml.core.VERSION)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Install the Azure ML TensorBoard package."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "!pip install azureml-contrib-tensorboard"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Diagnostics\n",
- "Opt-in diagnostics for better experience, quality, and security of future releases."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "Diagnostics"
- ]
- },
- "outputs": [],
- "source": [
- "from azureml.telemetry import set_diagnostics_collection\n",
- "\n",
- "set_diagnostics_collection(send_diagnostics=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Initialize Workspace\n",
- "\n",
- "Initialize a workspace object from persisted configuration."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core import Workspace\n",
- "\n",
- "ws = Workspace.from_config()\n",
- "print('Workspace name: ' + ws.name, \n",
- " 'Azure region: ' + ws.location, \n",
- " 'Subscription id: ' + ws.subscription_id, \n",
- " 'Resource group: ' + ws.resource_group, sep = '\\n')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Set experiment name and create project\n",
- "Choose a name for your run history container in the workspace, and create a folder for the project."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from os import path, makedirs\n",
- "experiment_name = 'tensorboard-demo'\n",
- "\n",
- "# experiment folder\n",
- "exp_dir = './sample_projects/' + experiment_name\n",
- "\n",
- "if not path.exists(exp_dir):\n",
- " makedirs(exp_dir)\n",
- "\n",
- "# runs we started in this session, for the finale\n",
- "runs = []"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Download Tensorflow Tensorboard demo code\n",
- "\n",
- "Tensorflow's repository has an MNIST demo with extensive Tensorboard instrumentation. We'll use it here for our purposes.\n",
- "\n",
- "Note that we don't need to make any code changes at all - the code works without modification from the Tensorflow repository."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import requests\n",
- "import os\n",
- "import tempfile\n",
- "tf_code = requests.get(\"https://raw.githubusercontent.com/tensorflow/tensorflow/r1.8/tensorflow/examples/tutorials/mnist/mnist_with_summaries.py\")\n",
- "with open(os.path.join(exp_dir, \"mnist_with_summaries.py\"), \"w\") as file:\n",
- " file.write(tf_code.text)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Configure and run locally\n",
- "\n",
- "We'll start by running this locally. While it might not initially seem that useful to use this for a local run - why not just run TB against the files generated locally? - even in this case there is some value to using this feature. Your local run will be registered in the run history, and your Tensorboard logs will be uploaded to the artifact store associated with this run. Later, you'll be able to restore the logs from any run, regardless of where it happened.\n",
- "\n",
- "Note that for this run, you will need to install Tensorflow on your local machine by yourself. Further, the Tensorboard module (that is, the one included with Tensorflow) must be accessible to this notebook's kernel, as the local machine is what runs Tensorboard."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.runconfig import RunConfiguration\n",
- "\n",
- "# Create a run configuration.\n",
- "run_config = RunConfiguration()\n",
- "run_config.environment.python.user_managed_dependencies = True\n",
- "\n",
- "# You can choose a specific Python environment by pointing to a Python path \n",
- "#run_config.environment.python.interpreter_path = '/home/ninghai/miniconda3/envs/sdk2/bin/python'"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core import Experiment, Run\n",
- "from azureml.core.script_run_config import ScriptRunConfig\n",
- "import tensorflow as tf\n",
- "\n",
- "logs_dir = os.path.join(os.curdir, \"logs\")\n",
- "data_dir = os.path.abspath(os.path.join(os.curdir, \"mnist_data\"))\n",
- "\n",
- "if not path.exists(data_dir):\n",
- " makedirs(data_dir)\n",
- "\n",
- "os.environ[\"TEST_TMPDIR\"] = data_dir\n",
- "\n",
- "# Writing logs to ./logs results in their being uploaded to Artifact Service,\n",
- "# and thus, made accessible to our Tensorboard instance.\n",
- "arguments_list = [\"--log_dir\", logs_dir]\n",
- "\n",
- "# Create an experiment\n",
- "exp = Experiment(ws, experiment_name)\n",
- "\n",
- "# If you would like the run to go for longer, add --max_steps 5000 to the arguments list:\n",
- "# arguments_list += [\"--max_steps\", \"5000\"]\n",
- "\n",
- "script = ScriptRunConfig(exp_dir,\n",
- " script=\"mnist_with_summaries.py\",\n",
- " run_config=run_config,\n",
- " arguments=arguments_list)\n",
- "\n",
- "run = exp.submit(script)\n",
- "# You can also wait for the run to complete\n",
- "# run.wait_for_completion(show_output=True)\n",
- "runs.append(run)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Start Tensorboard\n",
- "\n",
- "Now, while the run is in progress, we just need to start Tensorboard with the run as its target, and it will begin streaming logs."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.contrib.tensorboard import Tensorboard\n",
- "\n",
- "# The Tensorboard constructor takes an array of runs, so be sure and pass it in as a single-element array here\n",
- "tb = Tensorboard([run])\n",
- "\n",
- "# If successful, start() returns a string with the URI of the instance.\n",
- "tb.start()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Stop Tensorboard\n",
- "\n",
- "When you're done, make sure to call the `stop()` method of the Tensorboard object, or it will stay running even after your job completes."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "tb.stop()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Now, with a DSVM\n",
- "\n",
- "Tensorboard uploading works with all compute targets. Here we demonstrate it from a DSVM.\n",
- "Note that the Tensorboard instance itself will be run by the notebook kernel. Again, this means this notebook's kernel must have access to the Tensorboard module.\n",
- "\n",
- "If you are unfamiliar with DSVM configuration, check [04. Train in a remote VM](04.train-on-remote-vm.ipynb) for a more detailed breakdown.\n",
- "\n",
- "**Note**: To streamline the compute that Azure Machine Learning creates, we are making updates to support creating only single to multi-node `AmlCompute`. The `DSVMCompute` class will be deprecated in a later release, but the DSVM can be created using the below single line command and then attached(like any VM) using the sample code below. Also note, that we only support Linux VMs for remote execution from AML and the commands below will spin a Linux VM only.\n",
- "\n",
- "```shell\n",
- "# create a DSVM in your resource group\n",
- "# note you need to be at least a contributor to the resource group in order to execute this command successfully.\n",
- "(myenv) $ az vm create --resource-group --name --image microsoft-dsvm:linux-data-science-vm-ubuntu:linuxdsvmubuntu:latest --admin-username --admin-password --generate-ssh-keys --authentication-type password\n",
- "```\n",
- "You can also use [this url](https://portal.azure.com/#create/microsoft-dsvm.linux-data-science-vm-ubuntulinuxdsvmubuntu) to create the VM using the Azure Portal."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.compute import RemoteCompute\n",
- "from azureml.core.compute_target import ComputeTargetException\n",
- "import os\n",
- "\n",
- "username = os.getenv('AZUREML_DSVM_USERNAME', default='')\n",
- "address = os.getenv('AZUREML_DSVM_ADDRESS', default='')\n",
- "\n",
- "compute_target_name = 'cpudsvm'\n",
- "# if you want to connect using SSH key instead of username/password you can provide parameters private_key_file and private_key_passphrase \n",
- "try:\n",
- " attached_dsvm_compute = RemoteCompute(workspace=ws, name=compute_target_name)\n",
- " print('found existing:', attached_dsvm_compute.name)\n",
- "except ComputeTargetException:\n",
- " attached_dsvm_compute = RemoteCompute.attach(workspace=ws,\n",
- " name=compute_target_name,\n",
- " username=username,\n",
- " address=address,\n",
- " ssh_port=22,\n",
- " private_key_file='./.ssh/id_rsa')\n",
- " \n",
- " attached_dsvm_compute.wait_for_completion(show_output=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Submit run using TensorFlow estimator\n",
- "\n",
- "Instead of manually configuring the DSVM environment, we can use the TensorFlow estimator and everything is set up automatically."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.train.dnn import TensorFlow\n",
- "\n",
- "script_params = {\"--log_dir\": \"./logs\"}\n",
- "\n",
- "# If you want the run to go longer, set --max-steps to a higher number.\n",
- "# script_params[\"--max_steps\"] = \"5000\"\n",
- "\n",
- "tf_estimator = TensorFlow(source_directory=exp_dir,\n",
- " compute_target=attached_dsvm_compute,\n",
- " entry_script='mnist_with_summaries.py',\n",
- " script_params=script_params)\n",
- "\n",
- "run = exp.submit(tf_estimator)\n",
- "\n",
- "runs.append(run)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Start Tensorboard with this run\n",
- "\n",
- "Just like before."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# The Tensorboard constructor takes an array of runs, so be sure and pass it in as a single-element array here\n",
- "tb = Tensorboard([run])\n",
- "\n",
- "# If successful, start() returns a string with the URI of the instance.\n",
- "tb.start()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Stop Tensorboard\n",
- "\n",
- "When you're done, make sure to call the `stop()` method of the Tensorboard object, or it will stay running even after your job completes."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "tb.stop()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Once more, with an AmlCompute cluster\n",
- "\n",
- "Just to prove we can, let's create an AmlCompute CPU cluster, and run our demo there, as well."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.compute import ComputeTarget, AmlCompute\n",
- "from azureml.core.compute_target import ComputeTargetException\n",
- "\n",
- "# choose a name for your cluster\n",
- "cluster_name = \"cpucluster\"\n",
- "\n",
- "try:\n",
- " compute_target = ComputeTarget(workspace=ws, name=cluster_name)\n",
- " print('Found existing compute target.')\n",
- "except ComputeTargetException:\n",
- " print('Creating a new compute target...')\n",
- " compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_D2_V2', \n",
- " max_nodes=4)\n",
- "\n",
- " # create the cluster\n",
- " compute_target = ComputeTarget.create(ws, cluster_name, compute_config)\n",
- "\n",
- "compute_target.wait_for_completion(show_output=True, min_node_count=1, timeout_in_minutes=20)\n",
- "\n",
- "# Use the 'status' property to get a detailed status for the current cluster. \n",
- "print(compute_target.status.serialize())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Submit run using TensorFlow estimator\n",
- "\n",
- "Again, we can use the TensorFlow estimator and everything is set up automatically."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "script_params = {\"--log_dir\": \"./logs\"}\n",
- "\n",
- "# If you want the run to go longer, set --max-steps to a higher number.\n",
- "# script_params[\"--max_steps\"] = \"5000\"\n",
- "\n",
- "tf_estimator = TensorFlow(source_directory=exp_dir,\n",
- " compute_target=compute_target,\n",
- " entry_script='mnist_with_summaries.py',\n",
- " script_params=script_params)\n",
- "\n",
- "run = exp.submit(tf_estimator)\n",
- "\n",
- "runs.append(run)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Start Tensorboard with this run\n",
- "\n",
- "Once more..."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# The Tensorboard constructor takes an array of runs, so be sure and pass it in as a single-element array here\n",
- "tb = Tensorboard([run])\n",
- "\n",
- "# If successful, start() returns a string with the URI of the instance.\n",
- "tb.start()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Stop Tensorboard\n",
- "\n",
- "When you're done, make sure to call the `stop()` method of the Tensorboard object, or it will stay running even after your job completes."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "tb.stop()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Finale\n",
- "\n",
- "If you've paid close attention, you'll have noticed that we've been saving the run objects in an array as we went along. We can start a Tensorboard instance that combines all of these run objects into a single process. This way, you can compare historical runs. You can even do this with live runs; if you made some of those previous runs longer via the `--max_steps` parameter, they might still be running, and you'll see them live in this instance as well."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# The Tensorboard constructor takes an array of runs...\n",
- "# and it turns out that we have been building one of those all along.\n",
- "tb = Tensorboard(runs)\n",
- "\n",
- "# If successful, start() returns a string with the URI of the instance.\n",
- "tb.start()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Stop Tensorboard\n",
- "\n",
- "As you might already know, make sure to call the `stop()` method of the Tensorboard object, or it will stay running (until you kill the kernel associated with this notebook, at least)."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "tb.stop()"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "roastala"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Tensorboard Integration with Run History\n",
+ "\n",
+ "1. Run a Tensorflow job locally and view its TB output live.\n",
+ "2. The same, for a DSVM.\n",
+ "3. And once more, with an AmlCompute cluster.\n",
+ "4. Finally, we'll collect all of these historical runs together into a single Tensorboard graph."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Prerequisites\n",
+ "* Understand the [architecture and terms](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture) introduced by Azure Machine Learning\n",
+ "* Go through the [configuration notebook](../../../configuration.ipynb) notebook to:\n",
+ " * install the AML SDK\n",
+ " * create a workspace and its configuration file (`config.json`)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Check core SDK version number\n",
+ "import azureml.core\n",
+ "\n",
+ "print(\"SDK version:\", azureml.core.VERSION)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Install the Azure ML TensorBoard package."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "!pip install azureml-contrib-tensorboard"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Diagnostics\n",
+ "Opt-in diagnostics for better experience, quality, and security of future releases."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "Diagnostics"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "from azureml.telemetry import set_diagnostics_collection\n",
+ "\n",
+ "set_diagnostics_collection(send_diagnostics=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Initialize Workspace\n",
+ "\n",
+ "Initialize a workspace object from persisted configuration."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core import Workspace\n",
+ "\n",
+ "ws = Workspace.from_config()\n",
+ "print('Workspace name: ' + ws.name, \n",
+ " 'Azure region: ' + ws.location, \n",
+ " 'Subscription id: ' + ws.subscription_id, \n",
+ " 'Resource group: ' + ws.resource_group, sep = '\\n')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Set experiment name and create project\n",
+ "Choose a name for your run history container in the workspace, and create a folder for the project."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from os import path, makedirs\n",
+ "experiment_name = 'tensorboard-demo'\n",
+ "\n",
+ "# experiment folder\n",
+ "exp_dir = './sample_projects/' + experiment_name\n",
+ "\n",
+ "if not path.exists(exp_dir):\n",
+ " makedirs(exp_dir)\n",
+ "\n",
+ "# runs we started in this session, for the finale\n",
+ "runs = []"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Download Tensorflow Tensorboard demo code\n",
+ "\n",
+ "Tensorflow's repository has an MNIST demo with extensive Tensorboard instrumentation. We'll use it here for our purposes.\n",
+ "\n",
+ "Note that we don't need to make any code changes at all - the code works without modification from the Tensorflow repository."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import requests\n",
+ "import os\n",
+ "import tempfile\n",
+ "tf_code = requests.get(\"https://raw.githubusercontent.com/tensorflow/tensorflow/r1.8/tensorflow/examples/tutorials/mnist/mnist_with_summaries.py\")\n",
+ "with open(os.path.join(exp_dir, \"mnist_with_summaries.py\"), \"w\") as file:\n",
+ " file.write(tf_code.text)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Configure and run locally\n",
+ "\n",
+ "We'll start by running this locally. While it might not initially seem that useful to use this for a local run - why not just run TB against the files generated locally? - even in this case there is some value to using this feature. Your local run will be registered in the run history, and your Tensorboard logs will be uploaded to the artifact store associated with this run. Later, you'll be able to restore the logs from any run, regardless of where it happened.\n",
+ "\n",
+ "Note that for this run, you will need to install Tensorflow on your local machine by yourself. Further, the Tensorboard module (that is, the one included with Tensorflow) must be accessible to this notebook's kernel, as the local machine is what runs Tensorboard."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.runconfig import RunConfiguration\n",
+ "\n",
+ "# Create a run configuration.\n",
+ "run_config = RunConfiguration()\n",
+ "run_config.environment.python.user_managed_dependencies = True\n",
+ "\n",
+ "# You can choose a specific Python environment by pointing to a Python path \n",
+ "#run_config.environment.python.interpreter_path = '/home/ninghai/miniconda3/envs/sdk2/bin/python'"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core import Experiment, Run\n",
+ "from azureml.core.script_run_config import ScriptRunConfig\n",
+ "import tensorflow as tf\n",
+ "\n",
+ "logs_dir = os.path.join(os.curdir, \"logs\")\n",
+ "data_dir = os.path.abspath(os.path.join(os.curdir, \"mnist_data\"))\n",
+ "\n",
+ "if not path.exists(data_dir):\n",
+ " makedirs(data_dir)\n",
+ "\n",
+ "os.environ[\"TEST_TMPDIR\"] = data_dir\n",
+ "\n",
+ "# Writing logs to ./logs results in their being uploaded to Artifact Service,\n",
+ "# and thus, made accessible to our Tensorboard instance.\n",
+ "arguments_list = [\"--log_dir\", logs_dir]\n",
+ "\n",
+ "# Create an experiment\n",
+ "exp = Experiment(ws, experiment_name)\n",
+ "\n",
+ "# If you would like the run to go for longer, add --max_steps 5000 to the arguments list:\n",
+ "# arguments_list += [\"--max_steps\", \"5000\"]\n",
+ "\n",
+ "script = ScriptRunConfig(exp_dir,\n",
+ " script=\"mnist_with_summaries.py\",\n",
+ " run_config=run_config,\n",
+ " arguments=arguments_list)\n",
+ "\n",
+ "run = exp.submit(script)\n",
+ "# You can also wait for the run to complete\n",
+ "# run.wait_for_completion(show_output=True)\n",
+ "runs.append(run)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Start Tensorboard\n",
+ "\n",
+ "Now, while the run is in progress, we just need to start Tensorboard with the run as its target, and it will begin streaming logs."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.contrib.tensorboard import Tensorboard\n",
+ "\n",
+ "# The Tensorboard constructor takes an array of runs, so be sure and pass it in as a single-element array here\n",
+ "tb = Tensorboard([run])\n",
+ "\n",
+ "# If successful, start() returns a string with the URI of the instance.\n",
+ "tb.start()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Stop Tensorboard\n",
+ "\n",
+ "When you're done, make sure to call the `stop()` method of the Tensorboard object, or it will stay running even after your job completes."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "tb.stop()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Now, with a DSVM\n",
+ "\n",
+ "Tensorboard uploading works with all compute targets. Here we demonstrate it from a DSVM.\n",
+ "Note that the Tensorboard instance itself will be run by the notebook kernel. Again, this means this notebook's kernel must have access to the Tensorboard module.\n",
+ "\n",
+ "If you are unfamiliar with DSVM configuration, check [04. Train in a remote VM](04.train-on-remote-vm.ipynb) for a more detailed breakdown.\n",
+ "\n",
+ "**Note**: To streamline the compute that Azure Machine Learning creates, we are making updates to support creating only single to multi-node `AmlCompute`. The `DSVMCompute` class will be deprecated in a later release, but the DSVM can be created using the below single line command and then attached(like any VM) using the sample code below. Also note, that we only support Linux VMs for remote execution from AML and the commands below will spin a Linux VM only.\n",
+ "\n",
+ "```shell\n",
+ "# create a DSVM in your resource group\n",
+ "# note you need to be at least a contributor to the resource group in order to execute this command successfully.\n",
+ "(myenv) $ az vm create --resource-group --name --image microsoft-dsvm:linux-data-science-vm-ubuntu:linuxdsvmubuntu:latest --admin-username --admin-password --generate-ssh-keys --authentication-type password\n",
+ "```\n",
+ "You can also use [this url](https://portal.azure.com/#create/microsoft-dsvm.linux-data-science-vm-ubuntulinuxdsvmubuntu) to create the VM using the Azure Portal."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.compute import RemoteCompute\n",
+ "from azureml.core.compute_target import ComputeTargetException\n",
+ "import os\n",
+ "\n",
+ "username = os.getenv('AZUREML_DSVM_USERNAME', default='')\n",
+ "address = os.getenv('AZUREML_DSVM_ADDRESS', default='')\n",
+ "\n",
+ "compute_target_name = 'cpudsvm'\n",
+ "# if you want to connect using SSH key instead of username/password you can provide parameters private_key_file and private_key_passphrase \n",
+ "try:\n",
+ " attached_dsvm_compute = RemoteCompute(workspace=ws, name=compute_target_name)\n",
+ " print('found existing:', attached_dsvm_compute.name)\n",
+ "except ComputeTargetException:\n",
+ " attached_dsvm_compute = RemoteCompute.attach(workspace=ws,\n",
+ " name=compute_target_name,\n",
+ " username=username,\n",
+ " address=address,\n",
+ " ssh_port=22,\n",
+ " private_key_file='./.ssh/id_rsa')\n",
+ " \n",
+ " attached_dsvm_compute.wait_for_completion(show_output=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Submit run using TensorFlow estimator\n",
+ "\n",
+ "Instead of manually configuring the DSVM environment, we can use the TensorFlow estimator and everything is set up automatically."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.train.dnn import TensorFlow\n",
+ "\n",
+ "script_params = {\"--log_dir\": \"./logs\"}\n",
+ "\n",
+ "# If you want the run to go longer, set --max-steps to a higher number.\n",
+ "# script_params[\"--max_steps\"] = \"5000\"\n",
+ "\n",
+ "tf_estimator = TensorFlow(source_directory=exp_dir,\n",
+ " compute_target=attached_dsvm_compute,\n",
+ " entry_script='mnist_with_summaries.py',\n",
+ " script_params=script_params)\n",
+ "\n",
+ "run = exp.submit(tf_estimator)\n",
+ "\n",
+ "runs.append(run)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Start Tensorboard with this run\n",
+ "\n",
+ "Just like before."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# The Tensorboard constructor takes an array of runs, so be sure and pass it in as a single-element array here\n",
+ "tb = Tensorboard([run])\n",
+ "\n",
+ "# If successful, start() returns a string with the URI of the instance.\n",
+ "tb.start()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Stop Tensorboard\n",
+ "\n",
+ "When you're done, make sure to call the `stop()` method of the Tensorboard object, or it will stay running even after your job completes."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "tb.stop()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Once more, with an AmlCompute cluster\n",
+ "\n",
+ "Just to prove we can, let's create an AmlCompute CPU cluster, and run our demo there, as well."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.compute import ComputeTarget, AmlCompute\n",
+ "from azureml.core.compute_target import ComputeTargetException\n",
+ "\n",
+ "# choose a name for your cluster\n",
+ "cluster_name = \"cpucluster\"\n",
+ "\n",
+ "try:\n",
+ " compute_target = ComputeTarget(workspace=ws, name=cluster_name)\n",
+ " print('Found existing compute target.')\n",
+ "except ComputeTargetException:\n",
+ " print('Creating a new compute target...')\n",
+ " compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_D2_V2', \n",
+ " max_nodes=4)\n",
+ "\n",
+ " # create the cluster\n",
+ " compute_target = ComputeTarget.create(ws, cluster_name, compute_config)\n",
+ "\n",
+ "compute_target.wait_for_completion(show_output=True, min_node_count=1, timeout_in_minutes=20)\n",
+ "\n",
+ "# use get_status() to get a detailed status for the current cluster. \n",
+ "print(compute_target.get_status().serialize())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Submit run using TensorFlow estimator\n",
+ "\n",
+ "Again, we can use the TensorFlow estimator and everything is set up automatically."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "script_params = {\"--log_dir\": \"./logs\"}\n",
+ "\n",
+ "# If you want the run to go longer, set --max-steps to a higher number.\n",
+ "# script_params[\"--max_steps\"] = \"5000\"\n",
+ "\n",
+ "tf_estimator = TensorFlow(source_directory=exp_dir,\n",
+ " compute_target=compute_target,\n",
+ " entry_script='mnist_with_summaries.py',\n",
+ " script_params=script_params)\n",
+ "\n",
+ "run = exp.submit(tf_estimator)\n",
+ "\n",
+ "runs.append(run)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Start Tensorboard with this run\n",
+ "\n",
+ "Once more..."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# The Tensorboard constructor takes an array of runs, so be sure and pass it in as a single-element array here\n",
+ "tb = Tensorboard([run])\n",
+ "\n",
+ "# If successful, start() returns a string with the URI of the instance.\n",
+ "tb.start()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Stop Tensorboard\n",
+ "\n",
+ "When you're done, make sure to call the `stop()` method of the Tensorboard object, or it will stay running even after your job completes."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "tb.stop()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Finale\n",
+ "\n",
+ "If you've paid close attention, you'll have noticed that we've been saving the run objects in an array as we went along. We can start a Tensorboard instance that combines all of these run objects into a single process. This way, you can compare historical runs. You can even do this with live runs; if you made some of those previous runs longer via the `--max_steps` parameter, they might still be running, and you'll see them live in this instance as well."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# The Tensorboard constructor takes an array of runs...\n",
+ "# and it turns out that we have been building one of those all along.\n",
+ "tb = Tensorboard(runs)\n",
+ "\n",
+ "# If successful, start() returns a string with the URI of the instance.\n",
+ "tb.start()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Stop Tensorboard\n",
+ "\n",
+ "As you might already know, make sure to call the `stop()` method of the Tensorboard object, or it will stay running (until you kill the kernel associated with this notebook, at least)."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "tb.stop()"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "roastala"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.6"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/training-with-deep-learning/train-hyperparameter-tune-deploy-with-pytorch/train-hyperparameter-tune-deploy-with-pytorch.ipynb b/how-to-use-azureml/training-with-deep-learning/train-hyperparameter-tune-deploy-with-pytorch/train-hyperparameter-tune-deploy-with-pytorch.ipynb
index 4347c209..e5d0d3bb 100644
--- a/how-to-use-azureml/training-with-deep-learning/train-hyperparameter-tune-deploy-with-pytorch/train-hyperparameter-tune-deploy-with-pytorch.ipynb
+++ b/how-to-use-azureml/training-with-deep-learning/train-hyperparameter-tune-deploy-with-pytorch/train-hyperparameter-tune-deploy-with-pytorch.ipynb
@@ -1,740 +1,740 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved. \n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Train, hyperparameter tune, and deploy with PyTorch\n",
- "\n",
- "In this tutorial, you will train, hyperparameter tune, and deploy a PyTorch model using the Azure Machine Learning (Azure ML) Python SDK.\n",
- "\n",
- "This tutorial will train an image classification model using transfer learning, based on PyTorch's [Transfer Learning tutorial](https://pytorch.org/tutorials/beginner/transfer_learning_tutorial.html). The model is trained to classify ants and bees by first using a pretrained ResNet18 model that has been trained on the [ImageNet](http://image-net.org/index) dataset."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Prerequisites\n",
- "* Go through the [Configuration](https://github.com/Azure/MachineLearningNotebooks/blob/master/configuration.ipynb) notebook to install the Azure Machine Learning Python SDK and create an Azure ML `Workspace`"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Check core SDK version number\n",
- "import azureml.core\n",
- "\n",
- "print(\"SDK version:\", azureml.core.VERSION)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Diagnostics\n",
- "Opt-in diagnostics for better experience, quality, and security of future releases."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "Diagnostics"
- ]
- },
- "outputs": [],
- "source": [
- "from azureml.telemetry import set_diagnostics_collection\n",
- "\n",
- "set_diagnostics_collection(send_diagnostics=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Initialize workspace\n",
- "Initialize a [Workspace](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#workspace) object from the existing workspace you created in the Prerequisites step. `Workspace.from_config()` creates a workspace object from the details stored in `config.json`."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.workspace import Workspace\n",
- "\n",
- "ws = Workspace.from_config()\n",
- "print('Workspace name: ' + ws.name, \n",
- " 'Azure region: ' + ws.location, \n",
- " 'Subscription id: ' + ws.subscription_id, \n",
- " 'Resource group: ' + ws.resource_group, sep = '\\n')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Create or Attach existing AmlCompute\n",
- "You will need to create a [compute target](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#compute-target) for training your model. In this tutorial, we use Azure ML managed compute ([AmlCompute](https://docs.microsoft.com/azure/machine-learning/service/how-to-set-up-training-targets#amlcompute)) for our remote training compute resource.\n",
- "\n",
- "**Creation of AmlCompute takes approximately 5 minutes.** If the AmlCompute with that name is already in your workspace, this code will skip the creation process.\n",
- "\n",
- "As with other Azure services, there are limits on certain resources (e.g. AmlCompute) associated with the Azure Machine Learning service. Please read [this article](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-manage-quotas) on the default limits and how to request more quota."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.compute import ComputeTarget, AmlCompute\n",
- "from azureml.core.compute_target import ComputeTargetException\n",
- "\n",
- "# choose a name for your cluster\n",
- "cluster_name = \"gpucluster\"\n",
- "\n",
- "try:\n",
- " compute_target = ComputeTarget(workspace=ws, name=cluster_name)\n",
- " print('Found existing compute target.')\n",
- "except ComputeTargetException:\n",
- " print('Creating a new compute target...')\n",
- " compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_NC6', \n",
- " max_nodes=4)\n",
- "\n",
- " # create the cluster\n",
- " compute_target = ComputeTarget.create(ws, cluster_name, compute_config)\n",
- "\n",
- " compute_target.wait_for_completion(show_output=True)\n",
- "\n",
- "# Use the 'status' property to get a detailed status for the current cluster. \n",
- "print(compute_target.status.serialize())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "The above code creates a GPU cluster. If you instead want to create a CPU cluster, provide a different VM size to the `vm_size` parameter, such as `STANDARD_D2_V2`."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Train model on the remote compute\n",
- "Now that you have your data and training script prepared, you are ready to train on your remote compute cluster. You can take advantage of Azure compute to leverage GPUs to cut down your training time. "
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create a project directory\n",
- "Create a directory that will contain all the necessary code from your local machine that you will need access to on the remote resource. This includes the training script and any additional files your training script depends on."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import os\n",
- "\n",
- "project_folder = './pytorch-hymenoptera'\n",
- "os.makedirs(project_folder, exist_ok=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Download training data\n",
- "The dataset we will use (located [here](https://download.pytorch.org/tutorial/hymenoptera_data.zip) as a zip file) consists of about 120 training images each for ants and bees, with 75 validation images for each class. [Hymenoptera](https://en.wikipedia.org/wiki/Hymenoptera) is the order of insects that includes ants and bees. We will download and extract the dataset as part of our training script `pytorch_train.py`"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Prepare training script\n",
- "Now you will need to create your training script. In this tutorial, the training script is already provided for you at `pytorch_train.py`. In practice, you should be able to take any custom training script as is and run it with Azure ML without having to modify your code.\n",
- "\n",
- "However, if you would like to use Azure ML's [tracking and metrics](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#metrics) capabilities, you will have to add a small amount of Azure ML code inside your training script. \n",
- "\n",
- "In `pytorch_train.py`, we will log some metrics to our Azure ML run. To do so, we will access the Azure ML `Run` object within the script:\n",
- "```Python\n",
- "from azureml.core.run import Run\n",
- "run = Run.get_context()\n",
- "```\n",
- "Further within `pytorch_train.py`, we log the learning rate and momentum parameters, and the best validation accuracy the model achieves:\n",
- "```Python\n",
- "run.log('lr', np.float(learning_rate))\n",
- "run.log('momentum', np.float(momentum))\n",
- "\n",
- "run.log('best_val_acc', np.float(best_acc))\n",
- "```\n",
- "These run metrics will become particularly important when we begin hyperparameter tuning our model in the \"Tune model hyperparameters\" section."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Once your script is ready, copy the training script `pytorch_train.py` into your project directory."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import shutil\n",
- "\n",
- "shutil.copy('pytorch_train.py', project_folder)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create an experiment\n",
- "Create an [Experiment](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#experiment) to track all the runs in your workspace for this transfer learning PyTorch tutorial. "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core import Experiment\n",
- "\n",
- "experiment_name = 'pytorch-hymenoptera'\n",
- "experiment = Experiment(ws, name=experiment_name)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create a PyTorch estimator\n",
- "The Azure ML SDK's PyTorch estimator enables you to easily submit PyTorch training jobs for both single-node and distributed runs. For more information on the PyTorch estimator, refer [here](https://docs.microsoft.com/azure/machine-learning/service/how-to-train-pytorch). The following code will define a single-node PyTorch job."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.train.dnn import PyTorch\n",
- "\n",
- "script_params = {\n",
- " '--num_epochs': 30,\n",
- " '--output_dir': './outputs'\n",
- "}\n",
- "\n",
- "estimator = PyTorch(source_directory=project_folder, \n",
- " script_params=script_params,\n",
- " compute_target=compute_target,\n",
- " entry_script='pytorch_train.py',\n",
- " use_gpu=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "The `script_params` parameter is a dictionary containing the command-line arguments to your training script `entry_script`. Please note the following:\n",
- "- We passed our training data reference `ds_data` to our script's `--data_dir` argument. This will 1) mount our datastore on the remote compute and 2) provide the path to the training data `hymenoptera_data` on our datastore.\n",
- "- We specified the output directory as `./outputs`. The `outputs` directory is specially treated by Azure ML in that all the content in this directory gets uploaded to your workspace as part of your run history. The files written to this directory are therefore accessible even once your remote run is over. In this tutorial, we will save our trained model to this output directory.\n",
- "\n",
- "To leverage the Azure VM's GPU for training, we set `use_gpu=True`."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Submit job\n",
- "Run your experiment by submitting your estimator object. Note that this call is asynchronous."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run = experiment.submit(estimator)\n",
- "print(run)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# to get more details of your run\n",
- "print(run.get_details())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Monitor your run\n",
- "You can monitor the progress of the run with a Jupyter widget. Like the run submission, the widget is asynchronous and provides live updates every 10-15 seconds until the job completes."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.widgets import RunDetails\n",
- "\n",
- "RunDetails(run).show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Alternatively, you can block until the script has completed training before running more code."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run.wait_for_completion(show_output=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Tune model hyperparameters\n",
- "Now that we've seen how to do a simple PyTorch training run using the SDK, let's see if we can further improve the accuracy of our model. We can optimize our model's hyperparameters using Azure Machine Learning's hyperparameter tuning capabilities."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Start a hyperparameter sweep\n",
- "First, we will define the hyperparameter space to sweep over. Since our training script uses a learning rate schedule to decay the learning rate every several epochs, let's tune the initial learning rate and the momentum parameters. In this example we will use random sampling to try different configuration sets of hyperparameters to maximize our primary metric, the best validation accuracy (`best_val_acc`).\n",
- "\n",
- "Then, we specify the early termination policy to use to early terminate poorly performing runs. Here we use the `BanditPolicy`, which will terminate any run that doesn't fall within the slack factor of our primary evaluation metric. In this tutorial, we will apply this policy every epoch (since we report our `best_val_acc` metric every epoch and `evaluation_interval=1`). Notice we will delay the first policy evaluation until after the first `10` epochs (`delay_evaluation=10`).\n",
- "Refer [here](https://docs.microsoft.com/azure/machine-learning/service/how-to-tune-hyperparameters#specify-an-early-termination-policy) for more information on the BanditPolicy and other policies available."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.train.hyperdrive import *\n",
- "\n",
- "param_sampling = RandomParameterSampling( {\n",
- " 'learning_rate': uniform(0.0005, 0.005),\n",
- " 'momentum': uniform(0.9, 0.99)\n",
- " }\n",
- ")\n",
- "\n",
- "early_termination_policy = BanditPolicy(slack_factor=0.15, evaluation_interval=1, delay_evaluation=10)\n",
- "\n",
- "hyperdrive_run_config = HyperDriveRunConfig(estimator=estimator,\n",
- " hyperparameter_sampling=param_sampling, \n",
- " policy=early_termination_policy,\n",
- " primary_metric_name='best_val_acc',\n",
- " primary_metric_goal=PrimaryMetricGoal.MAXIMIZE,\n",
- " max_total_runs=8,\n",
- " max_concurrent_runs=4)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Finally, lauch the hyperparameter tuning job."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# start the HyperDrive run\n",
- "hyperdrive_run = experiment.submit(hyperdrive_run_config)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Monitor HyperDrive runs\n",
- "You can monitor the progress of the runs with the following Jupyter widget. "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.widgets import RunDetails\n",
- "\n",
- "RunDetails(hyperdrive_run).show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Or block until the HyperDrive sweep has completed:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "hyperdrive_run.wait_for_completion(show_output=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Find and register the best model\n",
- "Once all the runs complete, we can find the run that produced the model with the highest accuracy."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "best_run = hyperdrive_run.get_best_run_by_primary_metric()\n",
- "best_run_metrics = best_run.get_metrics()\n",
- "print(best_run)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "print('Best Run is:\\n Validation accuracy: {0:.5f} \\n Learning rate: {1:.5f} \\n Momentum: {2:.5f}'.format(\n",
- " best_run_metrics['best_val_acc'][-1],\n",
- " best_run_metrics['lr'],\n",
- " best_run_metrics['momentum'])\n",
- " )"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Finally, register the model from your best-performing run to your workspace. The `model_path` parameter takes in the relative path on the remote VM to the model file in your `outputs` directory. In the next section, we will deploy this registered model as a web service."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "model = best_run.register_model(model_name = 'pytorch-hymenoptera', model_path = 'outputs/model.pt')\n",
- "print(model.name, model.id, model.version, sep = '\\t')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Deploy model as web service\n",
- "Once you have your trained model, you can deploy the model on Azure. In this tutorial, we will deploy the model as a web service in [Azure Container Instances](https://docs.microsoft.com/en-us/azure/container-instances/) (ACI). For more information on deploying models using Azure ML, refer [here](https://docs.microsoft.com/azure/machine-learning/service/how-to-deploy-and-where)."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create scoring script\n",
- "\n",
- "First, we will create a scoring script that will be invoked by the web service call. Note that the scoring script must have two required functions:\n",
- "* `init()`: In this function, you typically load the model into a `global` object. This function is executed only once when the Docker container is started. \n",
- "* `run(input_data)`: In this function, the model is used to predict a value based on the input data. The input and output typically use JSON as serialization and deserialization format, but you are not limited to that.\n",
- "\n",
- "Refer to the scoring script `pytorch_score.py` for this tutorial. Our web service will use this file to predict whether an image is an ant or a bee. When writing your own scoring script, don't forget to test it locally first before you go and deploy the web service."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create environment file\n",
- "Then, we will need to create an environment file (`myenv.yml`) that specifies all of the scoring script's package dependencies. This file is used to ensure that all of those dependencies are installed in the Docker image by Azure ML. In this case, we need to specify `azureml-core`, `torch` and `torchvision`."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.conda_dependencies import CondaDependencies \n",
- "\n",
- "myenv = CondaDependencies.create(pip_packages=['azureml-defaults', 'torch', 'torchvision'])\n",
- "\n",
- "with open(\"myenv.yml\",\"w\") as f:\n",
- " f.write(myenv.serialize_to_string())\n",
- " \n",
- "print(myenv.serialize_to_string())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Configure the container image\n",
- "Now configure the Docker image that you will use to build your ACI container."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.image import ContainerImage\n",
- "\n",
- "image_config = ContainerImage.image_configuration(execution_script='pytorch_score.py', \n",
- " runtime='python', \n",
- " conda_file='myenv.yml',\n",
- " description='Image with hymenoptera model')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Configure the ACI container\n",
- "We are almost ready to deploy. Create a deployment configuration file to specify the number of CPUs and gigabytes of RAM needed for your ACI container. While it depends on your model, the default of `1` core and `1` gigabyte of RAM is usually sufficient for many models."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.webservice import AciWebservice\n",
- "\n",
- "aciconfig = AciWebservice.deploy_configuration(cpu_cores=1, \n",
- " memory_gb=1, \n",
- " tags={'data': 'hymenoptera', 'method':'transfer learning', 'framework':'pytorch'},\n",
- " description='Classify ants/bees using transfer learning with PyTorch')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Deploy the registered model\n",
- "Finally, let's deploy a web service from our registered model. Deploy the web service using the ACI config and image config files created in the previous steps. We pass the `model` object in a list to the `models` parameter. If you would like to deploy more than one registered model, append the additional models to this list."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%time\n",
- "from azureml.core.webservice import Webservice\n",
- "\n",
- "service_name = 'aci-hymenoptera'\n",
- "service = Webservice.deploy_from_model(workspace=ws,\n",
- " name=service_name,\n",
- " models=[model],\n",
- " image_config=image_config,\n",
- " deployment_config=aciconfig,)\n",
- "\n",
- "service.wait_for_deployment(show_output=True)\n",
- "print(service.state)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "If your deployment fails for any reason and you need to redeploy, make sure to delete the service before you do so: `service.delete()`"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "**Tip: If something goes wrong with the deployment, the first thing to look at is the logs from the service by running the following command:**"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "service.get_logs()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Get the web service's HTTP endpoint, which accepts REST client calls. This endpoint can be shared with anyone who wants to test the web service or integrate it into an application."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "print(service.scoring_uri)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Test the web service\n",
- "Finally, let's test our deployed web service. We will send the data as a JSON string to the web service hosted in ACI and use the SDK's `run` API to invoke the service. Here we will take an image from our validation data to predict on."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import os, json\n",
- "from PIL import Image\n",
- "import matplotlib.pyplot as plt\n",
- "\n",
- "plt.imshow(Image.open('test_img.jpg'))"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import torch\n",
- "from torchvision import transforms\n",
- " \n",
- "def preprocess(image_file):\n",
- " \"\"\"Preprocess the input image.\"\"\"\n",
- " data_transforms = transforms.Compose([\n",
- " transforms.Resize(256),\n",
- " transforms.CenterCrop(224),\n",
- " transforms.ToTensor(),\n",
- " transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])\n",
- " ])\n",
- "\n",
- " image = Image.open(image_file)\n",
- " image = data_transforms(image).float()\n",
- " image = torch.tensor(image)\n",
- " image = image.unsqueeze(0)\n",
- " return image.numpy()"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "input_data = preprocess('test_img.jpg')\n",
- "result = service.run(input_data=json.dumps({'data': input_data.tolist()}))\n",
- "print(result)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Clean up\n",
- "Once you no longer need the web service, you can delete it with a simple API call."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "service.delete()"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "minxia"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved. \n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Train, hyperparameter tune, and deploy with PyTorch\n",
+ "\n",
+ "In this tutorial, you will train, hyperparameter tune, and deploy a PyTorch model using the Azure Machine Learning (Azure ML) Python SDK.\n",
+ "\n",
+ "This tutorial will train an image classification model using transfer learning, based on PyTorch's [Transfer Learning tutorial](https://pytorch.org/tutorials/beginner/transfer_learning_tutorial.html). The model is trained to classify ants and bees by first using a pretrained ResNet18 model that has been trained on the [ImageNet](http://image-net.org/index) dataset."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Prerequisites\n",
+ "* Go through the [Configuration](../../../configuration.ipynb) notebook to install the Azure Machine Learning Python SDK and create an Azure ML `Workspace`"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Check core SDK version number\n",
+ "import azureml.core\n",
+ "\n",
+ "print(\"SDK version:\", azureml.core.VERSION)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Diagnostics\n",
+ "Opt-in diagnostics for better experience, quality, and security of future releases."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "Diagnostics"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "from azureml.telemetry import set_diagnostics_collection\n",
+ "\n",
+ "set_diagnostics_collection(send_diagnostics=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Initialize workspace\n",
+ "Initialize a [Workspace](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#workspace) object from the existing workspace you created in the Prerequisites step. `Workspace.from_config()` creates a workspace object from the details stored in `config.json`."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.workspace import Workspace\n",
+ "\n",
+ "ws = Workspace.from_config()\n",
+ "print('Workspace name: ' + ws.name, \n",
+ " 'Azure region: ' + ws.location, \n",
+ " 'Subscription id: ' + ws.subscription_id, \n",
+ " 'Resource group: ' + ws.resource_group, sep = '\\n')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Create or Attach existing AmlCompute\n",
+ "You will need to create a [compute target](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#compute-target) for training your model. In this tutorial, we use Azure ML managed compute ([AmlCompute](https://docs.microsoft.com/azure/machine-learning/service/how-to-set-up-training-targets#amlcompute)) for our remote training compute resource.\n",
+ "\n",
+ "**Creation of AmlCompute takes approximately 5 minutes.** If the AmlCompute with that name is already in your workspace, this code will skip the creation process.\n",
+ "\n",
+ "As with other Azure services, there are limits on certain resources (e.g. AmlCompute) associated with the Azure Machine Learning service. Please read [this article](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-manage-quotas) on the default limits and how to request more quota."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.compute import ComputeTarget, AmlCompute\n",
+ "from azureml.core.compute_target import ComputeTargetException\n",
+ "\n",
+ "# choose a name for your cluster\n",
+ "cluster_name = \"gpucluster\"\n",
+ "\n",
+ "try:\n",
+ " compute_target = ComputeTarget(workspace=ws, name=cluster_name)\n",
+ " print('Found existing compute target.')\n",
+ "except ComputeTargetException:\n",
+ " print('Creating a new compute target...')\n",
+ " compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_NC6', \n",
+ " max_nodes=4)\n",
+ "\n",
+ " # create the cluster\n",
+ " compute_target = ComputeTarget.create(ws, cluster_name, compute_config)\n",
+ "\n",
+ " compute_target.wait_for_completion(show_output=True)\n",
+ "\n",
+ "# use get_status() to get a detailed status for the current cluster. \n",
+ "print(compute_target.get_status().serialize())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "The above code creates a GPU cluster. If you instead want to create a CPU cluster, provide a different VM size to the `vm_size` parameter, such as `STANDARD_D2_V2`."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Train model on the remote compute\n",
+ "Now that you have your data and training script prepared, you are ready to train on your remote compute cluster. You can take advantage of Azure compute to leverage GPUs to cut down your training time. "
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create a project directory\n",
+ "Create a directory that will contain all the necessary code from your local machine that you will need access to on the remote resource. This includes the training script and any additional files your training script depends on."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import os\n",
+ "\n",
+ "project_folder = './pytorch-hymenoptera'\n",
+ "os.makedirs(project_folder, exist_ok=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Download training data\n",
+ "The dataset we will use (located [here](https://download.pytorch.org/tutorial/hymenoptera_data.zip) as a zip file) consists of about 120 training images each for ants and bees, with 75 validation images for each class. [Hymenoptera](https://en.wikipedia.org/wiki/Hymenoptera) is the order of insects that includes ants and bees. We will download and extract the dataset as part of our training script `pytorch_train.py`"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Prepare training script\n",
+ "Now you will need to create your training script. In this tutorial, the training script is already provided for you at `pytorch_train.py`. In practice, you should be able to take any custom training script as is and run it with Azure ML without having to modify your code.\n",
+ "\n",
+ "However, if you would like to use Azure ML's [tracking and metrics](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#metrics) capabilities, you will have to add a small amount of Azure ML code inside your training script. \n",
+ "\n",
+ "In `pytorch_train.py`, we will log some metrics to our Azure ML run. To do so, we will access the Azure ML `Run` object within the script:\n",
+ "```Python\n",
+ "from azureml.core.run import Run\n",
+ "run = Run.get_context()\n",
+ "```\n",
+ "Further within `pytorch_train.py`, we log the learning rate and momentum parameters, and the best validation accuracy the model achieves:\n",
+ "```Python\n",
+ "run.log('lr', np.float(learning_rate))\n",
+ "run.log('momentum', np.float(momentum))\n",
+ "\n",
+ "run.log('best_val_acc', np.float(best_acc))\n",
+ "```\n",
+ "These run metrics will become particularly important when we begin hyperparameter tuning our model in the \"Tune model hyperparameters\" section."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Once your script is ready, copy the training script `pytorch_train.py` into your project directory."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import shutil\n",
+ "\n",
+ "shutil.copy('pytorch_train.py', project_folder)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create an experiment\n",
+ "Create an [Experiment](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#experiment) to track all the runs in your workspace for this transfer learning PyTorch tutorial. "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core import Experiment\n",
+ "\n",
+ "experiment_name = 'pytorch-hymenoptera'\n",
+ "experiment = Experiment(ws, name=experiment_name)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create a PyTorch estimator\n",
+ "The Azure ML SDK's PyTorch estimator enables you to easily submit PyTorch training jobs for both single-node and distributed runs. For more information on the PyTorch estimator, refer [here](https://docs.microsoft.com/azure/machine-learning/service/how-to-train-pytorch). The following code will define a single-node PyTorch job."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.train.dnn import PyTorch\n",
+ "\n",
+ "script_params = {\n",
+ " '--num_epochs': 30,\n",
+ " '--output_dir': './outputs'\n",
+ "}\n",
+ "\n",
+ "estimator = PyTorch(source_directory=project_folder, \n",
+ " script_params=script_params,\n",
+ " compute_target=compute_target,\n",
+ " entry_script='pytorch_train.py',\n",
+ " use_gpu=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "The `script_params` parameter is a dictionary containing the command-line arguments to your training script `entry_script`. Please note the following:\n",
+ "- We passed our training data reference `ds_data` to our script's `--data_dir` argument. This will 1) mount our datastore on the remote compute and 2) provide the path to the training data `hymenoptera_data` on our datastore.\n",
+ "- We specified the output directory as `./outputs`. The `outputs` directory is specially treated by Azure ML in that all the content in this directory gets uploaded to your workspace as part of your run history. The files written to this directory are therefore accessible even once your remote run is over. In this tutorial, we will save our trained model to this output directory.\n",
+ "\n",
+ "To leverage the Azure VM's GPU for training, we set `use_gpu=True`."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Submit job\n",
+ "Run your experiment by submitting your estimator object. Note that this call is asynchronous."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run = experiment.submit(estimator)\n",
+ "print(run)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# to get more details of your run\n",
+ "print(run.get_details())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Monitor your run\n",
+ "You can monitor the progress of the run with a Jupyter widget. Like the run submission, the widget is asynchronous and provides live updates every 10-15 seconds until the job completes."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.widgets import RunDetails\n",
+ "\n",
+ "RunDetails(run).show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Alternatively, you can block until the script has completed training before running more code."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run.wait_for_completion(show_output=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Tune model hyperparameters\n",
+ "Now that we've seen how to do a simple PyTorch training run using the SDK, let's see if we can further improve the accuracy of our model. We can optimize our model's hyperparameters using Azure Machine Learning's hyperparameter tuning capabilities."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Start a hyperparameter sweep\n",
+ "First, we will define the hyperparameter space to sweep over. Since our training script uses a learning rate schedule to decay the learning rate every several epochs, let's tune the initial learning rate and the momentum parameters. In this example we will use random sampling to try different configuration sets of hyperparameters to maximize our primary metric, the best validation accuracy (`best_val_acc`).\n",
+ "\n",
+ "Then, we specify the early termination policy to use to early terminate poorly performing runs. Here we use the `BanditPolicy`, which will terminate any run that doesn't fall within the slack factor of our primary evaluation metric. In this tutorial, we will apply this policy every epoch (since we report our `best_val_acc` metric every epoch and `evaluation_interval=1`). Notice we will delay the first policy evaluation until after the first `10` epochs (`delay_evaluation=10`).\n",
+ "Refer [here](https://docs.microsoft.com/azure/machine-learning/service/how-to-tune-hyperparameters#specify-an-early-termination-policy) for more information on the BanditPolicy and other policies available."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.train.hyperdrive import *\n",
+ "\n",
+ "param_sampling = RandomParameterSampling( {\n",
+ " 'learning_rate': uniform(0.0005, 0.005),\n",
+ " 'momentum': uniform(0.9, 0.99)\n",
+ " }\n",
+ ")\n",
+ "\n",
+ "early_termination_policy = BanditPolicy(slack_factor=0.15, evaluation_interval=1, delay_evaluation=10)\n",
+ "\n",
+ "hyperdrive_run_config = HyperDriveRunConfig(estimator=estimator,\n",
+ " hyperparameter_sampling=param_sampling, \n",
+ " policy=early_termination_policy,\n",
+ " primary_metric_name='best_val_acc',\n",
+ " primary_metric_goal=PrimaryMetricGoal.MAXIMIZE,\n",
+ " max_total_runs=8,\n",
+ " max_concurrent_runs=4)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Finally, lauch the hyperparameter tuning job."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# start the HyperDrive run\n",
+ "hyperdrive_run = experiment.submit(hyperdrive_run_config)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Monitor HyperDrive runs\n",
+ "You can monitor the progress of the runs with the following Jupyter widget. "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.widgets import RunDetails\n",
+ "\n",
+ "RunDetails(hyperdrive_run).show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Or block until the HyperDrive sweep has completed:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "hyperdrive_run.wait_for_completion(show_output=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Find and register the best model\n",
+ "Once all the runs complete, we can find the run that produced the model with the highest accuracy."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "best_run = hyperdrive_run.get_best_run_by_primary_metric()\n",
+ "best_run_metrics = best_run.get_metrics()\n",
+ "print(best_run)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "print('Best Run is:\\n Validation accuracy: {0:.5f} \\n Learning rate: {1:.5f} \\n Momentum: {2:.5f}'.format(\n",
+ " best_run_metrics['best_val_acc'][-1],\n",
+ " best_run_metrics['lr'],\n",
+ " best_run_metrics['momentum'])\n",
+ " )"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Finally, register the model from your best-performing run to your workspace. The `model_path` parameter takes in the relative path on the remote VM to the model file in your `outputs` directory. In the next section, we will deploy this registered model as a web service."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "model = best_run.register_model(model_name = 'pytorch-hymenoptera', model_path = 'outputs/model.pt')\n",
+ "print(model.name, model.id, model.version, sep = '\\t')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Deploy model as web service\n",
+ "Once you have your trained model, you can deploy the model on Azure. In this tutorial, we will deploy the model as a web service in [Azure Container Instances](https://docs.microsoft.com/en-us/azure/container-instances/) (ACI). For more information on deploying models using Azure ML, refer [here](https://docs.microsoft.com/azure/machine-learning/service/how-to-deploy-and-where)."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create scoring script\n",
+ "\n",
+ "First, we will create a scoring script that will be invoked by the web service call. Note that the scoring script must have two required functions:\n",
+ "* `init()`: In this function, you typically load the model into a `global` object. This function is executed only once when the Docker container is started. \n",
+ "* `run(input_data)`: In this function, the model is used to predict a value based on the input data. The input and output typically use JSON as serialization and deserialization format, but you are not limited to that.\n",
+ "\n",
+ "Refer to the scoring script `pytorch_score.py` for this tutorial. Our web service will use this file to predict whether an image is an ant or a bee. When writing your own scoring script, don't forget to test it locally first before you go and deploy the web service."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create environment file\n",
+ "Then, we will need to create an environment file (`myenv.yml`) that specifies all of the scoring script's package dependencies. This file is used to ensure that all of those dependencies are installed in the Docker image by Azure ML. In this case, we need to specify `azureml-core`, `torch` and `torchvision`."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.conda_dependencies import CondaDependencies \n",
+ "\n",
+ "myenv = CondaDependencies.create(pip_packages=['azureml-defaults', 'torch', 'torchvision'])\n",
+ "\n",
+ "with open(\"myenv.yml\",\"w\") as f:\n",
+ " f.write(myenv.serialize_to_string())\n",
+ " \n",
+ "print(myenv.serialize_to_string())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Configure the container image\n",
+ "Now configure the Docker image that you will use to build your ACI container."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.image import ContainerImage\n",
+ "\n",
+ "image_config = ContainerImage.image_configuration(execution_script='pytorch_score.py', \n",
+ " runtime='python', \n",
+ " conda_file='myenv.yml',\n",
+ " description='Image with hymenoptera model')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Configure the ACI container\n",
+ "We are almost ready to deploy. Create a deployment configuration file to specify the number of CPUs and gigabytes of RAM needed for your ACI container. While it depends on your model, the default of `1` core and `1` gigabyte of RAM is usually sufficient for many models."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.webservice import AciWebservice\n",
+ "\n",
+ "aciconfig = AciWebservice.deploy_configuration(cpu_cores=1, \n",
+ " memory_gb=1, \n",
+ " tags={'data': 'hymenoptera', 'method':'transfer learning', 'framework':'pytorch'},\n",
+ " description='Classify ants/bees using transfer learning with PyTorch')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Deploy the registered model\n",
+ "Finally, let's deploy a web service from our registered model. Deploy the web service using the ACI config and image config files created in the previous steps. We pass the `model` object in a list to the `models` parameter. If you would like to deploy more than one registered model, append the additional models to this list."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%time\n",
+ "from azureml.core.webservice import Webservice\n",
+ "\n",
+ "service_name = 'aci-hymenoptera'\n",
+ "service = Webservice.deploy_from_model(workspace=ws,\n",
+ " name=service_name,\n",
+ " models=[model],\n",
+ " image_config=image_config,\n",
+ " deployment_config=aciconfig,)\n",
+ "\n",
+ "service.wait_for_deployment(show_output=True)\n",
+ "print(service.state)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "If your deployment fails for any reason and you need to redeploy, make sure to delete the service before you do so: `service.delete()`"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "**Tip: If something goes wrong with the deployment, the first thing to look at is the logs from the service by running the following command:**"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "service.get_logs()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Get the web service's HTTP endpoint, which accepts REST client calls. This endpoint can be shared with anyone who wants to test the web service or integrate it into an application."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "print(service.scoring_uri)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Test the web service\n",
+ "Finally, let's test our deployed web service. We will send the data as a JSON string to the web service hosted in ACI and use the SDK's `run` API to invoke the service. Here we will take an image from our validation data to predict on."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import os, json\n",
+ "from PIL import Image\n",
+ "import matplotlib.pyplot as plt\n",
+ "\n",
+ "plt.imshow(Image.open('test_img.jpg'))"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import torch\n",
+ "from torchvision import transforms\n",
+ " \n",
+ "def preprocess(image_file):\n",
+ " \"\"\"Preprocess the input image.\"\"\"\n",
+ " data_transforms = transforms.Compose([\n",
+ " transforms.Resize(256),\n",
+ " transforms.CenterCrop(224),\n",
+ " transforms.ToTensor(),\n",
+ " transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])\n",
+ " ])\n",
+ "\n",
+ " image = Image.open(image_file)\n",
+ " image = data_transforms(image).float()\n",
+ " image = torch.tensor(image)\n",
+ " image = image.unsqueeze(0)\n",
+ " return image.numpy()"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "input_data = preprocess('test_img.jpg')\n",
+ "result = service.run(input_data=json.dumps({'data': input_data.tolist()}))\n",
+ "print(result)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Clean up\n",
+ "Once you no longer need the web service, you can delete it with a simple API call."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "service.delete()"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "minxia"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.6"
+ },
+ "msauthor": "minxia"
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- },
- "msauthor": "minxia"
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/training-with-deep-learning/train-hyperparameter-tune-deploy-with-tensorflow/train-hyperparameter-tune-deploy-with-tensorflow.ipynb b/how-to-use-azureml/training-with-deep-learning/train-hyperparameter-tune-deploy-with-tensorflow/train-hyperparameter-tune-deploy-with-tensorflow.ipynb
index b986ac4b..816d0a09 100644
--- a/how-to-use-azureml/training-with-deep-learning/train-hyperparameter-tune-deploy-with-tensorflow/train-hyperparameter-tune-deploy-with-tensorflow.ipynb
+++ b/how-to-use-azureml/training-with-deep-learning/train-hyperparameter-tune-deploy-with-tensorflow/train-hyperparameter-tune-deploy-with-tensorflow.ipynb
@@ -1,1172 +1,1172 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {
- "nbpresent": {
- "id": "bf74d2e9-2708-49b1-934b-e0ede342f475"
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "nbpresent": {
+ "id": "bf74d2e9-2708-49b1-934b-e0ede342f475"
+ }
+ },
+ "source": [
+ "# Training, hyperparameter tune, and deploy with TensorFlow\n",
+ "\n",
+ "## Introduction\n",
+ "This tutorial shows how to train a simple deep neural network using the MNIST dataset and TensorFlow on Azure Machine Learning. MNIST is a popular dataset consisting of 70,000 grayscale images. Each image is a handwritten digit of `28x28` pixels, representing number from 0 to 9. The goal is to create a multi-class classifier to identify the digit each image represents, and deploy it as a web service in Azure.\n",
+ "\n",
+ "For more information about the MNIST dataset, please visit [Yan LeCun's website](http://yann.lecun.com/exdb/mnist/).\n",
+ "\n",
+ "## Prerequisite:\n",
+ "* Understand the [architecture and terms](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture) introduced by Azure Machine Learning\n",
+ "* Go through the [configuration notebook](../../../configuration.ipynb) to:\n",
+ " * install the AML SDK\n",
+ " * create a workspace and its configuration file (`config.json`)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Let's get started. First let's import some Python libraries."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "nbpresent": {
+ "id": "c377ea0c-0cd9-4345-9be2-e20fb29c94c3"
+ }
+ },
+ "outputs": [],
+ "source": [
+ "%matplotlib inline\n",
+ "import numpy as np\n",
+ "import os\n",
+ "import matplotlib\n",
+ "import matplotlib.pyplot as plt"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "nbpresent": {
+ "id": "edaa7f2f-2439-4148-b57a-8c794c0945ec"
+ }
+ },
+ "outputs": [],
+ "source": [
+ "import azureml\n",
+ "from azureml.core import Workspace, Run\n",
+ "\n",
+ "# check core SDK version number\n",
+ "print(\"Azure ML SDK Version: \", azureml.core.VERSION)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Diagnostics\n",
+ "Opt-in diagnostics for better experience, quality, and security of future releases."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "Diagnostics"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "from azureml.telemetry import set_diagnostics_collection\n",
+ "\n",
+ "set_diagnostics_collection(send_diagnostics=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Initialize workspace\n",
+ "Initialize a [Workspace](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#workspace) object from the existing workspace you created in the Prerequisites step. `Workspace.from_config()` creates a workspace object from the details stored in `config.json`."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.workspace import Workspace\n",
+ "\n",
+ "ws = Workspace.from_config()\n",
+ "print('Workspace name: ' + ws.name, \n",
+ " 'Azure region: ' + ws.location, \n",
+ " 'Subscription id: ' + ws.subscription_id, \n",
+ " 'Resource group: ' + ws.resource_group, sep = '\\n')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "nbpresent": {
+ "id": "59f52294-4a25-4c92-bab8-3b07f0f44d15"
+ }
+ },
+ "source": [
+ "## Create an Azure ML experiment\n",
+ "Let's create an experiment named \"tf-mnist\" and a folder to hold the training scripts. The script runs will be recorded under the experiment in Azure."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "nbpresent": {
+ "id": "bc70f780-c240-4779-96f3-bc5ef9a37d59"
+ }
+ },
+ "outputs": [],
+ "source": [
+ "from azureml.core import Experiment\n",
+ "\n",
+ "script_folder = './tf-mnist'\n",
+ "os.makedirs(script_folder, exist_ok=True)\n",
+ "\n",
+ "exp = Experiment(workspace=ws, name='tf-mnist')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "nbpresent": {
+ "id": "defe921f-8097-44c3-8336-8af6700804a7"
+ }
+ },
+ "source": [
+ "## Download MNIST dataset\n",
+ "In order to train on the MNIST dataset we will first need to download it from Yan LeCun's web site directly and save them in a `data` folder locally."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import os\n",
+ "import urllib\n",
+ "\n",
+ "os.makedirs('./data/mnist', exist_ok=True)\n",
+ "\n",
+ "urllib.request.urlretrieve('http://yann.lecun.com/exdb/mnist/train-images-idx3-ubyte.gz', filename = './data/mnist/train-images.gz')\n",
+ "urllib.request.urlretrieve('http://yann.lecun.com/exdb/mnist/train-labels-idx1-ubyte.gz', filename = './data/mnist/train-labels.gz')\n",
+ "urllib.request.urlretrieve('http://yann.lecun.com/exdb/mnist/t10k-images-idx3-ubyte.gz', filename = './data/mnist/test-images.gz')\n",
+ "urllib.request.urlretrieve('http://yann.lecun.com/exdb/mnist/t10k-labels-idx1-ubyte.gz', filename = './data/mnist/test-labels.gz')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "nbpresent": {
+ "id": "c3f2f57c-7454-4d3e-b38d-b0946cf066ea"
+ }
+ },
+ "source": [
+ "## Show some sample images\n",
+ "Let's load the downloaded compressed file into numpy arrays using some utility functions included in the `utils.py` library file from the current folder. Then we use `matplotlib` to plot 30 random images from the dataset along with their labels."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "nbpresent": {
+ "id": "396d478b-34aa-4afa-9898-cdce8222a516"
+ }
+ },
+ "outputs": [],
+ "source": [
+ "from utils import load_data\n",
+ "\n",
+ "# note we also shrink the intensity values (X) from 0-255 to 0-1. This helps the neural network converge faster.\n",
+ "X_train = load_data('./data/mnist/train-images.gz', False) / 255.0\n",
+ "y_train = load_data('./data/mnist/train-labels.gz', True).reshape(-1)\n",
+ "\n",
+ "X_test = load_data('./data/mnist/test-images.gz', False) / 255.0\n",
+ "y_test = load_data('./data/mnist/test-labels.gz', True).reshape(-1)\n",
+ "\n",
+ "count = 0\n",
+ "sample_size = 30\n",
+ "plt.figure(figsize = (16, 6))\n",
+ "for i in np.random.permutation(X_train.shape[0])[:sample_size]:\n",
+ " count = count + 1\n",
+ " plt.subplot(1, sample_size, count)\n",
+ " plt.axhline('')\n",
+ " plt.axvline('')\n",
+ " plt.text(x = 10, y = -10, s = y_train[i], fontsize = 18)\n",
+ " plt.imshow(X_train[i].reshape(28, 28), cmap = plt.cm.Greys)\n",
+ "plt.show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Upload MNIST dataset to default datastore \n",
+ "A [datastore](https://docs.microsoft.com/azure/machine-learning/service/how-to-access-data) is a place where data can be stored that is then made accessible to a Run either by means of mounting or copying the data to the compute target. A datastore can either be backed by an Azure Blob Storage or and Azure File Share (ADLS will be supported in the future). For simple data handling, each workspace provides a default datastore that can be used, in case the data is not already in Blob Storage or File Share."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "ds = ws.get_default_datastore()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "In this next step, we will upload the training and test set into the workspace's default datastore, which we will then later be mount on an `AmlCompute` cluster for training."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "ds.upload(src_dir='./data/mnist', target_path='mnist', overwrite=True, show_progress=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Create or Attach existing AmlCompute\n",
+ "You will need to create a [compute target](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#compute-target) for training your model. In this tutorial, you create `AmlCompute` as your training compute resource."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "If we could not find the cluster with the given name, then we will create a new cluster here. We will create an `AmlCompute` cluster of `STANDARD_NC6` GPU VMs. This process is broken down into 3 steps:\n",
+ "1. create the configuration (this step is local and only takes a second)\n",
+ "2. create the cluster (this step will take about **20 seconds**)\n",
+ "3. provision the VMs to bring the cluster to the initial size (of 1 in this case). This step will take about **3-5 minutes** and is providing only sparse output in the process. Please make sure to wait until the call returns before moving to the next cell"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.compute import ComputeTarget, AmlCompute\n",
+ "from azureml.core.compute_target import ComputeTargetException\n",
+ "\n",
+ "# choose a name for your cluster\n",
+ "cluster_name = \"gpucluster\"\n",
+ "\n",
+ "try:\n",
+ " compute_target = ComputeTarget(workspace=ws, name=cluster_name)\n",
+ " print('Found existing compute target')\n",
+ "except ComputeTargetException:\n",
+ " print('Creating a new compute target...')\n",
+ " compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_NC6', \n",
+ " max_nodes=4)\n",
+ "\n",
+ " # create the cluster\n",
+ " compute_target = ComputeTarget.create(ws, cluster_name, compute_config)\n",
+ "\n",
+ " # can poll for a minimum number of nodes and for a specific timeout. \n",
+ " # if no min node count is provided it uses the scale settings for the cluster\n",
+ " compute_target.wait_for_completion(show_output=True, min_node_count=None, timeout_in_minutes=20)\n",
+ "\n",
+ "# use get_status() to get a detailed status for the current cluster. \n",
+ "print(compute_target.get_status().serialize())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Now that you have created the compute target, let's see what the workspace's `compute_targets` property returns. You should now see one entry named 'gpucluster' of type `AmlCompute`."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "compute_targets = ws.compute_targets\n",
+ "for name, ct in compute_targets.items():\n",
+ " print(name, ct.type, ct.provisioning_state)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Copy the training files into the script folder\n",
+ "The TensorFlow training script is already created for you. You can simply copy it into the script folder, together with the utility library used to load compressed data file into numpy array."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import shutil\n",
+ "\n",
+ "# the training logic is in the tf_mnist.py file.\n",
+ "shutil.copy('./tf_mnist.py', script_folder)\n",
+ "\n",
+ "# the utils.py just helps loading data from the downloaded MNIST dataset into numpy arrays.\n",
+ "shutil.copy('./utils.py', script_folder)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {
+ "nbpresent": {
+ "id": "2039d2d5-aca6-4f25-a12f-df9ae6529cae"
+ }
+ },
+ "source": [
+ "## Construct neural network in TensorFlow\n",
+ "In the training script `tf_mnist.py`, it creates a very simple DNN (deep neural network), with just 2 hidden layers. The input layer has 28 * 28 = 784 neurons, each representing a pixel in an image. The first hidden layer has 300 neurons, and the second hidden layer has 100 neurons. The output layer has 10 neurons, each representing a targeted label from 0 to 9.\n",
+ "\n",
+ ""
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Azure ML concepts \n",
+ "Please note the following three things in the code below:\n",
+ "1. The script accepts arguments using the argparse package. In this case there is one argument `--data_folder` which specifies the file system folder in which the script can find the MNIST data\n",
+ "```\n",
+ " parser = argparse.ArgumentParser()\n",
+ " parser.add_argument('--data_folder')\n",
+ "```\n",
+ "2. The script is accessing the Azure ML `Run` object by executing `run = Run.get_context()`. Further down the script is using the `run` to report the training accuracy and the validation accuracy as training progresses.\n",
+ "```\n",
+ " run.log('training_acc', np.float(acc_train))\n",
+ " run.log('validation_acc', np.float(acc_val))\n",
+ "```\n",
+ "3. When running the script on Azure ML, you can write files out to a folder `./outputs` that is relative to the root directory. This folder is specially tracked by Azure ML in the sense that any files written to that folder during script execution on the remote target will be picked up by Run History; these files (known as artifacts) will be available as part of the run history record."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "The next cell will print out the training code for you to inspect it."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "with open(os.path.join(script_folder, './tf_mnist.py'), 'r') as f:\n",
+ " print(f.read())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Create TensorFlow estimator\n",
+ "Next, we construct an `azureml.train.dnn.TensorFlow` estimator object, use the Batch AI cluster as compute target, and pass the mount-point of the datastore to the training code as a parameter.\n",
+ "The TensorFlow estimator is providing a simple way of launching a TensorFlow training job on a compute target. It will automatically provide a docker image that has TensorFlow installed -- if additional pip or conda packages are required, their names can be passed in via the `pip_packages` and `conda_packages` arguments and they will be included in the resulting docker."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.train.dnn import TensorFlow\n",
+ "\n",
+ "script_params = {\n",
+ " '--data-folder': ws.get_default_datastore().as_mount(),\n",
+ " '--batch-size': 50,\n",
+ " '--first-layer-neurons': 300,\n",
+ " '--second-layer-neurons': 100,\n",
+ " '--learning-rate': 0.01\n",
+ "}\n",
+ "\n",
+ "est = TensorFlow(source_directory=script_folder,\n",
+ " script_params=script_params,\n",
+ " compute_target=compute_target,\n",
+ " entry_script='tf_mnist.py', \n",
+ " use_gpu=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Submit job to run\n",
+ "Calling the `fit` function on the estimator submits the job to Azure ML for execution. Submitting the job should only take a few seconds."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run = exp.submit(est)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Monitor the Run\n",
+ "As the Run is executed, it will go through the following stages:\n",
+ "1. Preparing: A docker image is created matching the Python environment specified by the TensorFlow estimator and it will be uploaded to the workspace's Azure Container Registry. This step will only happen once for each Python environment -- the container will then be cached for subsequent runs. Creating and uploading the image takes about **5 minutes**. While the job is preparing, logs are streamed to the run history and can be viewed to monitor the progress of the image creation.\n",
+ "\n",
+ "2. Scaling: If the compute needs to be scaled up (i.e. the Batch AI cluster requires more nodes to execute the run than currently available), the cluster will attempt to scale up in order to make the required amount of nodes available. Scaling typically takes about **5 minutes**.\n",
+ "\n",
+ "3. Running: All scripts in the script folder are uploaded to the compute target, data stores are mounted/copied and the `entry_script` is executed. While the job is running, stdout and the `./logs` folder are streamed to the run history and can be viewed to monitor the progress of the run.\n",
+ "\n",
+ "4. Post-Processing: The `./outputs` folder of the run is copied over to the run history\n",
+ "\n",
+ "There are multiple ways to check the progress of a running job. We can use a Jupyter notebook widget. \n",
+ "\n",
+ "**Note: The widget will automatically update ever 10-15 seconds, always showing you the most up-to-date information about the run**"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.widgets import RunDetails\n",
+ "\n",
+ "RunDetails(run).show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "We can also periodically check the status of the run object, and navigate to Azure portal to monitor the run."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run.wait_for_completion(show_output=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### The Run object\n",
+ "The Run object provides the interface to the run history -- both to the job and to the control plane (this notebook), and both while the job is running and after it has completed. It provides a number of interesting features for instance:\n",
+ "* `run.get_details()`: Provides a rich set of properties of the run\n",
+ "* `run.get_metrics()`: Provides a dictionary with all the metrics that were reported for the Run\n",
+ "* `run.get_file_names()`: List all the files that were uploaded to the run history for this Run. This will include the `outputs` and `logs` folder, azureml-logs and other logs, as well as files that were explicitly uploaded to the run using `run.upload_file()`\n",
+ "\n",
+ "Below are some examples -- please run through them and inspect their output. "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run.get_details()"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run.get_metrics()"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run.get_file_names()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Plot accuracy over epochs\n",
+ "Since we can retrieve the metrics from the run, we can easily make plots using `matplotlib` in the notebook. Then we can add the plotted image to the run using `run.log_image()`, so all information about the run is kept together."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import os\n",
+ "\n",
+ "os.makedirs('./imgs', exist_ok=True)\n",
+ "metrics = run.get_metrics()\n",
+ "\n",
+ "plt.figure(figsize = (13,5))\n",
+ "plt.plot(metrics['validation_acc'], 'r-', lw=4, alpha=.6)\n",
+ "plt.plot(metrics['training_acc'], 'b--', alpha=0.5)\n",
+ "plt.legend(['Full evaluation set', 'Training set mini-batch'])\n",
+ "plt.xlabel('epochs', fontsize=14)\n",
+ "plt.ylabel('accuracy', fontsize=14)\n",
+ "plt.title('Accuracy over Epochs', fontsize=16)\n",
+ "run.log_image(name='acc_over_epochs.png', plot=plt)\n",
+ "plt.show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Download the saved model"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "In the training script, a TensorFlow `saver` object is used to persist the model in a local folder (local to the compute target). The model was saved to the `./outputs` folder on the disk of the Batch AI cluster node where the job is run. Azure ML automatically uploaded anything written in the `./outputs` folder into run history file store. Subsequently, we can use the `Run` object to download the model files the `saver` object saved. They are under the the `outputs/model` folder in the run history file store, and are downloaded into a local folder named `model`. Note the TensorFlow model consists of four files in binary format and they are not human-readable."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# create a model folder in the current directory\n",
+ "os.makedirs('./model', exist_ok=True)\n",
+ "\n",
+ "for f in run.get_file_names():\n",
+ " if f.startswith('outputs/model'):\n",
+ " output_file_path = os.path.join('./model', f.split('/')[-1])\n",
+ " print('Downloading from {} to {} ...'.format(f, output_file_path))\n",
+ " run.download_file(name=f, output_file_path=output_file_path)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Predict on the test set\n",
+ "Now load the saved TensorFlow graph, and list all operations under the `network` scope. This way we can discover the input tensor `network/X:0` and the output tensor `network/output/MatMul:0`, and use them in the scoring script in the next step.\n",
+ "\n",
+ "Note: if your local TensorFlow version is different than the version running in the cluster where the model is trained, you might see a \"compiletime version mismatch\" warning. You can ignore it."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import tensorflow as tf\n",
+ "\n",
+ "tf.reset_default_graph()\n",
+ "\n",
+ "saver = tf.train.import_meta_graph(\"./model/mnist-tf.model.meta\")\n",
+ "graph = tf.get_default_graph()\n",
+ "\n",
+ "for op in graph.get_operations():\n",
+ " if op.name.startswith('network'):\n",
+ " print(op.name)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Feed test dataset to the persisted model to get predictions."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# input tensor. this is an array of 784 elements, each representing the intensity of a pixel in the digit image.\n",
+ "X = tf.get_default_graph().get_tensor_by_name(\"network/X:0\")\n",
+ "# output tensor. this is an array of 10 elements, each representing the probability of predicted value of the digit.\n",
+ "output = tf.get_default_graph().get_tensor_by_name(\"network/output/MatMul:0\")\n",
+ "\n",
+ "with tf.Session() as sess:\n",
+ " saver.restore(sess, './model/mnist-tf.model')\n",
+ " k = output.eval(feed_dict={X : X_test})\n",
+ "# get the prediction, which is the index of the element that has the largest probability value.\n",
+ "y_hat = np.argmax(k, axis=1)\n",
+ "\n",
+ "# print the first 30 labels and predictions\n",
+ "print('labels: \\t', y_test[:30])\n",
+ "print('predictions:\\t', y_hat[:30])"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Calculate the overall accuracy by comparing the predicted value against the test set."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "print(\"Accuracy on the test set:\", np.average(y_hat == y_test))"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Intelligent hyperparameter tuning\n",
+ "We have trained the model with one set of hyperparameters, now let's how we can do hyperparameter tuning by launching multiple runs on the cluster. First let's define the parameter space using random sampling."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.train.hyperdrive import *\n",
+ "\n",
+ "ps = RandomParameterSampling(\n",
+ " {\n",
+ " '--batch-size': choice(25, 50, 100),\n",
+ " '--first-layer-neurons': choice(10, 50, 200, 300, 500),\n",
+ " '--second-layer-neurons': choice(10, 50, 200, 500),\n",
+ " '--learning-rate': loguniform(-6, -1)\n",
+ " }\n",
+ ")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Next, we will create a new estimator without the above parameters since they will be passed in later. Note we still need to keep the `data-folder` parameter since that's not a hyperparamter we will sweep."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "est = TensorFlow(source_directory=script_folder,\n",
+ " script_params={'--data-folder': ws.get_default_datastore().as_mount()},\n",
+ " compute_target=compute_target,\n",
+ " entry_script='tf_mnist.py', \n",
+ " use_gpu=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Now we will define an early termnination policy. The `BanditPolicy` basically states to check the job every 2 iterations. If the primary metric (defined later) falls outside of the top 10% range, Azure ML terminate the job. This saves us from continuing to explore hyperparameters that don't show promise of helping reach our target metric."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "policy = BanditPolicy(evaluation_interval=2, slack_factor=0.1)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Now we are ready to configure a run configuration object, and specify the primary metric `validation_acc` that's recorded in your training runs. If you go back to visit the training script, you will notice that this value is being logged after every epoch (a full batch set). We also want to tell the service that we are looking to maximizing this value. We also set the number of samples to 20, and maximal concurrent job to 4, which is the same as the number of nodes in our computer cluster."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "htc = HyperDriveRunConfig(estimator=est, \n",
+ " hyperparameter_sampling=ps, \n",
+ " policy=policy, \n",
+ " primary_metric_name='validation_acc', \n",
+ " primary_metric_goal=PrimaryMetricGoal.MAXIMIZE, \n",
+ " max_total_runs=8,\n",
+ " max_concurrent_runs=4)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Finally, let's launch the hyperparameter tuning job."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "htr = exp.submit(config=htc)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "We can use a run history widget to show the progress. Be patient as this might take a while to complete."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "RunDetails(htr).show()"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "htr.wait_for_completion(show_output=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Find and register best model\n",
+ "When all the jobs finish, we can find out the one that has the highest accuracy."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "best_run = htr.get_best_run_by_primary_metric()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Now let's list the model files uploaded during the run."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "print(best_run.get_file_names())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "We can then register the folder (and all files in it) as a model named `tf-dnn-mnist` under the workspace for deployment."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "model = best_run.register_model(model_name='tf-dnn-mnist', model_path='outputs/model')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Deploy the model in ACI\n",
+ "Now we are ready to deploy the model as a web service running in Azure Container Instance [ACI](https://azure.microsoft.com/en-us/services/container-instances/). Azure Machine Learning accomplishes this by constructing a Docker image with the scoring logic and model baked in.\n",
+ "### Create score.py\n",
+ "First, we will create a scoring script that will be invoked by the web service call. \n",
+ "\n",
+ "* Note that the scoring script must have two required functions, `init()` and `run(input_data)`. \n",
+ " * In `init()` function, you typically load the model into a global object. This function is executed only once when the Docker container is started. \n",
+ " * In `run(input_data)` function, the model is used to predict a value based on the input data. The input and output to `run` typically use JSON as serialization and de-serialization format but you are not limited to that."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%writefile score.py\n",
+ "import json\n",
+ "import numpy as np\n",
+ "import os\n",
+ "import tensorflow as tf\n",
+ "\n",
+ "from azureml.core.model import Model\n",
+ "\n",
+ "def init():\n",
+ " global X, output, sess\n",
+ " tf.reset_default_graph()\n",
+ " model_root = Model.get_model_path('tf-dnn-mnist')\n",
+ " saver = tf.train.import_meta_graph(os.path.join(model_root, 'mnist-tf.model.meta'))\n",
+ " X = tf.get_default_graph().get_tensor_by_name(\"network/X:0\")\n",
+ " output = tf.get_default_graph().get_tensor_by_name(\"network/output/MatMul:0\")\n",
+ " \n",
+ " sess = tf.Session()\n",
+ " saver.restore(sess, os.path.join(model_root, 'mnist-tf.model'))\n",
+ "\n",
+ "def run(raw_data):\n",
+ " data = np.array(json.loads(raw_data)['data'])\n",
+ " # make prediction\n",
+ " out = output.eval(session=sess, feed_dict={X: data})\n",
+ " y_hat = np.argmax(out, axis=1)\n",
+ " return y_hat.tolist()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create myenv.yml\n",
+ "We also need to create an environment file so that Azure Machine Learning can install the necessary packages in the Docker image which are required by your scoring script. In this case, we need to specify packages `numpy`, `tensorflow`."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.runconfig import CondaDependencies\n",
+ "\n",
+ "cd = CondaDependencies.create()\n",
+ "cd.add_conda_package('numpy')\n",
+ "cd.add_tensorflow_conda_package()\n",
+ "cd.save_to_file(base_directory='./', conda_file_path='myenv.yml')\n",
+ "\n",
+ "print(cd.serialize_to_string())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Deploy to ACI\n",
+ "We are almost ready to deploy. Create a deployment configuration and specify the number of CPUs and gigbyte of RAM needed for your ACI container. "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.webservice import AciWebservice\n",
+ "\n",
+ "aciconfig = AciWebservice.deploy_configuration(cpu_cores=1, \n",
+ " memory_gb=1, \n",
+ " tags={'name':'mnist', 'framework': 'TensorFlow DNN'},\n",
+ " description='Tensorflow DNN on MNIST')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Deployment Process\n",
+ "Now we can deploy. **This cell will run for about 7-8 minutes**. Behind the scene, it will do the following:\n",
+ "1. **Register model** \n",
+ "Take the local `model` folder (which contains our previously downloaded trained model files) and register it (and the files inside that folder) as a model named `model` under the workspace. Azure ML will register the model directory or model file(s) we specify to the `model_paths` parameter of the `Webservice.deploy` call.\n",
+ "2. **Build Docker image** \n",
+ "Build a Docker image using the scoring file (`score.py`), the environment file (`myenv.yml`), and the `model` folder containing the TensorFlow model files. \n",
+ "3. **Register image** \n",
+ "Register that image under the workspace. \n",
+ "4. **Ship to ACI** \n",
+ "And finally ship the image to the ACI infrastructure, start up a container in ACI using that image, and expose an HTTP endpoint to accept REST client calls."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.image import ContainerImage\n",
+ "\n",
+ "imgconfig = ContainerImage.image_configuration(execution_script=\"score.py\", \n",
+ " runtime=\"python\", \n",
+ " conda_file=\"myenv.yml\")"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%time\n",
+ "from azureml.core.webservice import Webservice\n",
+ "\n",
+ "service = Webservice.deploy_from_model(workspace=ws,\n",
+ " name='tf-mnist-svc',\n",
+ " deployment_config=aciconfig,\n",
+ " models=[model],\n",
+ " image_config=imgconfig)\n",
+ "\n",
+ "service.wait_for_deployment(show_output=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "**Tip: If something goes wrong with the deployment, the first thing to look at is the logs from the service by running the following command:**"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "print(service.get_logs())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "This is the scoring web service endpoint:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "print(service.scoring_uri)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Test the deployed model\n",
+ "Let's test the deployed model. Pick 30 random samples from the test set, and send it to the web service hosted in ACI. Note here we are using the `run` API in the SDK to invoke the service. You can also make raw HTTP calls using any HTTP tool such as curl.\n",
+ "\n",
+ "After the invocation, we print the returned predictions and plot them along with the input images. Use red font color and inversed image (white on black) to highlight the misclassified samples. Note since the model accuracy is pretty high, you might have to run the below cell a few times before you can see a misclassified sample."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import json\n",
+ "\n",
+ "# find 30 random samples from test set\n",
+ "n = 30\n",
+ "sample_indices = np.random.permutation(X_test.shape[0])[0:n]\n",
+ "\n",
+ "test_samples = json.dumps({\"data\": X_test[sample_indices].tolist()})\n",
+ "test_samples = bytes(test_samples, encoding='utf8')\n",
+ "\n",
+ "# predict using the deployed model\n",
+ "result = service.run(input_data=test_samples)\n",
+ "\n",
+ "# compare actual value vs. the predicted values:\n",
+ "i = 0\n",
+ "plt.figure(figsize = (20, 1))\n",
+ "\n",
+ "for s in sample_indices:\n",
+ " plt.subplot(1, n, i + 1)\n",
+ " plt.axhline('')\n",
+ " plt.axvline('')\n",
+ " \n",
+ " # use different color for misclassified sample\n",
+ " font_color = 'red' if y_test[s] != result[i] else 'black'\n",
+ " clr_map = plt.cm.gray if y_test[s] != result[i] else plt.cm.Greys\n",
+ " \n",
+ " plt.text(x=10, y=-10, s=y_hat[s], fontsize=18, color=font_color)\n",
+ " plt.imshow(X_test[s].reshape(28, 28), cmap=clr_map)\n",
+ " \n",
+ " i = i + 1\n",
+ "plt.show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "We can also send raw HTTP request to the service."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import requests\n",
+ "import json\n",
+ "\n",
+ "# send a random row from the test set to score\n",
+ "random_index = np.random.randint(0, len(X_test)-1)\n",
+ "input_data = \"{\\\"data\\\": [\" + str(list(X_test[random_index])) + \"]}\"\n",
+ "\n",
+ "headers = {'Content-Type':'application/json'}\n",
+ "\n",
+ "resp = requests.post(service.scoring_uri, input_data, headers=headers)\n",
+ "\n",
+ "print(\"POST to url\", service.scoring_uri)\n",
+ "#print(\"input data:\", input_data)\n",
+ "print(\"label:\", y_test[random_index])\n",
+ "print(\"prediction:\", resp.text)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Let's look at the workspace after the web service was deployed. You should see \n",
+ "* a registered model named 'model' and with the id 'model:1'\n",
+ "* an image called 'tf-mnist' and with a docker image location pointing to your workspace's Azure Container Registry (ACR) \n",
+ "* a webservice called 'tf-mnist' with some scoring URL"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "models = ws.models\n",
+ "for name, model in models.items():\n",
+ " print(\"Model: {}, ID: {}\".format(name, model.id))\n",
+ " \n",
+ "images = ws.images\n",
+ "for name, image in images.items():\n",
+ " print(\"Image: {}, location: {}\".format(name, image.image_location))\n",
+ " \n",
+ "webservices = ws.webservices\n",
+ "for name, webservice in webservices.items():\n",
+ " print(\"Webservice: {}, scoring URI: {}\".format(name, webservice.scoring_uri))"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Clean up\n",
+ "You can delete the ACI deployment with a simple delete API call."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "service.delete()"
+ ]
}
- },
- "source": [
- "# Training, hyperparameter tune, and deploy with TensorFlow\n",
- "\n",
- "## Introduction\n",
- "This tutorial shows how to train a simple deep neural network using the MNIST dataset and TensorFlow on Azure Machine Learning. MNIST is a popular dataset consisting of 70,000 grayscale images. Each image is a handwritten digit of `28x28` pixels, representing number from 0 to 9. The goal is to create a multi-class classifier to identify the digit each image represents, and deploy it as a web service in Azure.\n",
- "\n",
- "For more information about the MNIST dataset, please visit [Yan LeCun's website](http://yann.lecun.com/exdb/mnist/).\n",
- "\n",
- "## Prerequisite:\n",
- "* Understand the [architecture and terms](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture) introduced by Azure Machine Learning\n",
- "* Go through the [00.configuration.ipynb](https://github.com/Azure/MachineLearningNotebooks/blob/master/00.configuration.ipynb) notebook to:\n",
- " * install the AML SDK\n",
- " * create a workspace and its configuration file (`config.json`)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Let's get started. First let's import some Python libraries."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "nbpresent": {
- "id": "c377ea0c-0cd9-4345-9be2-e20fb29c94c3"
- }
- },
- "outputs": [],
- "source": [
- "%matplotlib inline\n",
- "import numpy as np\n",
- "import os\n",
- "import matplotlib\n",
- "import matplotlib.pyplot as plt"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "nbpresent": {
- "id": "edaa7f2f-2439-4148-b57a-8c794c0945ec"
- }
- },
- "outputs": [],
- "source": [
- "import azureml\n",
- "from azureml.core import Workspace, Run\n",
- "\n",
- "# check core SDK version number\n",
- "print(\"Azure ML SDK Version: \", azureml.core.VERSION)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Diagnostics\n",
- "Opt-in diagnostics for better experience, quality, and security of future releases."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "Diagnostics"
- ]
- },
- "outputs": [],
- "source": [
- "from azureml.telemetry import set_diagnostics_collection\n",
- "\n",
- "set_diagnostics_collection(send_diagnostics=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Initialize workspace\n",
- "Initialize a [Workspace](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#workspace) object from the existing workspace you created in the Prerequisites step. `Workspace.from_config()` creates a workspace object from the details stored in `config.json`."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.workspace import Workspace\n",
- "\n",
- "ws = Workspace.from_config()\n",
- "print('Workspace name: ' + ws.name, \n",
- " 'Azure region: ' + ws.location, \n",
- " 'Subscription id: ' + ws.subscription_id, \n",
- " 'Resource group: ' + ws.resource_group, sep = '\\n')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {
- "nbpresent": {
- "id": "59f52294-4a25-4c92-bab8-3b07f0f44d15"
- }
- },
- "source": [
- "## Create an Azure ML experiment\n",
- "Let's create an experiment named \"tf-mnist\" and a folder to hold the training scripts. The script runs will be recorded under the experiment in Azure."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "nbpresent": {
- "id": "bc70f780-c240-4779-96f3-bc5ef9a37d59"
- }
- },
- "outputs": [],
- "source": [
- "from azureml.core import Experiment\n",
- "\n",
- "script_folder = './tf-mnist'\n",
- "os.makedirs(script_folder, exist_ok=True)\n",
- "\n",
- "exp = Experiment(workspace=ws, name='tf-mnist')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {
- "nbpresent": {
- "id": "defe921f-8097-44c3-8336-8af6700804a7"
- }
- },
- "source": [
- "## Download MNIST dataset\n",
- "In order to train on the MNIST dataset we will first need to download it from Yan LeCun's web site directly and save them in a `data` folder locally."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import os\n",
- "import urllib\n",
- "\n",
- "os.makedirs('./data/mnist', exist_ok=True)\n",
- "\n",
- "urllib.request.urlretrieve('http://yann.lecun.com/exdb/mnist/train-images-idx3-ubyte.gz', filename = './data/mnist/train-images.gz')\n",
- "urllib.request.urlretrieve('http://yann.lecun.com/exdb/mnist/train-labels-idx1-ubyte.gz', filename = './data/mnist/train-labels.gz')\n",
- "urllib.request.urlretrieve('http://yann.lecun.com/exdb/mnist/t10k-images-idx3-ubyte.gz', filename = './data/mnist/test-images.gz')\n",
- "urllib.request.urlretrieve('http://yann.lecun.com/exdb/mnist/t10k-labels-idx1-ubyte.gz', filename = './data/mnist/test-labels.gz')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {
- "nbpresent": {
- "id": "c3f2f57c-7454-4d3e-b38d-b0946cf066ea"
- }
- },
- "source": [
- "## Show some sample images\n",
- "Let's load the downloaded compressed file into numpy arrays using some utility functions included in the `utils.py` library file from the current folder. Then we use `matplotlib` to plot 30 random images from the dataset along with their labels."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "nbpresent": {
- "id": "396d478b-34aa-4afa-9898-cdce8222a516"
- }
- },
- "outputs": [],
- "source": [
- "from utils import load_data\n",
- "\n",
- "# note we also shrink the intensity values (X) from 0-255 to 0-1. This helps the neural network converge faster.\n",
- "X_train = load_data('./data/mnist/train-images.gz', False) / 255.0\n",
- "y_train = load_data('./data/mnist/train-labels.gz', True).reshape(-1)\n",
- "\n",
- "X_test = load_data('./data/mnist/test-images.gz', False) / 255.0\n",
- "y_test = load_data('./data/mnist/test-labels.gz', True).reshape(-1)\n",
- "\n",
- "count = 0\n",
- "sample_size = 30\n",
- "plt.figure(figsize = (16, 6))\n",
- "for i in np.random.permutation(X_train.shape[0])[:sample_size]:\n",
- " count = count + 1\n",
- " plt.subplot(1, sample_size, count)\n",
- " plt.axhline('')\n",
- " plt.axvline('')\n",
- " plt.text(x = 10, y = -10, s = y_train[i], fontsize = 18)\n",
- " plt.imshow(X_train[i].reshape(28, 28), cmap = plt.cm.Greys)\n",
- "plt.show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Upload MNIST dataset to default datastore \n",
- "A [datastore](https://docs.microsoft.com/azure/machine-learning/service/how-to-access-data) is a place where data can be stored that is then made accessible to a Run either by means of mounting or copying the data to the compute target. A datastore can either be backed by an Azure Blob Storage or and Azure File Share (ADLS will be supported in the future). For simple data handling, each workspace provides a default datastore that can be used, in case the data is not already in Blob Storage or File Share."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "ds = ws.get_default_datastore()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "In this next step, we will upload the training and test set into the workspace's default datastore, which we will then later be mount on an `AmlCompute` cluster for training."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "ds.upload(src_dir='./data/mnist', target_path='mnist', overwrite=True, show_progress=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Create or Attach existing AmlCompute\n",
- "You will need to create a [compute target](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#compute-target) for training your model. In this tutorial, you create `AmlCompute` as your training compute resource."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "If we could not find the cluster with the given name, then we will create a new cluster here. We will create an `AmlCompute` cluster of `STANDARD_NC6` GPU VMs. This process is broken down into 3 steps:\n",
- "1. create the configuration (this step is local and only takes a second)\n",
- "2. create the cluster (this step will take about **20 seconds**)\n",
- "3. provision the VMs to bring the cluster to the initial size (of 1 in this case). This step will take about **3-5 minutes** and is providing only sparse output in the process. Please make sure to wait until the call returns before moving to the next cell"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.compute import ComputeTarget, AmlCompute\n",
- "from azureml.core.compute_target import ComputeTargetException\n",
- "\n",
- "# choose a name for your cluster\n",
- "cluster_name = \"gpucluster\"\n",
- "\n",
- "try:\n",
- " compute_target = ComputeTarget(workspace=ws, name=cluster_name)\n",
- " print('Found existing compute target')\n",
- "except ComputeTargetException:\n",
- " print('Creating a new compute target...')\n",
- " compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_NC6', \n",
- " max_nodes=4)\n",
- "\n",
- " # create the cluster\n",
- " compute_target = ComputeTarget.create(ws, cluster_name, compute_config)\n",
- "\n",
- " # can poll for a minimum number of nodes and for a specific timeout. \n",
- " # if no min node count is provided it uses the scale settings for the cluster\n",
- " compute_target.wait_for_completion(show_output=True, min_node_count=None, timeout_in_minutes=20)\n",
- "\n",
- "# Use the 'status' property to get a detailed status for the current cluster. \n",
- "print(compute_target.status.serialize())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Now that you have created the compute target, let's see what the workspace's `compute_targets` property returns. You should now see one entry named 'gpucluster' of type `AmlCompute`."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "compute_targets = ws.compute_targets\n",
- "for name, ct in compute_targets.items():\n",
- " print(name, ct.type, ct.provisioning_state)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Copy the training files into the script folder\n",
- "The TensorFlow training script is already created for you. You can simply copy it into the script folder, together with the utility library used to load compressed data file into numpy array."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import shutil\n",
- "\n",
- "# the training logic is in the tf_mnist.py file.\n",
- "shutil.copy('./tf_mnist.py', script_folder)\n",
- "\n",
- "# the utils.py just helps loading data from the downloaded MNIST dataset into numpy arrays.\n",
- "shutil.copy('./utils.py', script_folder)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {
- "nbpresent": {
- "id": "2039d2d5-aca6-4f25-a12f-df9ae6529cae"
- }
- },
- "source": [
- "## Construct neural network in TensorFlow\n",
- "In the training script `tf_mnist.py`, it creates a very simple DNN (deep neural network), with just 2 hidden layers. The input layer has 28 * 28 = 784 neurons, each representing a pixel in an image. The first hidden layer has 300 neurons, and the second hidden layer has 100 neurons. The output layer has 10 neurons, each representing a targeted label from 0 to 9.\n",
- "\n",
- ""
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Azure ML concepts \n",
- "Please note the following three things in the code below:\n",
- "1. The script accepts arguments using the argparse package. In this case there is one argument `--data_folder` which specifies the file system folder in which the script can find the MNIST data\n",
- "```\n",
- " parser = argparse.ArgumentParser()\n",
- " parser.add_argument('--data_folder')\n",
- "```\n",
- "2. The script is accessing the Azure ML `Run` object by executing `run = Run.get_context()`. Further down the script is using the `run` to report the training accuracy and the validation accuracy as training progresses.\n",
- "```\n",
- " run.log('training_acc', np.float(acc_train))\n",
- " run.log('validation_acc', np.float(acc_val))\n",
- "```\n",
- "3. When running the script on Azure ML, you can write files out to a folder `./outputs` that is relative to the root directory. This folder is specially tracked by Azure ML in the sense that any files written to that folder during script execution on the remote target will be picked up by Run History; these files (known as artifacts) will be available as part of the run history record."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "The next cell will print out the training code for you to inspect it."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "with open(os.path.join(script_folder, './tf_mnist.py'), 'r') as f:\n",
- " print(f.read())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Create TensorFlow estimator\n",
- "Next, we construct an `azureml.train.dnn.TensorFlow` estimator object, use the Batch AI cluster as compute target, and pass the mount-point of the datastore to the training code as a parameter.\n",
- "The TensorFlow estimator is providing a simple way of launching a TensorFlow training job on a compute target. It will automatically provide a docker image that has TensorFlow installed -- if additional pip or conda packages are required, their names can be passed in via the `pip_packages` and `conda_packages` arguments and they will be included in the resulting docker."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.train.dnn import TensorFlow\n",
- "\n",
- "script_params = {\n",
- " '--data-folder': ws.get_default_datastore().as_mount(),\n",
- " '--batch-size': 50,\n",
- " '--first-layer-neurons': 300,\n",
- " '--second-layer-neurons': 100,\n",
- " '--learning-rate': 0.01\n",
- "}\n",
- "\n",
- "est = TensorFlow(source_directory=script_folder,\n",
- " script_params=script_params,\n",
- " compute_target=compute_target,\n",
- " entry_script='tf_mnist.py', \n",
- " use_gpu=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Submit job to run\n",
- "Calling the `fit` function on the estimator submits the job to Azure ML for execution. Submitting the job should only take a few seconds."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run = exp.submit(est)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Monitor the Run\n",
- "As the Run is executed, it will go through the following stages:\n",
- "1. Preparing: A docker image is created matching the Python environment specified by the TensorFlow estimator and it will be uploaded to the workspace's Azure Container Registry. This step will only happen once for each Python environment -- the container will then be cached for subsequent runs. Creating and uploading the image takes about **5 minutes**. While the job is preparing, logs are streamed to the run history and can be viewed to monitor the progress of the image creation.\n",
- "\n",
- "2. Scaling: If the compute needs to be scaled up (i.e. the Batch AI cluster requires more nodes to execute the run than currently available), the cluster will attempt to scale up in order to make the required amount of nodes available. Scaling typically takes about **5 minutes**.\n",
- "\n",
- "3. Running: All scripts in the script folder are uploaded to the compute target, data stores are mounted/copied and the `entry_script` is executed. While the job is running, stdout and the `./logs` folder are streamed to the run history and can be viewed to monitor the progress of the run.\n",
- "\n",
- "4. Post-Processing: The `./outputs` folder of the run is copied over to the run history\n",
- "\n",
- "There are multiple ways to check the progress of a running job. We can use a Jupyter notebook widget. \n",
- "\n",
- "**Note: The widget will automatically update ever 10-15 seconds, always showing you the most up-to-date information about the run**"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.widgets import RunDetails\n",
- "\n",
- "RunDetails(run).show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "We can also periodically check the status of the run object, and navigate to Azure portal to monitor the run."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run.wait_for_completion(show_output=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### The Run object\n",
- "The Run object provides the interface to the run history -- both to the job and to the control plane (this notebook), and both while the job is running and after it has completed. It provides a number of interesting features for instance:\n",
- "* `run.get_details()`: Provides a rich set of properties of the run\n",
- "* `run.get_metrics()`: Provides a dictionary with all the metrics that were reported for the Run\n",
- "* `run.get_file_names()`: List all the files that were uploaded to the run history for this Run. This will include the `outputs` and `logs` folder, azureml-logs and other logs, as well as files that were explicitly uploaded to the run using `run.upload_file()`\n",
- "\n",
- "Below are some examples -- please run through them and inspect their output. "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run.get_details()"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run.get_metrics()"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run.get_file_names()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Plot accuracy over epochs\n",
- "Since we can retrieve the metrics from the run, we can easily make plots using `matplotlib` in the notebook. Then we can add the plotted image to the run using `run.log_image()`, so all information about the run is kept together."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import os\n",
- "\n",
- "os.makedirs('./imgs', exist_ok=True)\n",
- "metrics = run.get_metrics()\n",
- "\n",
- "plt.figure(figsize = (13,5))\n",
- "plt.plot(metrics['validation_acc'], 'r-', lw=4, alpha=.6)\n",
- "plt.plot(metrics['training_acc'], 'b--', alpha=0.5)\n",
- "plt.legend(['Full evaluation set', 'Training set mini-batch'])\n",
- "plt.xlabel('epochs', fontsize=14)\n",
- "plt.ylabel('accuracy', fontsize=14)\n",
- "plt.title('Accuracy over Epochs', fontsize=16)\n",
- "run.log_image(name='acc_over_epochs.png', plot=plt)\n",
- "plt.show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Download the saved model"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "In the training script, a TensorFlow `saver` object is used to persist the model in a local folder (local to the compute target). The model was saved to the `./outputs` folder on the disk of the Batch AI cluster node where the job is run. Azure ML automatically uploaded anything written in the `./outputs` folder into run history file store. Subsequently, we can use the `Run` object to download the model files the `saver` object saved. They are under the the `outputs/model` folder in the run history file store, and are downloaded into a local folder named `model`. Note the TensorFlow model consists of four files in binary format and they are not human-readable."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# create a model folder in the current directory\n",
- "os.makedirs('./model', exist_ok=True)\n",
- "\n",
- "for f in run.get_file_names():\n",
- " if f.startswith('outputs/model'):\n",
- " output_file_path = os.path.join('./model', f.split('/')[-1])\n",
- " print('Downloading from {} to {} ...'.format(f, output_file_path))\n",
- " run.download_file(name=f, output_file_path=output_file_path)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Predict on the test set\n",
- "Now load the saved TensorFlow graph, and list all operations under the `network` scope. This way we can discover the input tensor `network/X:0` and the output tensor `network/output/MatMul:0`, and use them in the scoring script in the next step.\n",
- "\n",
- "Note: if your local TensorFlow version is different than the version running in the cluster where the model is trained, you might see a \"compiletime version mismatch\" warning. You can ignore it."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import tensorflow as tf\n",
- "\n",
- "tf.reset_default_graph()\n",
- "\n",
- "saver = tf.train.import_meta_graph(\"./model/mnist-tf.model.meta\")\n",
- "graph = tf.get_default_graph()\n",
- "\n",
- "for op in graph.get_operations():\n",
- " if op.name.startswith('network'):\n",
- " print(op.name)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Feed test dataset to the persisted model to get predictions."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# input tensor. this is an array of 784 elements, each representing the intensity of a pixel in the digit image.\n",
- "X = tf.get_default_graph().get_tensor_by_name(\"network/X:0\")\n",
- "# output tensor. this is an array of 10 elements, each representing the probability of predicted value of the digit.\n",
- "output = tf.get_default_graph().get_tensor_by_name(\"network/output/MatMul:0\")\n",
- "\n",
- "with tf.Session() as sess:\n",
- " saver.restore(sess, './model/mnist-tf.model')\n",
- " k = output.eval(feed_dict={X : X_test})\n",
- "# get the prediction, which is the index of the element that has the largest probability value.\n",
- "y_hat = np.argmax(k, axis=1)\n",
- "\n",
- "# print the first 30 labels and predictions\n",
- "print('labels: \\t', y_test[:30])\n",
- "print('predictions:\\t', y_hat[:30])"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Calculate the overall accuracy by comparing the predicted value against the test set."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "print(\"Accuracy on the test set:\", np.average(y_hat == y_test))"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Intelligent hyperparameter tuning\n",
- "We have trained the model with one set of hyperparameters, now let's how we can do hyperparameter tuning by launching multiple runs on the cluster. First let's define the parameter space using random sampling."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.train.hyperdrive import *\n",
- "\n",
- "ps = RandomParameterSampling(\n",
- " {\n",
- " '--batch-size': choice(25, 50, 100),\n",
- " '--first-layer-neurons': choice(10, 50, 200, 300, 500),\n",
- " '--second-layer-neurons': choice(10, 50, 200, 500),\n",
- " '--learning-rate': loguniform(-6, -1)\n",
- " }\n",
- ")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Next, we will create a new estimator without the above parameters since they will be passed in later. Note we still need to keep the `data-folder` parameter since that's not a hyperparamter we will sweep."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "est = TensorFlow(source_directory=script_folder,\n",
- " script_params={'--data-folder': ws.get_default_datastore().as_mount()},\n",
- " compute_target=compute_target,\n",
- " entry_script='tf_mnist.py', \n",
- " use_gpu=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Now we will define an early termnination policy. The `BanditPolicy` basically states to check the job every 2 iterations. If the primary metric (defined later) falls outside of the top 10% range, Azure ML terminate the job. This saves us from continuing to explore hyperparameters that don't show promise of helping reach our target metric."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "policy = BanditPolicy(evaluation_interval=2, slack_factor=0.1)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Now we are ready to configure a run configuration object, and specify the primary metric `validation_acc` that's recorded in your training runs. If you go back to visit the training script, you will notice that this value is being logged after every epoch (a full batch set). We also want to tell the service that we are looking to maximizing this value. We also set the number of samples to 20, and maximal concurrent job to 4, which is the same as the number of nodes in our computer cluster."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "htc = HyperDriveRunConfig(estimator=est, \n",
- " hyperparameter_sampling=ps, \n",
- " policy=policy, \n",
- " primary_metric_name='validation_acc', \n",
- " primary_metric_goal=PrimaryMetricGoal.MAXIMIZE, \n",
- " max_total_runs=8,\n",
- " max_concurrent_runs=4)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Finally, let's launch the hyperparameter tuning job."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "htr = exp.submit(config=htc)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "We can use a run history widget to show the progress. Be patient as this might take a while to complete."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "RunDetails(htr).show()"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "htr.wait_for_completion(show_output=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Find and register best model\n",
- "When all the jobs finish, we can find out the one that has the highest accuracy."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "best_run = htr.get_best_run_by_primary_metric()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Now let's list the model files uploaded during the run."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "print(best_run.get_file_names())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "We can then register the folder (and all files in it) as a model named `tf-dnn-mnist` under the workspace for deployment."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "model = best_run.register_model(model_name='tf-dnn-mnist', model_path='outputs/model')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Deploy the model in ACI\n",
- "Now we are ready to deploy the model as a web service running in Azure Container Instance [ACI](https://azure.microsoft.com/en-us/services/container-instances/). Azure Machine Learning accomplishes this by constructing a Docker image with the scoring logic and model baked in.\n",
- "### Create score.py\n",
- "First, we will create a scoring script that will be invoked by the web service call. \n",
- "\n",
- "* Note that the scoring script must have two required functions, `init()` and `run(input_data)`. \n",
- " * In `init()` function, you typically load the model into a global object. This function is executed only once when the Docker container is started. \n",
- " * In `run(input_data)` function, the model is used to predict a value based on the input data. The input and output to `run` typically use JSON as serialization and de-serialization format but you are not limited to that."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%writefile score.py\n",
- "import json\n",
- "import numpy as np\n",
- "import os\n",
- "import tensorflow as tf\n",
- "\n",
- "from azureml.core.model import Model\n",
- "\n",
- "def init():\n",
- " global X, output, sess\n",
- " tf.reset_default_graph()\n",
- " model_root = Model.get_model_path('tf-dnn-mnist')\n",
- " saver = tf.train.import_meta_graph(os.path.join(model_root, 'mnist-tf.model.meta'))\n",
- " X = tf.get_default_graph().get_tensor_by_name(\"network/X:0\")\n",
- " output = tf.get_default_graph().get_tensor_by_name(\"network/output/MatMul:0\")\n",
- " \n",
- " sess = tf.Session()\n",
- " saver.restore(sess, os.path.join(model_root, 'mnist-tf.model'))\n",
- "\n",
- "def run(raw_data):\n",
- " data = np.array(json.loads(raw_data)['data'])\n",
- " # make prediction\n",
- " out = output.eval(session=sess, feed_dict={X: data})\n",
- " y_hat = np.argmax(out, axis=1)\n",
- " return y_hat.tolist()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create myenv.yml\n",
- "We also need to create an environment file so that Azure Machine Learning can install the necessary packages in the Docker image which are required by your scoring script. In this case, we need to specify packages `numpy`, `tensorflow`."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.runconfig import CondaDependencies\n",
- "\n",
- "cd = CondaDependencies.create()\n",
- "cd.add_conda_package('numpy')\n",
- "cd.add_tensorflow_conda_package()\n",
- "cd.save_to_file(base_directory='./', conda_file_path='myenv.yml')\n",
- "\n",
- "print(cd.serialize_to_string())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Deploy to ACI\n",
- "We are almost ready to deploy. Create a deployment configuration and specify the number of CPUs and gigbyte of RAM needed for your ACI container. "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.webservice import AciWebservice\n",
- "\n",
- "aciconfig = AciWebservice.deploy_configuration(cpu_cores=1, \n",
- " memory_gb=1, \n",
- " tags={'name':'mnist', 'framework': 'TensorFlow DNN'},\n",
- " description='Tensorflow DNN on MNIST')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Deployment Process\n",
- "Now we can deploy. **This cell will run for about 7-8 minutes**. Behind the scene, it will do the following:\n",
- "1. **Register model** \n",
- "Take the local `model` folder (which contains our previously downloaded trained model files) and register it (and the files inside that folder) as a model named `model` under the workspace. Azure ML will register the model directory or model file(s) we specify to the `model_paths` parameter of the `Webservice.deploy` call.\n",
- "2. **Build Docker image** \n",
- "Build a Docker image using the scoring file (`score.py`), the environment file (`myenv.yml`), and the `model` folder containing the TensorFlow model files. \n",
- "3. **Register image** \n",
- "Register that image under the workspace. \n",
- "4. **Ship to ACI** \n",
- "And finally ship the image to the ACI infrastructure, start up a container in ACI using that image, and expose an HTTP endpoint to accept REST client calls."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.image import ContainerImage\n",
- "\n",
- "imgconfig = ContainerImage.image_configuration(execution_script=\"score.py\", \n",
- " runtime=\"python\", \n",
- " conda_file=\"myenv.yml\")"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%time\n",
- "from azureml.core.webservice import Webservice\n",
- "\n",
- "service = Webservice.deploy_from_model(workspace=ws,\n",
- " name='tf-mnist-svc',\n",
- " deployment_config=aciconfig,\n",
- " models=[model],\n",
- " image_config=imgconfig)\n",
- "\n",
- "service.wait_for_deployment(show_output=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "**Tip: If something goes wrong with the deployment, the first thing to look at is the logs from the service by running the following command:**"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "print(service.get_logs())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "This is the scoring web service endpoint:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "print(service.scoring_uri)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Test the deployed model\n",
- "Let's test the deployed model. Pick 30 random samples from the test set, and send it to the web service hosted in ACI. Note here we are using the `run` API in the SDK to invoke the service. You can also make raw HTTP calls using any HTTP tool such as curl.\n",
- "\n",
- "After the invocation, we print the returned predictions and plot them along with the input images. Use red font color and inversed image (white on black) to highlight the misclassified samples. Note since the model accuracy is pretty high, you might have to run the below cell a few times before you can see a misclassified sample."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import json\n",
- "\n",
- "# find 30 random samples from test set\n",
- "n = 30\n",
- "sample_indices = np.random.permutation(X_test.shape[0])[0:n]\n",
- "\n",
- "test_samples = json.dumps({\"data\": X_test[sample_indices].tolist()})\n",
- "test_samples = bytes(test_samples, encoding='utf8')\n",
- "\n",
- "# predict using the deployed model\n",
- "result = service.run(input_data=test_samples)\n",
- "\n",
- "# compare actual value vs. the predicted values:\n",
- "i = 0\n",
- "plt.figure(figsize = (20, 1))\n",
- "\n",
- "for s in sample_indices:\n",
- " plt.subplot(1, n, i + 1)\n",
- " plt.axhline('')\n",
- " plt.axvline('')\n",
- " \n",
- " # use different color for misclassified sample\n",
- " font_color = 'red' if y_test[s] != result[i] else 'black'\n",
- " clr_map = plt.cm.gray if y_test[s] != result[i] else plt.cm.Greys\n",
- " \n",
- " plt.text(x=10, y=-10, s=y_hat[s], fontsize=18, color=font_color)\n",
- " plt.imshow(X_test[s].reshape(28, 28), cmap=clr_map)\n",
- " \n",
- " i = i + 1\n",
- "plt.show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "We can also send raw HTTP request to the service."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import requests\n",
- "import json\n",
- "\n",
- "# send a random row from the test set to score\n",
- "random_index = np.random.randint(0, len(X_test)-1)\n",
- "input_data = \"{\\\"data\\\": [\" + str(list(X_test[random_index])) + \"]}\"\n",
- "\n",
- "headers = {'Content-Type':'application/json'}\n",
- "\n",
- "resp = requests.post(service.scoring_uri, input_data, headers=headers)\n",
- "\n",
- "print(\"POST to url\", service.scoring_uri)\n",
- "#print(\"input data:\", input_data)\n",
- "print(\"label:\", y_test[random_index])\n",
- "print(\"prediction:\", resp.text)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Let's look at the workspace after the web service was deployed. You should see \n",
- "* a registered model named 'model' and with the id 'model:1'\n",
- "* an image called 'tf-mnist' and with a docker image location pointing to your workspace's Azure Container Registry (ACR) \n",
- "* a webservice called 'tf-mnist' with some scoring URL"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "models = ws.models\n",
- "for name, model in models.items():\n",
- " print(\"Model: {}, ID: {}\".format(name, model.id))\n",
- " \n",
- "images = ws.images\n",
- "for name, image in images.items():\n",
- " print(\"Image: {}, location: {}\".format(name, image.image_location))\n",
- " \n",
- "webservices = ws.webservices\n",
- "for name, webservice in webservices.items():\n",
- " print(\"Webservice: {}, scoring URI: {}\".format(name, webservice.scoring_uri))"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Clean up\n",
- "You can delete the ACI deployment with a simple delete API call."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "service.delete()"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "minxia"
- }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "minxia"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.6"
+ },
+ "msauthor": "minxia"
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- },
- "msauthor": "minxia"
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/training/logging-api/logging-api.ipynb b/how-to-use-azureml/training/logging-api/logging-api.ipynb
index 81013269..4c8041f3 100644
--- a/how-to-use-azureml/training/logging-api/logging-api.ipynb
+++ b/how-to-use-azureml/training/logging-api/logging-api.ipynb
@@ -1,328 +1,328 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# 06. Logging APIs\n",
- "This notebook showcase various ways to use the Azure Machine Learning service run logging APIs, and view the results in the Azure portal."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Prerequisites\n",
- "Make sure you go through the [00. Installation and Configuration](../../00.configuration.ipynb) Notebook first if you haven't. Also make sure you have tqdm and matplotlib installed in the current kernel.\n",
- "\n",
- "```\n",
- "(myenv) $ conda install -y tqdm matplotlib\n",
- "```"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Validate Azure ML SDK installation and get version number for debugging purposes"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "install"
- ]
- },
- "outputs": [],
- "source": [
- "from azureml.core import Experiment, Run, Workspace\n",
- "import azureml.core\n",
- "import numpy as np\n",
- "\n",
- "# Check core SDK version number\n",
- "print(\"SDK version:\", azureml.core.VERSION)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Initialize Workspace\n",
- "\n",
- "Initialize a workspace object from persisted configuration."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "create workspace"
- ]
- },
- "outputs": [],
- "source": [
- "ws = Workspace.from_config()\n",
- "print('Workspace name: ' + ws.name, \n",
- " 'Azure region: ' + ws.location, \n",
- " 'Subscription id: ' + ws.subscription_id, \n",
- " 'Resource group: ' + ws.resource_group, sep='\\n')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Set experiment\n",
- "Create a new experiment (or get the one with such name)."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "exp = Experiment(workspace=ws, name='logging-api-test')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Log metrics\n",
- "We will start a run, and use the various logging APIs to record different types of metrics during the run."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from tqdm import tqdm\n",
- "\n",
- "# start logging for the run\n",
- "run = exp.start_logging()\n",
- "\n",
- "# log a string value\n",
- "run.log(name='Name', value='Logging API run')\n",
- "\n",
- "# log a numerical value\n",
- "run.log(name='Magic Number', value=42)\n",
- "\n",
- "# Log a list of values. Note this will generate a single-variable line chart.\n",
- "run.log_list(name='Fibonacci', value=[0, 1, 1, 2, 3, 5, 8, 13, 21, 34, 55, 89])\n",
- "\n",
- "# create a dictionary to hold a table of values\n",
- "sines = {}\n",
- "sines['angle'] = []\n",
- "sines['sine'] = []\n",
- "\n",
- "for i in tqdm(range(-10, 10)):\n",
- " # log a metric value repeatedly, this will generate a single-variable line chart.\n",
- " run.log(name='Sigmoid', value=1 / (1 + np.exp(-i)))\n",
- " angle = i / 2.0\n",
- " \n",
- " # log a 2 (or more) values as a metric repeatedly. This will generate a 2-variable line chart if you have 2 numerical columns.\n",
- " run.log_row(name='Cosine Wave', angle=angle, cos=np.cos(angle))\n",
- " \n",
- " sines['angle'].append(angle)\n",
- " sines['sine'].append(np.sin(angle))\n",
- "\n",
- "# log a dictionary as a table, this will generate a 2-variable chart if you have 2 numerical columns\n",
- "run.log_table(name='Sine Wave', value=sines)\n",
- "\n",
- "run.complete()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Even after the run is marked completed, you can still log things."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Log an image\n",
- "This is how to log a _matplotlib_ pyplot object."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%matplotlib inline\n",
- "import matplotlib.pyplot as plt\n",
- "angle = np.linspace(-3, 3, 50)\n",
- "plt.plot(angle, np.tanh(angle), label='tanh')\n",
- "plt.legend(fontsize=12)\n",
- "plt.title('Hyperbolic Tangent', fontsize=16)\n",
- "plt.grid(True)\n",
- "\n",
- "run.log_image(name='Hyperbolic Tangent', plot=plt)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Upload a file"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "You can also upload an abitrary file. First, let's create a dummy file locally."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%writefile myfile.txt\n",
- "\n",
- "This is a dummy file."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Now let's upload this file into the run record as a run artifact, and display the properties after the upload."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "props = run.upload_file(name='myfile_in_the_cloud.txt', path_or_stream='./myfile.txt')\n",
- "props.serialize()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Examine the run"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Now let's take a look at the run detail page in Azure portal. Make sure you checkout the various charts and plots generated/uploaded."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "You can get all the metrics in that run back."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run.get_metrics()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "You can also see the files uploaded for this run."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run.get_file_names()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "You can also download all the files locally."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import os\n",
- "os.makedirs('files', exist_ok=True)\n",
- "\n",
- "for f in run.get_file_names():\n",
- " dest = os.path.join('files', f.split('/')[-1])\n",
- " print('Downloading file {} to {}...'.format(f, dest))\n",
- " run.download_file(f, dest) "
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "haining"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# 06. Logging APIs\n",
+ "This notebook showcase various ways to use the Azure Machine Learning service run logging APIs, and view the results in the Azure portal."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Prerequisites\n",
+ "Make sure you go through the [configuration notebook](../../../configuration.ipynb) first if you haven't. Also make sure you have tqdm and matplotlib installed in the current kernel.\n",
+ "\n",
+ "```\n",
+ "(myenv) $ conda install -y tqdm matplotlib\n",
+ "```"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Validate Azure ML SDK installation and get version number for debugging purposes"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "install"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "from azureml.core import Experiment, Run, Workspace\n",
+ "import azureml.core\n",
+ "import numpy as np\n",
+ "\n",
+ "# Check core SDK version number\n",
+ "print(\"SDK version:\", azureml.core.VERSION)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Initialize Workspace\n",
+ "\n",
+ "Initialize a workspace object from persisted configuration."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "create workspace"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "ws = Workspace.from_config()\n",
+ "print('Workspace name: ' + ws.name, \n",
+ " 'Azure region: ' + ws.location, \n",
+ " 'Subscription id: ' + ws.subscription_id, \n",
+ " 'Resource group: ' + ws.resource_group, sep='\\n')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Set experiment\n",
+ "Create a new experiment (or get the one with such name)."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "exp = Experiment(workspace=ws, name='logging-api-test')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Log metrics\n",
+ "We will start a run, and use the various logging APIs to record different types of metrics during the run."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from tqdm import tqdm\n",
+ "\n",
+ "# start logging for the run\n",
+ "run = exp.start_logging()\n",
+ "\n",
+ "# log a string value\n",
+ "run.log(name='Name', value='Logging API run')\n",
+ "\n",
+ "# log a numerical value\n",
+ "run.log(name='Magic Number', value=42)\n",
+ "\n",
+ "# Log a list of values. Note this will generate a single-variable line chart.\n",
+ "run.log_list(name='Fibonacci', value=[0, 1, 1, 2, 3, 5, 8, 13, 21, 34, 55, 89])\n",
+ "\n",
+ "# create a dictionary to hold a table of values\n",
+ "sines = {}\n",
+ "sines['angle'] = []\n",
+ "sines['sine'] = []\n",
+ "\n",
+ "for i in tqdm(range(-10, 10)):\n",
+ " # log a metric value repeatedly, this will generate a single-variable line chart.\n",
+ " run.log(name='Sigmoid', value=1 / (1 + np.exp(-i)))\n",
+ " angle = i / 2.0\n",
+ " \n",
+ " # log a 2 (or more) values as a metric repeatedly. This will generate a 2-variable line chart if you have 2 numerical columns.\n",
+ " run.log_row(name='Cosine Wave', angle=angle, cos=np.cos(angle))\n",
+ " \n",
+ " sines['angle'].append(angle)\n",
+ " sines['sine'].append(np.sin(angle))\n",
+ "\n",
+ "# log a dictionary as a table, this will generate a 2-variable chart if you have 2 numerical columns\n",
+ "run.log_table(name='Sine Wave', value=sines)\n",
+ "\n",
+ "run.complete()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Even after the run is marked completed, you can still log things."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Log an image\n",
+ "This is how to log a _matplotlib_ pyplot object."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%matplotlib inline\n",
+ "import matplotlib.pyplot as plt\n",
+ "angle = np.linspace(-3, 3, 50)\n",
+ "plt.plot(angle, np.tanh(angle), label='tanh')\n",
+ "plt.legend(fontsize=12)\n",
+ "plt.title('Hyperbolic Tangent', fontsize=16)\n",
+ "plt.grid(True)\n",
+ "\n",
+ "run.log_image(name='Hyperbolic Tangent', plot=plt)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Upload a file"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "You can also upload an abitrary file. First, let's create a dummy file locally."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%writefile myfile.txt\n",
+ "\n",
+ "This is a dummy file."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Now let's upload this file into the run record as a run artifact, and display the properties after the upload."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "props = run.upload_file(name='myfile_in_the_cloud.txt', path_or_stream='./myfile.txt')\n",
+ "props.serialize()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Examine the run"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Now let's take a look at the run detail page in Azure portal. Make sure you checkout the various charts and plots generated/uploaded."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "You can get all the metrics in that run back."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run.get_metrics()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "You can also see the files uploaded for this run."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run.get_file_names()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "You can also download all the files locally."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import os\n",
+ "os.makedirs('files', exist_ok=True)\n",
+ "\n",
+ "for f in run.get_file_names():\n",
+ " dest = os.path.join('files', f.split('/')[-1])\n",
+ " print('Downloading file {} to {}...'.format(f, dest))\n",
+ " run.download_file(f, dest) "
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "haining"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.6"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/training/train-on-amlcompute/train-on-amlcompute.ipynb b/how-to-use-azureml/training/train-on-amlcompute/train-on-amlcompute.ipynb
index 4635716d..1e4a09cc 100644
--- a/how-to-use-azureml/training/train-on-amlcompute/train-on-amlcompute.ipynb
+++ b/how-to-use-azureml/training/train-on-amlcompute/train-on-amlcompute.ipynb
@@ -1,516 +1,516 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Train using Azure Machine Learning Compute\n",
- "\n",
- "* Initialize a Workspace\n",
- "* Create an Experiment\n",
- "* Introduction to AmlCompute\n",
- "* Submit an AmlCompute run in a few different ways\n",
- " - Provision as a run based compute target \n",
- " - Provision as a persistent compute target (Basic)\n",
- " - Provision as a persistent compute target (Advanced)\n",
- "* Additional operations to perform on AmlCompute\n",
- "* Find the best model in the run"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Prerequisites\n",
- "Make sure you go through the [00.configuration.ipynb](https://github.com/Azure/MachineLearningNotebooks/blob/master/00.configuration.ipynb) Notebook first if you haven't."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Check core SDK version number\n",
- "import azureml.core\n",
- "\n",
- "print(\"SDK version:\", azureml.core.VERSION)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Initialize a Workspace\n",
- "\n",
- "Initialize a workspace object from persisted configuration"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "create workspace"
- ]
- },
- "outputs": [],
- "source": [
- "from azureml.core import Workspace\n",
- "\n",
- "ws = Workspace.from_config()\n",
- "print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep = '\\n')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Create An Experiment\n",
- "\n",
- "**Experiment** is a logical container in an Azure ML Workspace. It hosts run records which can include run metrics and output artifacts from your experiments."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core import Experiment\n",
- "experiment_name = 'train-on-amlcompute'\n",
- "experiment = Experiment(workspace = ws, name = experiment_name)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Introduction to AmlCompute\n",
- "\n",
- "Azure Machine Learning Compute is managed compute infrastructure that allows the user to easily create single to multi-node compute of the appropriate VM Family. It is created **within your workspace region** and is a resource that can be used by other users in your workspace. It autoscales by default to the max_nodes, when a job is submitted, and executes in a containerized environment packaging the dependencies as specified by the user. \n",
- "\n",
- "Since it is managed compute, job scheduling and cluster management are handled internally by Azure Machine Learning service. \n",
- "\n",
- "For more information on Azure Machine Learning Compute, please read [this article](https://docs.microsoft.com/azure/machine-learning/service/how-to-set-up-training-targets#amlcompute)\n",
- "\n",
- "If you are an existing BatchAI customer who is migrating to Azure Machine Learning, please read [this article](https://aka.ms/batchai-retirement)\n",
- "\n",
- "**Note**: As with other Azure services, there are limits on certain resources (for eg. AmlCompute quota) associated with the Azure Machine Learning service. Please read [this article](https://docs.microsoft.com/azure/machine-learning/service/how-to-manage-quotas) on the default limits and how to request more quota.\n",
- "\n",
- "\n",
- "The training script `train.py` is already created for you. Let's have a look."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Submit an AmlCompute run in a few different ways\n",
- "\n",
- "First lets check which VM families are available in your region. Azure is a regional service and some specialized SKUs (especially GPUs) are only available in certain regions. Since AmlCompute is created in the region of your workspace, we will use the supported_vms () function to see if the VM family we want to use ('STANDARD_D2_V2') is supported.\n",
- "\n",
- "You can also pass a different region to check availability and then re-create your workspace in that region through the [00. Installation and Configuration](00.configuration.ipynb)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.compute import ComputeTarget, AmlCompute\n",
- "\n",
- "AmlCompute.supported_vmsizes(workspace = ws)\n",
- "#AmlCompute.supported_vmsizes(workspace = ws, location='southcentralus')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create project directory\n",
- "\n",
- "Create a directory that will contain all the necessary code from your local machine that you will need access to on the remote resource. This includes the training script, and any additional files your training script depends on"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import os\n",
- "import shutil\n",
- "\n",
- "project_folder = './train-on-amlcompute'\n",
- "os.makedirs(project_folder, exist_ok=True)\n",
- "shutil.copy('train.py', project_folder)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Provision as a run based compute target\n",
- "\n",
- "You can provision AmlCompute as a compute target at run-time. In this case, the compute is auto-created for your run, scales up to max_nodes that you specify, and then **deleted automatically** after the run completes."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.runconfig import RunConfiguration\n",
- "from azureml.core.conda_dependencies import CondaDependencies\n",
- "from azureml.core.runconfig import DEFAULT_CPU_IMAGE\n",
- "\n",
- "# create a new runconfig object\n",
- "run_config = RunConfiguration()\n",
- "\n",
- "# signal that you want to use AmlCompute to execute script.\n",
- "run_config.target = \"amlcompute\"\n",
- "\n",
- "# AmlCompute will be created in the same region as workspace\n",
- "# Set vm size for AmlCompute\n",
- "run_config.amlcompute.vm_size = 'STANDARD_D2_V2'\n",
- "\n",
- "# enable Docker \n",
- "run_config.environment.docker.enabled = True\n",
- "\n",
- "# set Docker base image to the default CPU-based image\n",
- "run_config.environment.docker.base_image = DEFAULT_CPU_IMAGE\n",
- "\n",
- "# use conda_dependencies.yml to create a conda environment in the Docker image for execution\n",
- "run_config.environment.python.user_managed_dependencies = False\n",
- "\n",
- "# auto-prepare the Docker image when used for execution (if it is not already prepared)\n",
- "run_config.auto_prepare_environment = True\n",
- "\n",
- "# specify CondaDependencies obj\n",
- "run_config.environment.python.conda_dependencies = CondaDependencies.create(conda_packages=['scikit-learn'])\n",
- "\n",
- "# Now submit a run on AmlCompute\n",
- "from azureml.core.script_run_config import ScriptRunConfig\n",
- "\n",
- "script_run_config = ScriptRunConfig(source_directory=project_folder,\n",
- " script='train.py',\n",
- " run_config=run_config)\n",
- "\n",
- "run = experiment.submit(script_run_config)\n",
- "\n",
- "# Show run details\n",
- "run"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%time\n",
- "# Shows output of the run on stdout.\n",
- "run.wait_for_completion(show_output=True)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run.get_metrics()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Provision as a persistent compute target (Basic)\n",
- "\n",
- "You can provision a persistent AmlCompute resource by simply defining two parameters thanks to smart defaults. By default it autoscales from 0 nodes and provisions dedicated VMs to run your job in a container. This is useful when you want to continously re-use the same target, debug it between jobs or simply share the resource with other users of your workspace.\n",
- "\n",
- "* `vm_size`: VM family of the nodes provisioned by AmlCompute. Simply choose from the supported_vmsizes() above\n",
- "* `max_nodes`: Maximum nodes to autoscale to while running a job on AmlCompute"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.compute import ComputeTarget, AmlCompute\n",
- "from azureml.core.compute_target import ComputeTargetException\n",
- "\n",
- "# Choose a name for your CPU cluster\n",
- "cpu_cluster_name = \"cpucluster\"\n",
- "\n",
- "# Verify that cluster does not exist already\n",
- "try:\n",
- " cpu_cluster = ComputeTarget(workspace=ws, name=cpu_cluster_name)\n",
- " print('Found existing cluster, use it.')\n",
- "except ComputeTargetException:\n",
- " compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_D2_V2',\n",
- " max_nodes=4)\n",
- " cpu_cluster = ComputeTarget.create(ws, cpu_cluster_name, compute_config)\n",
- "\n",
- "cpu_cluster.wait_for_completion(show_output=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Configure & Run"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.runconfig import RunConfiguration\n",
- "from azureml.core.conda_dependencies import CondaDependencies\n",
- "\n",
- "# create a new RunConfig object\n",
- "run_config = RunConfiguration(framework=\"python\")\n",
- "\n",
- "# Set compute target to AmlCompute target created in previous step\n",
- "run_config.target = cpu_cluster.name\n",
- "\n",
- "# enable Docker \n",
- "run_config.environment.docker.enabled = True\n",
- "\n",
- "# specify CondaDependencies obj\n",
- "run_config.environment.python.conda_dependencies = CondaDependencies.create(conda_packages=['scikit-learn'])\n",
- "\n",
- "from azureml.core import Run\n",
- "from azureml.core import ScriptRunConfig\n",
- "\n",
- "src = ScriptRunConfig(source_directory=project_folder, \n",
- " script='train.py', \n",
- " run_config=run_config) \n",
- "run = experiment.submit(config=src)\n",
- "run"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%time\n",
- "# Shows output of the run on stdout.\n",
- "run.wait_for_completion(show_output=True)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run.get_metrics()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Provision as a persistent compute target (Advanced)\n",
- "\n",
- "You can also specify additional properties or change defaults while provisioning AmlCompute using a more advanced configuration. This is useful when you want a dedicated cluster of 4 nodes (for example you can set the min_nodes and max_nodes to 4), or want the compute to be within an existing VNet in your subscription.\n",
- "\n",
- "In addition to `vm_size` and `max_nodes`, you can specify:\n",
- "* `min_nodes`: Minimum nodes (default 0 nodes) to downscale to while running a job on AmlCompute\n",
- "* `vm_priority`: Choose between 'dedicated' (default) and 'lowpriority' VMs when provisioning AmlCompute. Low Priority VMs use Azure's excess capacity and are thus cheaper but risk your run being pre-empted\n",
- "* `idle_seconds_before_scaledown`: Idle time (default 120 seconds) to wait after run completion before auto-scaling to min_nodes\n",
- "* `vnet_resourcegroup_name`: Resource group of the **existing** VNet within which AmlCompute should be provisioned\n",
- "* `vnet_name`: Name of VNet\n",
- "* `subnet_name`: Name of SubNet within the VNet"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.compute import ComputeTarget, AmlCompute\n",
- "from azureml.core.compute_target import ComputeTargetException\n",
- "\n",
- "# Choose a name for your CPU cluster\n",
- "cpu_cluster_name = \"cpucluster\"\n",
- "\n",
- "# Verify that cluster does not exist already\n",
- "try:\n",
- " cpu_cluster = ComputeTarget(workspace=ws, name=cpu_cluster_name)\n",
- " print('Found existing cluster, use it.')\n",
- "except ComputeTargetException:\n",
- " compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_D2_V2',\n",
- " vm_priority='lowpriority',\n",
- " min_nodes=2,\n",
- " max_nodes=4,\n",
- " idle_seconds_before_scaledown='300',\n",
- " vnet_resourcegroup_name='',\n",
- " vnet_name='',\n",
- " subnet_name='')\n",
- " cpu_cluster = ComputeTarget.create(ws, cpu_cluster_name, compute_config)\n",
- "\n",
- "cpu_cluster.wait_for_completion(show_output=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Configure & Run"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.runconfig import RunConfiguration\n",
- "from azureml.core.conda_dependencies import CondaDependencies\n",
- "\n",
- "# create a new RunConfig object\n",
- "run_config = RunConfiguration(framework=\"python\")\n",
- "\n",
- "# Set compute target to AmlCompute target created in previous step\n",
- "run_config.target = cpu_cluster.name\n",
- "\n",
- "# enable Docker \n",
- "run_config.environment.docker.enabled = True\n",
- "\n",
- "# specify CondaDependencies obj\n",
- "run_config.environment.python.conda_dependencies = CondaDependencies.create(conda_packages=['scikit-learn'])\n",
- "\n",
- "from azureml.core import Run\n",
- "from azureml.core import ScriptRunConfig\n",
- "\n",
- "src = ScriptRunConfig(source_directory=project_folder, \n",
- " script='train.py', \n",
- " run_config=run_config) \n",
- "run = experiment.submit(config=src)\n",
- "run"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%time\n",
- "# Shows output of the run on stdout.\n",
- "run.wait_for_completion(show_output=True)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run.get_metrics()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Additional operations to perform on AmlCompute\n",
- "\n",
- "You can perform more operations on AmlCompute such as updating the node counts or deleting the compute. "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "#Get_status () gets the latest status of the AmlCompute target\n",
- "cpu_cluster.get_status()\n",
- "cpu_cluster.serialize()"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "#Update () takes in the min_nodes, max_nodes and idle_seconds_before_scaledown and updates the AmlCompute target\n",
- "#cpu_cluster.update(min_nodes=1)\n",
- "#cpu_cluster.update(max_nodes=10)\n",
- "cpu_cluster.update(idle_seconds_before_scaledown=300)\n",
- "#cpu_cluster.update(min_nodes=2, max_nodes=4, idle_seconds_before_scaledown=600)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "#Delete () is used to deprovision and delete the AmlCompute target. Useful if you want to re-use the compute name \n",
- "#'cpucluster' in this case but use a different VM family for instance.\n",
- "\n",
- "#cpu_cluster.delete()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Success!\n",
- "Great, you are ready to move on to the remaining notebooks."
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "nigup"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Train using Azure Machine Learning Compute\n",
+ "\n",
+ "* Initialize a Workspace\n",
+ "* Create an Experiment\n",
+ "* Introduction to AmlCompute\n",
+ "* Submit an AmlCompute run in a few different ways\n",
+ " - Provision as a run based compute target \n",
+ " - Provision as a persistent compute target (Basic)\n",
+ " - Provision as a persistent compute target (Advanced)\n",
+ "* Additional operations to perform on AmlCompute\n",
+ "* Find the best model in the run"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Prerequisites\n",
+ "Make sure you go through the [configuration notebook](../../../configuration.ipynb) first if you haven't."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Check core SDK version number\n",
+ "import azureml.core\n",
+ "\n",
+ "print(\"SDK version:\", azureml.core.VERSION)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Initialize a Workspace\n",
+ "\n",
+ "Initialize a workspace object from persisted configuration"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "create workspace"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "from azureml.core import Workspace\n",
+ "\n",
+ "ws = Workspace.from_config()\n",
+ "print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep = '\\n')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Create An Experiment\n",
+ "\n",
+ "**Experiment** is a logical container in an Azure ML Workspace. It hosts run records which can include run metrics and output artifacts from your experiments."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core import Experiment\n",
+ "experiment_name = 'train-on-amlcompute'\n",
+ "experiment = Experiment(workspace = ws, name = experiment_name)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Introduction to AmlCompute\n",
+ "\n",
+ "Azure Machine Learning Compute is managed compute infrastructure that allows the user to easily create single to multi-node compute of the appropriate VM Family. It is created **within your workspace region** and is a resource that can be used by other users in your workspace. It autoscales by default to the max_nodes, when a job is submitted, and executes in a containerized environment packaging the dependencies as specified by the user. \n",
+ "\n",
+ "Since it is managed compute, job scheduling and cluster management are handled internally by Azure Machine Learning service. \n",
+ "\n",
+ "For more information on Azure Machine Learning Compute, please read [this article](https://docs.microsoft.com/azure/machine-learning/service/how-to-set-up-training-targets#amlcompute)\n",
+ "\n",
+ "If you are an existing BatchAI customer who is migrating to Azure Machine Learning, please read [this article](https://aka.ms/batchai-retirement)\n",
+ "\n",
+ "**Note**: As with other Azure services, there are limits on certain resources (for eg. AmlCompute quota) associated with the Azure Machine Learning service. Please read [this article](https://docs.microsoft.com/azure/machine-learning/service/how-to-manage-quotas) on the default limits and how to request more quota.\n",
+ "\n",
+ "\n",
+ "The training script `train.py` is already created for you. Let's have a look."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Submit an AmlCompute run in a few different ways\n",
+ "\n",
+ "First lets check which VM families are available in your region. Azure is a regional service and some specialized SKUs (especially GPUs) are only available in certain regions. Since AmlCompute is created in the region of your workspace, we will use the supported_vms () function to see if the VM family we want to use ('STANDARD_D2_V2') is supported.\n",
+ "\n",
+ "You can also pass a different region to check availability and then re-create your workspace in that region through the [configuration notebook](../../../configuration.ipynb)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.compute import ComputeTarget, AmlCompute\n",
+ "\n",
+ "AmlCompute.supported_vmsizes(workspace = ws)\n",
+ "#AmlCompute.supported_vmsizes(workspace = ws, location='southcentralus')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create project directory\n",
+ "\n",
+ "Create a directory that will contain all the necessary code from your local machine that you will need access to on the remote resource. This includes the training script, and any additional files your training script depends on"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import os\n",
+ "import shutil\n",
+ "\n",
+ "project_folder = './train-on-amlcompute'\n",
+ "os.makedirs(project_folder, exist_ok=True)\n",
+ "shutil.copy('train.py', project_folder)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Provision as a run based compute target\n",
+ "\n",
+ "You can provision AmlCompute as a compute target at run-time. In this case, the compute is auto-created for your run, scales up to max_nodes that you specify, and then **deleted automatically** after the run completes."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.runconfig import RunConfiguration\n",
+ "from azureml.core.conda_dependencies import CondaDependencies\n",
+ "from azureml.core.runconfig import DEFAULT_CPU_IMAGE\n",
+ "\n",
+ "# create a new runconfig object\n",
+ "run_config = RunConfiguration()\n",
+ "\n",
+ "# signal that you want to use AmlCompute to execute script.\n",
+ "run_config.target = \"amlcompute\"\n",
+ "\n",
+ "# AmlCompute will be created in the same region as workspace\n",
+ "# Set vm size for AmlCompute\n",
+ "run_config.amlcompute.vm_size = 'STANDARD_D2_V2'\n",
+ "\n",
+ "# enable Docker \n",
+ "run_config.environment.docker.enabled = True\n",
+ "\n",
+ "# set Docker base image to the default CPU-based image\n",
+ "run_config.environment.docker.base_image = DEFAULT_CPU_IMAGE\n",
+ "\n",
+ "# use conda_dependencies.yml to create a conda environment in the Docker image for execution\n",
+ "run_config.environment.python.user_managed_dependencies = False\n",
+ "\n",
+ "# auto-prepare the Docker image when used for execution (if it is not already prepared)\n",
+ "run_config.auto_prepare_environment = True\n",
+ "\n",
+ "# specify CondaDependencies obj\n",
+ "run_config.environment.python.conda_dependencies = CondaDependencies.create(conda_packages=['scikit-learn'])\n",
+ "\n",
+ "# Now submit a run on AmlCompute\n",
+ "from azureml.core.script_run_config import ScriptRunConfig\n",
+ "\n",
+ "script_run_config = ScriptRunConfig(source_directory=project_folder,\n",
+ " script='train.py',\n",
+ " run_config=run_config)\n",
+ "\n",
+ "run = experiment.submit(script_run_config)\n",
+ "\n",
+ "# Show run details\n",
+ "run"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%time\n",
+ "# Shows output of the run on stdout.\n",
+ "run.wait_for_completion(show_output=True)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run.get_metrics()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Provision as a persistent compute target (Basic)\n",
+ "\n",
+ "You can provision a persistent AmlCompute resource by simply defining two parameters thanks to smart defaults. By default it autoscales from 0 nodes and provisions dedicated VMs to run your job in a container. This is useful when you want to continously re-use the same target, debug it between jobs or simply share the resource with other users of your workspace.\n",
+ "\n",
+ "* `vm_size`: VM family of the nodes provisioned by AmlCompute. Simply choose from the supported_vmsizes() above\n",
+ "* `max_nodes`: Maximum nodes to autoscale to while running a job on AmlCompute"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.compute import ComputeTarget, AmlCompute\n",
+ "from azureml.core.compute_target import ComputeTargetException\n",
+ "\n",
+ "# Choose a name for your CPU cluster\n",
+ "cpu_cluster_name = \"cpucluster\"\n",
+ "\n",
+ "# Verify that cluster does not exist already\n",
+ "try:\n",
+ " cpu_cluster = ComputeTarget(workspace=ws, name=cpu_cluster_name)\n",
+ " print('Found existing cluster, use it.')\n",
+ "except ComputeTargetException:\n",
+ " compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_D2_V2',\n",
+ " max_nodes=4)\n",
+ " cpu_cluster = ComputeTarget.create(ws, cpu_cluster_name, compute_config)\n",
+ "\n",
+ "cpu_cluster.wait_for_completion(show_output=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Configure & Run"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.runconfig import RunConfiguration\n",
+ "from azureml.core.conda_dependencies import CondaDependencies\n",
+ "\n",
+ "# create a new RunConfig object\n",
+ "run_config = RunConfiguration(framework=\"python\")\n",
+ "\n",
+ "# Set compute target to AmlCompute target created in previous step\n",
+ "run_config.target = cpu_cluster.name\n",
+ "\n",
+ "# enable Docker \n",
+ "run_config.environment.docker.enabled = True\n",
+ "\n",
+ "# specify CondaDependencies obj\n",
+ "run_config.environment.python.conda_dependencies = CondaDependencies.create(conda_packages=['scikit-learn'])\n",
+ "\n",
+ "from azureml.core import Run\n",
+ "from azureml.core import ScriptRunConfig\n",
+ "\n",
+ "src = ScriptRunConfig(source_directory=project_folder, \n",
+ " script='train.py', \n",
+ " run_config=run_config) \n",
+ "run = experiment.submit(config=src)\n",
+ "run"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%time\n",
+ "# Shows output of the run on stdout.\n",
+ "run.wait_for_completion(show_output=True)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run.get_metrics()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Provision as a persistent compute target (Advanced)\n",
+ "\n",
+ "You can also specify additional properties or change defaults while provisioning AmlCompute using a more advanced configuration. This is useful when you want a dedicated cluster of 4 nodes (for example you can set the min_nodes and max_nodes to 4), or want the compute to be within an existing VNet in your subscription.\n",
+ "\n",
+ "In addition to `vm_size` and `max_nodes`, you can specify:\n",
+ "* `min_nodes`: Minimum nodes (default 0 nodes) to downscale to while running a job on AmlCompute\n",
+ "* `vm_priority`: Choose between 'dedicated' (default) and 'lowpriority' VMs when provisioning AmlCompute. Low Priority VMs use Azure's excess capacity and are thus cheaper but risk your run being pre-empted\n",
+ "* `idle_seconds_before_scaledown`: Idle time (default 120 seconds) to wait after run completion before auto-scaling to min_nodes\n",
+ "* `vnet_resourcegroup_name`: Resource group of the **existing** VNet within which AmlCompute should be provisioned\n",
+ "* `vnet_name`: Name of VNet\n",
+ "* `subnet_name`: Name of SubNet within the VNet"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.compute import ComputeTarget, AmlCompute\n",
+ "from azureml.core.compute_target import ComputeTargetException\n",
+ "\n",
+ "# Choose a name for your CPU cluster\n",
+ "cpu_cluster_name = \"cpucluster\"\n",
+ "\n",
+ "# Verify that cluster does not exist already\n",
+ "try:\n",
+ " cpu_cluster = ComputeTarget(workspace=ws, name=cpu_cluster_name)\n",
+ " print('Found existing cluster, use it.')\n",
+ "except ComputeTargetException:\n",
+ " compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_D2_V2',\n",
+ " vm_priority='lowpriority',\n",
+ " min_nodes=2,\n",
+ " max_nodes=4,\n",
+ " idle_seconds_before_scaledown='300',\n",
+ " vnet_resourcegroup_name='',\n",
+ " vnet_name='',\n",
+ " subnet_name='')\n",
+ " cpu_cluster = ComputeTarget.create(ws, cpu_cluster_name, compute_config)\n",
+ "\n",
+ "cpu_cluster.wait_for_completion(show_output=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Configure & Run"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.runconfig import RunConfiguration\n",
+ "from azureml.core.conda_dependencies import CondaDependencies\n",
+ "\n",
+ "# create a new RunConfig object\n",
+ "run_config = RunConfiguration(framework=\"python\")\n",
+ "\n",
+ "# Set compute target to AmlCompute target created in previous step\n",
+ "run_config.target = cpu_cluster.name\n",
+ "\n",
+ "# enable Docker \n",
+ "run_config.environment.docker.enabled = True\n",
+ "\n",
+ "# specify CondaDependencies obj\n",
+ "run_config.environment.python.conda_dependencies = CondaDependencies.create(conda_packages=['scikit-learn'])\n",
+ "\n",
+ "from azureml.core import Run\n",
+ "from azureml.core import ScriptRunConfig\n",
+ "\n",
+ "src = ScriptRunConfig(source_directory=project_folder, \n",
+ " script='train.py', \n",
+ " run_config=run_config) \n",
+ "run = experiment.submit(config=src)\n",
+ "run"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%time\n",
+ "# Shows output of the run on stdout.\n",
+ "run.wait_for_completion(show_output=True)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run.get_metrics()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Additional operations to perform on AmlCompute\n",
+ "\n",
+ "You can perform more operations on AmlCompute such as updating the node counts or deleting the compute. "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "#Get_status () gets the latest status of the AmlCompute target\n",
+ "cpu_cluster.get_status()\n",
+ "cpu_cluster.serialize()"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "#Update () takes in the min_nodes, max_nodes and idle_seconds_before_scaledown and updates the AmlCompute target\n",
+ "#cpu_cluster.update(min_nodes=1)\n",
+ "#cpu_cluster.update(max_nodes=10)\n",
+ "cpu_cluster.update(idle_seconds_before_scaledown=300)\n",
+ "#cpu_cluster.update(min_nodes=2, max_nodes=4, idle_seconds_before_scaledown=600)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "#Delete () is used to deprovision and delete the AmlCompute target. Useful if you want to re-use the compute name \n",
+ "#'cpucluster' in this case but use a different VM family for instance.\n",
+ "\n",
+ "#cpu_cluster.delete()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Success!\n",
+ "Great, you are ready to move on to the remaining notebooks."
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "nigup"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.6"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/training/train-on-local/train-on-local.ipynb b/how-to-use-azureml/training/train-on-local/train-on-local.ipynb
index a7aa7028..b4b0ffa8 100644
--- a/how-to-use-azureml/training/train-on-local/train-on-local.ipynb
+++ b/how-to-use-azureml/training/train-on-local/train-on-local.ipynb
@@ -1,478 +1,478 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# 02. Train locally\n",
- "* Create or load workspace.\n",
- "* Create scripts locally.\n",
- "* Create `train.py` in a folder, along with a `my.lib` file.\n",
- "* Configure & execute a local run in a user-managed Python environment.\n",
- "* Configure & execute a local run in a system-managed Python environment.\n",
- "* Configure & execute a local run in a Docker environment.\n",
- "* Query run metrics to find the best model\n",
- "* Register model for operationalization."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Prerequisites\n",
- "Make sure you go through the [00. Installation and Configuration](00.configuration.ipynb) Notebook first if you haven't."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Check core SDK version number\n",
- "import azureml.core\n",
- "\n",
- "print(\"SDK version:\", azureml.core.VERSION)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Initialize Workspace\n",
- "\n",
- "Initialize a workspace object from persisted configuration."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.workspace import Workspace\n",
- "\n",
- "ws = Workspace.from_config()\n",
- "print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep='\\n')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Create An Experiment\n",
- "**Experiment** is a logical container in an Azure ML Workspace. It hosts run records which can include run metrics and output artifacts from your experiments."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core import Experiment\n",
- "experiment_name = 'train-on-local'\n",
- "exp = Experiment(workspace=ws, name=experiment_name)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## View `train.py`\n",
- "\n",
- "`train.py` is already created for you."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "with open('./train.py', 'r') as f:\n",
- " print(f.read())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Note `train.py` also references a `mylib.py` file."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "with open('./mylib.py', 'r') as f:\n",
- " print(f.read())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Configure & Run\n",
- "### User-managed environment\n",
- "Below, we use a user-managed run, which means you are responsible to ensure all the necessary packages are available in the Python environment you choose to run the script."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.runconfig import RunConfiguration\n",
- "\n",
- "# Editing a run configuration property on-fly.\n",
- "run_config_user_managed = RunConfiguration()\n",
- "\n",
- "run_config_user_managed.environment.python.user_managed_dependencies = True\n",
- "\n",
- "# You can choose a specific Python environment by pointing to a Python path \n",
- "#run_config.environment.python.interpreter_path = '/home/johndoe/miniconda3/envs/sdk2/bin/python'"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Submit script to run in the user-managed environment\n",
- "Note whole script folder is submitted for execution, including the `mylib.py` file."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core import ScriptRunConfig\n",
- "\n",
- "src = ScriptRunConfig(source_directory='./', script='train.py', run_config=run_config_user_managed)\n",
- "run = exp.submit(src)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Get run history details"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Block to wait till run finishes."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run.wait_for_completion(show_output=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### System-managed environment\n",
- "You can also ask the system to build a new conda environment and execute your scripts in it. The environment is built once and will be reused in subsequent executions as long as the conda dependencies remain unchanged. "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.runconfig import RunConfiguration\n",
- "from azureml.core.conda_dependencies import CondaDependencies\n",
- "\n",
- "run_config_system_managed = RunConfiguration()\n",
- "\n",
- "run_config_system_managed.environment.python.user_managed_dependencies = False\n",
- "run_config_system_managed.auto_prepare_environment = True\n",
- "\n",
- "# Specify conda dependencies with scikit-learn\n",
- "cd = CondaDependencies.create(conda_packages=['scikit-learn'])\n",
- "run_config_system_managed.environment.python.conda_dependencies = cd"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Submit script to run in the system-managed environment\n",
- "A new conda environment is built based on the conda dependencies object. If you are running this for the first time, this might take up to 5 mninutes. But this conda environment is reused so long as you don't change the conda dependencies."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "src = ScriptRunConfig(source_directory=\"./\", script='train.py', run_config=run_config_system_managed)\n",
- "run = exp.submit(src)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "#### Get run history details"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Block and wait till run finishes."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run.wait_for_completion(show_output = True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Docker-based execution\n",
- "**IMPORTANT**: You must have Docker engine installed locally in order to use this execution mode. If your kernel is already running in a Docker container, such as **Azure Notebooks**, this mode will **NOT** work.\n",
- "NOTE: The GPU base image must be used on Microsoft Azure Services only such as ACI, AML Compute, Azure VMs, and AKS.\n",
- "\n",
- "You can also ask the system to pull down a Docker image and execute your scripts in it."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run_config_docker = RunConfiguration()\n",
- "run_config_docker.environment.python.user_managed_dependencies = False\n",
- "run_config_docker.auto_prepare_environment = True\n",
- "run_config_docker.environment.docker.enabled = True\n",
- "run_config_docker.environment.docker.base_image = azureml.core.runconfig.DEFAULT_CPU_IMAGE\n",
- "\n",
- "# Specify conda dependencies with scikit-learn\n",
- "cd = CondaDependencies.create(conda_packages=['scikit-learn'])\n",
- "run_config_docker.environment.python.conda_dependencies = cd\n",
- "\n",
- "src = ScriptRunConfig(source_directory=\"./\", script='train.py', run_config=run_config_docker)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Submit script to run in the system-managed environment\n",
- "A new conda environment is built based on the conda dependencies object. If you are running this for the first time, this might take up to 5 mninutes. But this conda environment is reused so long as you don't change the conda dependencies.\n",
- "\n",
- "\n"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import subprocess\n",
- "\n",
- "# Check if Docker is installed and Linux containers are enables\n",
- "if subprocess.run(\"docker -v\", shell=True) == 0:\n",
- " out = subprocess.check_output(\"docker system info\", shell=True, encoding=\"ascii\").split(\"\\n\")\n",
- " if not \"OSType: linux\" in out:\n",
- " print(\"Switch Docker engine to use Linux containers.\")\n",
- " else:\n",
- " run = exp.submit(src)\n",
- "else:\n",
- " print(\"Docker engine not installed.\")"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "#Get run history details\n",
- "run"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run.wait_for_completion(show_output=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Query run metrics"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "query history",
- "get metrics"
- ]
- },
- "outputs": [],
- "source": [
- "# get all metris logged in the run\n",
- "run.get_metrics()\n",
- "metrics = run.get_metrics()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Let's find the model that has the lowest MSE value logged."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import numpy as np\n",
- "\n",
- "best_alpha = metrics['alpha'][np.argmin(metrics['mse'])]\n",
- "\n",
- "print('When alpha is {1:0.2f}, we have min MSE {0:0.2f}.'.format(\n",
- " min(metrics['mse']), \n",
- " best_alpha\n",
- "))"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "You can also list all the files that are associated with this run record"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run.get_file_names()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "We know the model `ridge_0.40.pkl` is the best performing model from the eariler queries. So let's register it with the workspace."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# supply a model name, and the full path to the serialized model file.\n",
- "model = run.register_model(model_name='best_ridge_model', model_path='./outputs/ridge_0.40.pkl')"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "print(model.name, model.version, model.url)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Now you can deploy this model following the example in the 01 notebook."
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "roastala"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# 02. Train locally\n",
+ "* Create or load workspace.\n",
+ "* Create scripts locally.\n",
+ "* Create `train.py` in a folder, along with a `my.lib` file.\n",
+ "* Configure & execute a local run in a user-managed Python environment.\n",
+ "* Configure & execute a local run in a system-managed Python environment.\n",
+ "* Configure & execute a local run in a Docker environment.\n",
+ "* Query run metrics to find the best model\n",
+ "* Register model for operationalization."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Prerequisites\n",
+ "Make sure you go through the [configuration notebook](../../../configuration.ipynb) first if you haven't."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Check core SDK version number\n",
+ "import azureml.core\n",
+ "\n",
+ "print(\"SDK version:\", azureml.core.VERSION)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Initialize Workspace\n",
+ "\n",
+ "Initialize a workspace object from persisted configuration."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.workspace import Workspace\n",
+ "\n",
+ "ws = Workspace.from_config()\n",
+ "print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep='\\n')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Create An Experiment\n",
+ "**Experiment** is a logical container in an Azure ML Workspace. It hosts run records which can include run metrics and output artifacts from your experiments."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core import Experiment\n",
+ "experiment_name = 'train-on-local'\n",
+ "exp = Experiment(workspace=ws, name=experiment_name)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## View `train.py`\n",
+ "\n",
+ "`train.py` is already created for you."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "with open('./train.py', 'r') as f:\n",
+ " print(f.read())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Note `train.py` also references a `mylib.py` file."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "with open('./mylib.py', 'r') as f:\n",
+ " print(f.read())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Configure & Run\n",
+ "### User-managed environment\n",
+ "Below, we use a user-managed run, which means you are responsible to ensure all the necessary packages are available in the Python environment you choose to run the script."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.runconfig import RunConfiguration\n",
+ "\n",
+ "# Editing a run configuration property on-fly.\n",
+ "run_config_user_managed = RunConfiguration()\n",
+ "\n",
+ "run_config_user_managed.environment.python.user_managed_dependencies = True\n",
+ "\n",
+ "# You can choose a specific Python environment by pointing to a Python path \n",
+ "#run_config.environment.python.interpreter_path = '/home/johndoe/miniconda3/envs/sdk2/bin/python'"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Submit script to run in the user-managed environment\n",
+ "Note whole script folder is submitted for execution, including the `mylib.py` file."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core import ScriptRunConfig\n",
+ "\n",
+ "src = ScriptRunConfig(source_directory='./', script='train.py', run_config=run_config_user_managed)\n",
+ "run = exp.submit(src)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Get run history details"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Block to wait till run finishes."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run.wait_for_completion(show_output=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### System-managed environment\n",
+ "You can also ask the system to build a new conda environment and execute your scripts in it. The environment is built once and will be reused in subsequent executions as long as the conda dependencies remain unchanged. "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.runconfig import RunConfiguration\n",
+ "from azureml.core.conda_dependencies import CondaDependencies\n",
+ "\n",
+ "run_config_system_managed = RunConfiguration()\n",
+ "\n",
+ "run_config_system_managed.environment.python.user_managed_dependencies = False\n",
+ "run_config_system_managed.auto_prepare_environment = True\n",
+ "\n",
+ "# Specify conda dependencies with scikit-learn\n",
+ "cd = CondaDependencies.create(conda_packages=['scikit-learn'])\n",
+ "run_config_system_managed.environment.python.conda_dependencies = cd"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Submit script to run in the system-managed environment\n",
+ "A new conda environment is built based on the conda dependencies object. If you are running this for the first time, this might take up to 5 mninutes. But this conda environment is reused so long as you don't change the conda dependencies."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "src = ScriptRunConfig(source_directory=\"./\", script='train.py', run_config=run_config_system_managed)\n",
+ "run = exp.submit(src)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "#### Get run history details"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Block and wait till run finishes."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run.wait_for_completion(show_output = True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Docker-based execution\n",
+ "**IMPORTANT**: You must have Docker engine installed locally in order to use this execution mode. If your kernel is already running in a Docker container, such as **Azure Notebooks**, this mode will **NOT** work.\n",
+ "NOTE: The GPU base image must be used on Microsoft Azure Services only such as ACI, AML Compute, Azure VMs, and AKS.\n",
+ "\n",
+ "You can also ask the system to pull down a Docker image and execute your scripts in it."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run_config_docker = RunConfiguration()\n",
+ "run_config_docker.environment.python.user_managed_dependencies = False\n",
+ "run_config_docker.auto_prepare_environment = True\n",
+ "run_config_docker.environment.docker.enabled = True\n",
+ "run_config_docker.environment.docker.base_image = azureml.core.runconfig.DEFAULT_CPU_IMAGE\n",
+ "\n",
+ "# Specify conda dependencies with scikit-learn\n",
+ "cd = CondaDependencies.create(conda_packages=['scikit-learn'])\n",
+ "run_config_docker.environment.python.conda_dependencies = cd\n",
+ "\n",
+ "src = ScriptRunConfig(source_directory=\"./\", script='train.py', run_config=run_config_docker)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Submit script to run in the system-managed environment\n",
+ "A new conda environment is built based on the conda dependencies object. If you are running this for the first time, this might take up to 5 mninutes. But this conda environment is reused so long as you don't change the conda dependencies.\n",
+ "\n",
+ "\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import subprocess\n",
+ "\n",
+ "# Check if Docker is installed and Linux containers are enables\n",
+ "if subprocess.run(\"docker -v\", shell=True) == 0:\n",
+ " out = subprocess.check_output(\"docker system info\", shell=True, encoding=\"ascii\").split(\"\\n\")\n",
+ " if not \"OSType: linux\" in out:\n",
+ " print(\"Switch Docker engine to use Linux containers.\")\n",
+ " else:\n",
+ " run = exp.submit(src)\n",
+ "else:\n",
+ " print(\"Docker engine not installed.\")"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "#Get run history details\n",
+ "run"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run.wait_for_completion(show_output=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Query run metrics"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "query history",
+ "get metrics"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "# get all metris logged in the run\n",
+ "run.get_metrics()\n",
+ "metrics = run.get_metrics()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Let's find the model that has the lowest MSE value logged."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import numpy as np\n",
+ "\n",
+ "best_alpha = metrics['alpha'][np.argmin(metrics['mse'])]\n",
+ "\n",
+ "print('When alpha is {1:0.2f}, we have min MSE {0:0.2f}.'.format(\n",
+ " min(metrics['mse']), \n",
+ " best_alpha\n",
+ "))"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "You can also list all the files that are associated with this run record"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run.get_file_names()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "We know the model `ridge_0.40.pkl` is the best performing model from the eariler queries. So let's register it with the workspace."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# supply a model name, and the full path to the serialized model file.\n",
+ "model = run.register_model(model_name='best_ridge_model', model_path='./outputs/ridge_0.40.pkl')"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "print(model.name, model.version, model.url)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Now you can deploy this model following the example in the 01 notebook."
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "roastala"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.6"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/training/train-on-remote-vm/train-on-remote-vm.ipynb b/how-to-use-azureml/training/train-on-remote-vm/train-on-remote-vm.ipynb
index f0e76156..7c455d98 100644
--- a/how-to-use-azureml/training/train-on-remote-vm/train-on-remote-vm.ipynb
+++ b/how-to-use-azureml/training/train-on-remote-vm/train-on-remote-vm.ipynb
@@ -1,613 +1,607 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# 04. Train in a remote Linux VM\n",
- "* Create Workspace\n",
- "* Create `train.py` file\n",
- "* Create and Attach a Remote VM (eg. DSVM) as compute resource.\n",
- "* Upoad data files into default datastore\n",
- "* Configure & execute a run in a few different ways\n",
- " - Use system-built conda\n",
- " - Use existing Python environment\n",
- " - Use Docker \n",
- "* Find the best model in the run"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Prerequisites\n",
- "Make sure you go through the [00. Installation and Configuration](00.configuration.ipynb) Notebook first if you haven't."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# Check core SDK version number\n",
- "import azureml.core\n",
- "\n",
- "print(\"SDK version:\", azureml.core.VERSION)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Initialize Workspace\n",
- "\n",
- "Initialize a workspace object from persisted configuration."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core import Workspace\n",
- "\n",
- "ws = Workspace.from_config()\n",
- "print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep='\\n')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Create Experiment\n",
- "\n",
- "**Experiment** is a logical container in an Azure ML Workspace. It hosts run records which can include run metrics and output artifacts from your experiments."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "experiment_name = 'train-on-remote-vm'\n",
- "\n",
- "from azureml.core import Experiment\n",
- "exp = Experiment(workspace=ws, name=experiment_name)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Let's also create a local folder to hold the training script."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import os\n",
- "script_folder = './vm-run'\n",
- "os.makedirs(script_folder, exist_ok=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Upload data files into datastore\n",
- "Every workspace comes with a default datastore (and you can register more) which is backed by the Azure blob storage account associated with the workspace. We can use it to transfer data from local to the cloud, and access it from the compute target."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# get the default datastore\n",
- "ds = ws.get_default_datastore()\n",
- "print(ds.name, ds.datastore_type, ds.account_name, ds.container_name)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Load diabetes data from `scikit-learn` and save it as 2 local files."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from sklearn.datasets import load_diabetes\n",
- "import numpy as np\n",
- "\n",
- "training_data = load_diabetes()\n",
- "np.save(file='./features.npy', arr=training_data['data'])\n",
- "np.save(file='./labels.npy', arr=training_data['target'])"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Now let's upload the 2 files into the default datastore under a path named `diabetes`:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "ds.upload_files(['./features.npy', './labels.npy'], target_path='diabetes', overwrite=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## View `train.py`\n",
- "\n",
- "For convenience, we created a training script for you. It is printed below as a text, but you can also run `%pfile ./train.py` in a cell to show the file. Please pay special attention on how we are loading the features and labels from files in the `data_folder` path, which is passed in as an argument of the training script (shown later)."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# copy train.py into the script folder\n",
- "import shutil\n",
- "shutil.copy('./train.py', os.path.join(script_folder, 'train.py'))\n",
- "\n",
- "with open(os.path.join(script_folder, './train.py'), 'r') as training_script:\n",
- " print(training_script.read())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Create and Attach a DSVM as a compute target\n",
- "\n",
- "**Note**: To streamline the compute that Azure Machine Learning creates, we are making updates to support creating only single to multi-node `AmlCompute`. The `DSVMCompute` class will be deprecated in a later release, but the DSVM can be created using the below single line command and then attached(like any VM) using the sample code below. Also note, that we only support Linux VMs for remote execution from AML and the commands below will spin a Linux VM only.\n",
- "\n",
- "```shell\n",
- "# create a DSVM in your resource group\n",
- "# note you need to be at least a contributor to the resource group in order to execute this command successfully\n",
- "(myenv) $ az vm create --resource-group --name --image microsoft-dsvm:linux-data-science-vm-ubuntu:linuxdsvmubuntu:latest --admin-username --admin-password --generate-ssh-keys --authentication-type password\n",
- "```\n",
- "\n",
- "**Note**: You can also use [this url](https://portal.azure.com/#create/microsoft-dsvm.linux-data-science-vm-ubuntulinuxdsvmubuntu) to create the VM using the Azure Portal\n",
- "\n",
- "**Note**: By default SSH runs on port 22 and you don't need to specify it. But if for security reasons you switch to a different port (such as 5022), you can specify the port number in the provisioning configuration object."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.compute import RemoteCompute\n",
- "from azureml.core.compute_target import ComputeTargetException\n",
- "import os\n",
- "\n",
- "username = os.getenv('AZUREML_DSVM_USERNAME', default='')\n",
- "address = os.getenv('AZUREML_DSVM_ADDRESS', default='')\n",
- "\n",
- "compute_target_name = 'cpudsvm'\n",
- "# if you want to connect using SSH key instead of username/password you can provide parameters private_key_file and private_key_passphrase \n",
- "try:\n",
- " attached_dsvm_compute = RemoteCompute(workspace=ws, name=compute_target_name)\n",
- " print('found existing:', attached_dsvm_compute.name)\n",
- "except ComputeTargetException:\n",
- " attached_dsvm_compute = RemoteCompute.attach(workspace=ws,\n",
- " name=compute_target_name,\n",
- " username=username,\n",
- " address=address,\n",
- " ssh_port=22,\n",
- " private_key_file='./.ssh/id_rsa')\n",
- " \n",
- " attached_dsvm_compute.wait_for_completion(show_output=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Configure & Run\n",
- "First let's create a `DataReferenceConfiguration` object to inform the system what data folder to download to the copmute target."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.runconfig import DataReferenceConfiguration\n",
- "dr = DataReferenceConfiguration(datastore_name=ds.name, \n",
- " path_on_datastore='diabetes', \n",
- " mode='download', # download files from datastore to compute target\n",
- " overwrite=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Now we can try a few different ways to run the training script in the VM."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Conda run\n",
- "You can ask the system to build a conda environment based on your dependency specification, and submit your script to run there. Once the environment is built, and if you don't change your dependencies, it will be reused in subsequent runs."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.runconfig import RunConfiguration\n",
- "from azureml.core.conda_dependencies import CondaDependencies\n",
- "\n",
- "# create a new RunConfig object\n",
- "conda_run_config = RunConfiguration(framework=\"python\")\n",
- "\n",
- "# Set compute target to the Linux DSVM\n",
- "conda_run_config.target = attached_dsvm_compute.name\n",
- "\n",
- "# set the data reference of the run configuration\n",
- "conda_run_config.data_references = {ds.name: dr}\n",
- "\n",
- "# specify CondaDependencies obj\n",
- "conda_run_config.environment.python.conda_dependencies = CondaDependencies.create(conda_packages=['scikit-learn'])"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core import Run\n",
- "from azureml.core import ScriptRunConfig\n",
- "\n",
- "src = ScriptRunConfig(source_directory=script_folder, \n",
- " script='train.py', \n",
- " run_config=conda_run_config, \n",
- " # pass the datastore reference as a parameter to the training script\n",
- " arguments=['--data-folder', str(ds.as_download())] \n",
- " ) \n",
- "run = exp.submit(config=src)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run.wait_for_completion(show_output=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Show the run object. You can navigate to the Azure portal to see detailed information about the run."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Native VM run\n",
- "You can also configure to use an exiting Python environment in the VM to execute the script without asking the system to create a conda environment for you."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# create a new RunConfig object\n",
- "vm_run_config = RunConfiguration(framework=\"python\")\n",
- "\n",
- "# Set compute target to the Linux DSVM\n",
- "vm_run_config.target = attached_dsvm_compute.name\n",
- "\n",
- "# set the data reference of the run coonfiguration\n",
- "conda_run_config.data_references = {ds.name: dr}\n",
- "\n",
- "# Let system know that you will configure the Python environment yourself.\n",
- "vm_run_config.environment.python.user_managed_dependencies = True"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "The below run will likely fail because `train.py` needs dependency `azureml`, `scikit-learn` and others, which are not found in that Python environment. "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "src = ScriptRunConfig(source_directory=script_folder, \n",
- " script='train.py', \n",
- " run_config=vm_run_config,\n",
- " # pass the datastore reference as a parameter to the training script\n",
- " arguments=['--data-folder', str(ds.as_download())])\n",
- "run = exp.submit(config=src)\n",
- "run.wait_for_completion(show_output=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "You can choose to SSH into the VM and install Azure ML SDK, and any other missing dependencies, in that Python environment. For demonstration purposes, we simply are going to create another script `train2.py` that doesn't have azureml dependencies, and submit it instead."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%writefile $script_folder/train2.py\n",
- "\n",
- "print('####################################')\n",
- "print('Hello World (without Azure ML SDK)!')\n",
- "print('####################################')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Now let's try again. And this time it should work fine."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "src = ScriptRunConfig(source_directory=script_folder, \n",
- " script='train2.py', \n",
- " run_config=vm_run_config)\n",
- "run = exp.submit(config=src)\n",
- "run.wait_for_completion(show_output=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Note even in this case you get a run record with some basic statistics."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Configure a Docker run with new conda environment on the VM\n",
- "You can execute in a Docker container in the VM. If you choose this option, the system will pull down a base Docker image, build a new conda environment in it if you ask for (you can also skip this if you are using a customer Docker image when a preconfigured Python environment), start a container, and run your script in there. This image is also uploaded into your ACR (Azure Container Registry) assoicated with your workspace, an reused if your dependencies don't change in the subsequent runs."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.runconfig import RunConfiguration\n",
- "from azureml.core.conda_dependencies import CondaDependencies\n",
- "\n",
- "\n",
- "# Load the \"cpu-dsvm.runconfig\" file (created by the above attach operation) in memory\n",
- "docker_run_config = RunConfiguration(framework=\"python\")\n",
- "\n",
- "# Set compute target to the Linux DSVM\n",
- "docker_run_config.target = attached_dsvm_compute.name\n",
- "\n",
- "# Use Docker in the remote VM\n",
- "docker_run_config.environment.docker.enabled = True\n",
- "\n",
- "# Use CPU base image from DockerHub\n",
- "docker_run_config.environment.docker.base_image = azureml.core.runconfig.DEFAULT_CPU_IMAGE\n",
- "print('Base Docker image is:', docker_run_config.environment.docker.base_image)\n",
- "\n",
- "# set the data reference of the run coonfiguration\n",
- "docker_run_config.data_references = {ds.name: dr}\n",
- "\n",
- "# specify CondaDependencies obj\n",
- "docker_run_config.environment.python.conda_dependencies = CondaDependencies.create(conda_packages=['scikit-learn'])"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Submit the Experiment\n",
- "Submit script to run in the Docker image in the remote VM. If you run this for the first time, the system will download the base image, layer in packages specified in the `conda_dependencies.yml` file on top of the base image, create a container and then execute the script in the container."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "src = ScriptRunConfig(source_directory=script_folder, \n",
- " script='train.py', \n",
- " run_config=docker_run_config,\n",
- " # pass the datastore reference as a parameter to the training script\n",
- " arguments=['--data-folder', str(ds.as_download())])\n",
- "run = exp.submit(config=src)"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run.wait_for_completion(show_output=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### View run history details"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Find the best model"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Now we have tried various execution modes, we can find the best model from the last run."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# get all metris logged in the run\n",
- "run.get_metrics()\n",
- "metrics = run.get_metrics()"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# find the index where MSE is the smallest\n",
- "indices = list(range(0, len(metrics['mse'])))\n",
- "min_mse_index = min(indices, key=lambda x: metrics['mse'][x])\n",
- "\n",
- "print('When alpha is {1:0.2f}, we have min MSE {0:0.2f}.'.format(\n",
- " metrics['mse'][min_mse_index], \n",
- " metrics['alpha'][min_mse_index]\n",
- "))"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Clean up compute resource\n",
- "\n",
- "Use ```detach()``` to detach an existing DSVM from Workspace without deleting it. Use ```delete()``` if you created a new ```DsvmCompute``` and want to delete it."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# dsvm_compute.detach()\n",
- "# dsvm_compute.delete()"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "haining"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# 04. Train in a remote Linux VM\n",
+ "* Create Workspace\n",
+ "* Create `train.py` file\n",
+ "* Create and Attach a Remote VM (eg. DSVM) as compute resource.\n",
+ "* Upoad data files into default datastore\n",
+ "* Configure & execute a run in a few different ways\n",
+ " - Use system-built conda\n",
+ " - Use existing Python environment\n",
+ " - Use Docker \n",
+ "* Find the best model in the run"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Prerequisites\n",
+ "Make sure you go through the [configuration notebook](../../../configuration.ipynb) first if you haven't."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Check core SDK version number\n",
+ "import azureml.core\n",
+ "\n",
+ "print(\"SDK version:\", azureml.core.VERSION)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Initialize Workspace\n",
+ "\n",
+ "Initialize a workspace object from persisted configuration."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core import Workspace\n",
+ "\n",
+ "ws = Workspace.from_config()\n",
+ "print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep='\\n')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Create Experiment\n",
+ "\n",
+ "**Experiment** is a logical container in an Azure ML Workspace. It hosts run records which can include run metrics and output artifacts from your experiments."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "experiment_name = 'train-on-remote-vm'\n",
+ "\n",
+ "from azureml.core import Experiment\n",
+ "exp = Experiment(workspace=ws, name=experiment_name)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Let's also create a local folder to hold the training script."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import os\n",
+ "script_folder = './vm-run'\n",
+ "os.makedirs(script_folder, exist_ok=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Upload data files into datastore\n",
+ "Every workspace comes with a default datastore (and you can register more) which is backed by the Azure blob storage account associated with the workspace. We can use it to transfer data from local to the cloud, and access it from the compute target."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# get the default datastore\n",
+ "ds = ws.get_default_datastore()\n",
+ "print(ds.name, ds.datastore_type, ds.account_name, ds.container_name)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Load diabetes data from `scikit-learn` and save it as 2 local files."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from sklearn.datasets import load_diabetes\n",
+ "import numpy as np\n",
+ "\n",
+ "training_data = load_diabetes()\n",
+ "np.save(file='./features.npy', arr=training_data['data'])\n",
+ "np.save(file='./labels.npy', arr=training_data['target'])"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Now let's upload the 2 files into the default datastore under a path named `diabetes`:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "ds.upload_files(['./features.npy', './labels.npy'], target_path='diabetes', overwrite=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## View `train.py`\n",
+ "\n",
+ "For convenience, we created a training script for you. It is printed below as a text, but you can also run `%pfile ./train.py` in a cell to show the file. Please pay special attention on how we are loading the features and labels from files in the `data_folder` path, which is passed in as an argument of the training script (shown later)."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# copy train.py into the script folder\n",
+ "import shutil\n",
+ "shutil.copy('./train.py', os.path.join(script_folder, 'train.py'))\n",
+ "\n",
+ "with open(os.path.join(script_folder, './train.py'), 'r') as training_script:\n",
+ " print(training_script.read())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Create and Attach a DSVM as a compute target\n",
+ "\n",
+ "**Note**: To streamline the compute that Azure Machine Learning creates, we are making updates to support creating only single to multi-node `AmlCompute`. The DSVM can be created using the below single line command and then attached(like any VM) using the sample code below. Also note, that we only support Linux VMs for remote execution from AML and the commands below will spin a Linux VM only.\n",
+ "\n",
+ "```shell\n",
+ "# create a DSVM in your resource group\n",
+ "# note you need to be at least a contributor to the resource group in order to execute this command successfully\n",
+ "(myenv) $ az vm create --resource-group --name --image microsoft-dsvm:linux-data-science-vm-ubuntu:linuxdsvmubuntu:latest --admin-username --admin-password --generate-ssh-keys --authentication-type password\n",
+ "```\n",
+ "\n",
+ "**Note**: You can also use [this url](https://portal.azure.com/#create/microsoft-dsvm.linux-data-science-vm-ubuntulinuxdsvmubuntu) to create the VM using the Azure Portal\n",
+ "\n",
+ "**Note**: By default SSH runs on port 22 and you don't need to specify it. But if for security reasons you switch to a different port (such as 5022), you can specify the port number in the provisioning configuration object."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.compute import ComputeTarget, RemoteCompute\n",
+ "from azureml.core.compute_target import ComputeTargetException\n",
+ "\n",
+ "username = os.getenv('AZUREML_DSVM_USERNAME', default='')\n",
+ "address = os.getenv('AZUREML_DSVM_ADDRESS', default='')\n",
+ "\n",
+ "compute_target_name = 'cpudsvm'\n",
+ "# if you want to connect using SSH key instead of username/password you can provide parameters private_key_file and private_key_passphrase \n",
+ "try:\n",
+ " attached_dsvm_compute = RemoteCompute(workspace=ws, name=compute_target_name)\n",
+ " print('found existing:', attached_dsvm_compute.name)\n",
+ "except ComputeTargetException:\n",
+ " attach_config = RemoteCompute.attach_configuration(address=address,\n",
+ " ssh_port=22,\n",
+ " username=username,\n",
+ " private_key_file='./.ssh/id_rsa')\n",
+ " attached_dsvm_compute = ComputeTarget.attach(workspace=ws,\n",
+ " name=compute_target_name,\n",
+ " attach_config=attach_config)\n",
+ " attached_dsvm_compute.wait_for_completion(show_output=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Configure & Run\n",
+ "First let's create a `DataReferenceConfiguration` object to inform the system what data folder to download to the copmute target."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.runconfig import DataReferenceConfiguration\n",
+ "dr = DataReferenceConfiguration(datastore_name=ds.name, \n",
+ " path_on_datastore='diabetes', \n",
+ " mode='download', # download files from datastore to compute target\n",
+ " overwrite=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Now we can try a few different ways to run the training script in the VM."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Conda run\n",
+ "You can ask the system to build a conda environment based on your dependency specification, and submit your script to run there. Once the environment is built, and if you don't change your dependencies, it will be reused in subsequent runs."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.runconfig import RunConfiguration\n",
+ "from azureml.core.conda_dependencies import CondaDependencies\n",
+ "\n",
+ "# create a new RunConfig object\n",
+ "conda_run_config = RunConfiguration(framework=\"python\")\n",
+ "\n",
+ "# Set compute target to the Linux DSVM\n",
+ "conda_run_config.target = attached_dsvm_compute.name\n",
+ "\n",
+ "# set the data reference of the run configuration\n",
+ "conda_run_config.data_references = {ds.name: dr}\n",
+ "\n",
+ "# specify CondaDependencies obj\n",
+ "conda_run_config.environment.python.conda_dependencies = CondaDependencies.create(conda_packages=['scikit-learn'])"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core import ScriptRunConfig\n",
+ "\n",
+ "src = ScriptRunConfig(source_directory=script_folder, \n",
+ " script='train.py', \n",
+ " run_config=conda_run_config, \n",
+ " # pass the datastore reference as a parameter to the training script\n",
+ " arguments=['--data-folder', str(ds.as_download())] \n",
+ " ) \n",
+ "run = exp.submit(config=src)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run.wait_for_completion(show_output=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Show the run object. You can navigate to the Azure portal to see detailed information about the run."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Native VM run\n",
+ "You can also configure to use an exiting Python environment in the VM to execute the script without asking the system to create a conda environment for you."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# create a new RunConfig object\n",
+ "vm_run_config = RunConfiguration(framework=\"python\")\n",
+ "\n",
+ "# Set compute target to the Linux DSVM\n",
+ "vm_run_config.target = attached_dsvm_compute.name\n",
+ "\n",
+ "# set the data reference of the run coonfiguration\n",
+ "conda_run_config.data_references = {ds.name: dr}\n",
+ "\n",
+ "# Let system know that you will configure the Python environment yourself.\n",
+ "vm_run_config.environment.python.user_managed_dependencies = True"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "The below run will likely fail because `train.py` needs dependency `azureml`, `scikit-learn` and others, which are not found in that Python environment. "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "src = ScriptRunConfig(source_directory=script_folder, \n",
+ " script='train.py', \n",
+ " run_config=vm_run_config,\n",
+ " # pass the datastore reference as a parameter to the training script\n",
+ " arguments=['--data-folder', str(ds.as_download())])\n",
+ "run = exp.submit(config=src)\n",
+ "run.wait_for_completion(show_output=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "You can choose to SSH into the VM and install Azure ML SDK, and any other missing dependencies, in that Python environment. For demonstration purposes, we simply are going to use another script `train2.py` that doesn't have azureml dependencies, and submit it instead."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# copy train2.py into the script folder\n",
+ "shutil.copy('./train2.py', os.path.join(script_folder, 'train2.py'))\n",
+ "\n",
+ "with open(os.path.join(script_folder, './train2.py'), 'r') as training_script:\n",
+ " print(training_script.read())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Now let's try again. And this time it should work fine."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "src = ScriptRunConfig(source_directory=script_folder, \n",
+ " script='train2.py', \n",
+ " run_config=vm_run_config)\n",
+ "run = exp.submit(config=src)\n",
+ "run.wait_for_completion(show_output=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Note even in this case you get a run record with some basic statistics."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Configure a Docker run with new conda environment on the VM\n",
+ "You can execute in a Docker container in the VM. If you choose this option, the system will pull down a base Docker image, build a new conda environment in it if you ask for (you can also skip this if you are using a customer Docker image when a preconfigured Python environment), start a container, and run your script in there. This image is also uploaded into your ACR (Azure Container Registry) assoicated with your workspace, an reused if your dependencies don't change in the subsequent runs."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# Load the \"cpu-dsvm.runconfig\" file (created by the above attach operation) in memory\n",
+ "docker_run_config = RunConfiguration(framework=\"python\")\n",
+ "\n",
+ "# Set compute target to the Linux DSVM\n",
+ "docker_run_config.target = attached_dsvm_compute.name\n",
+ "\n",
+ "# Use Docker in the remote VM\n",
+ "docker_run_config.environment.docker.enabled = True\n",
+ "\n",
+ "# Use CPU base image from DockerHub\n",
+ "docker_run_config.environment.docker.base_image = azureml.core.runconfig.DEFAULT_CPU_IMAGE\n",
+ "print('Base Docker image is:', docker_run_config.environment.docker.base_image)\n",
+ "\n",
+ "# set the data reference of the run coonfiguration\n",
+ "docker_run_config.data_references = {ds.name: dr}\n",
+ "\n",
+ "# specify CondaDependencies obj\n",
+ "docker_run_config.environment.python.conda_dependencies = CondaDependencies.create(conda_packages=['scikit-learn'])"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Submit the Experiment\n",
+ "Submit script to run in the Docker image in the remote VM. If you run this for the first time, the system will download the base image, layer in packages specified in the `conda_dependencies.yml` file on top of the base image, create a container and then execute the script in the container."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "src = ScriptRunConfig(source_directory=script_folder, \n",
+ " script='train.py', \n",
+ " run_config=docker_run_config,\n",
+ " # pass the datastore reference as a parameter to the training script\n",
+ " arguments=['--data-folder', str(ds.as_download())])\n",
+ "run = exp.submit(config=src)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run.wait_for_completion(show_output=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### View run history details"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Find the best model"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Now we have tried various execution modes, we can find the best model from the last run."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# get all metris logged in the run\n",
+ "run.get_metrics()\n",
+ "metrics = run.get_metrics()"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# find the index where MSE is the smallest\n",
+ "indices = list(range(0, len(metrics['mse'])))\n",
+ "min_mse_index = min(indices, key=lambda x: metrics['mse'][x])\n",
+ "\n",
+ "print('When alpha is {1:0.2f}, we have min MSE {0:0.2f}.'.format(\n",
+ " metrics['mse'][min_mse_index], \n",
+ " metrics['alpha'][min_mse_index]\n",
+ "))"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Clean up compute resource\n",
+ "\n",
+ "Use ```detach()``` to detach an existing DSVM from Workspace without deleting it. Use ```delete()``` if you created a new ```DsvmCompute``` and want to delete it."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# dsvm_compute.detach()\n",
+ "# dsvm_compute.delete()"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "haining"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.6"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/how-to-use-azureml/training/train-on-remote-vm/train2.py b/how-to-use-azureml/training/train-on-remote-vm/train2.py
new file mode 100644
index 00000000..2cf812e0
--- /dev/null
+++ b/how-to-use-azureml/training/train-on-remote-vm/train2.py
@@ -0,0 +1,6 @@
+# Copyright (c) Microsoft. All rights reserved.
+# Licensed under the MIT license.
+
+print('####################################')
+print('Hello World (without Azure ML SDK)!')
+print('####################################')
diff --git a/how-to-use-azureml/training/train-within-notebook/train-within-notebook.ipynb b/how-to-use-azureml/training/train-within-notebook/train-within-notebook.ipynb
index 9aa6eff1..33a1ab74 100644
--- a/how-to-use-azureml/training/train-within-notebook/train-within-notebook.ipynb
+++ b/how-to-use-azureml/training/train-within-notebook/train-within-notebook.ipynb
@@ -1,708 +1,706 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Train and deploy a model\n",
- "_**Create and deploy a model directly from a notebook**_\n",
- "\n",
- "---\n",
- "---\n",
- "\n",
- "## Contents\n",
- "1. [Introduction](#Introduction)\n",
- "1. [Setup](#Setup)\n",
- "1. [Data](#Data)\n",
- "1. [Train](#Train)\n",
- " 1. Viewing run results\n",
- " 1. Simple parameter sweep\n",
- " 1. Viewing experiment results\n",
- " 1. Select the best model\n",
- "1. [Deploy](#Deploy)\n",
- " 1. Register the model\n",
- " 1. Create a scoring file\n",
- " 1. Describe your environment\n",
- " 1. Descrice your target compute\n",
- " 1. Deploy your webservice\n",
- " 1. Test your webservice\n",
- " 1. Clean up\n",
- "1. [Next Steps](#Next%20Steps)\n",
- "\n",
- "---\n",
- "\n",
- "## Introduction\n",
- "Azure Machine Learning provides capabilities to control all aspects of model training and deployment directly from a notebook using the AML Python SDK. In this notebook we will\n",
- "* connect to our AML Workspace\n",
- "* create an experiment that contains multiple runs with tracked metrics\n",
- "* choose the best model created across all runs\n",
- "* deploy that model as a service\n",
- "\n",
- "In the end we will have a model deployed as a web service which we can call from an HTTP endpoint"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "---\n",
- "\n",
- "## Setup\n",
- "Make sure you have completed the [Configuration](..\\..\\configuration.ipnyb) notebook to set up your Azure Machine Learning workspace and ensure other common prerequisites are met. From the configuration, the important sections are the workspace configuration and ACI regristration.\n",
- "\n",
- "We will also need the following libraries install to our conda environment. If these are not installed, use the following command to do so and restart the notebook.\n",
- "```shell\n",
- "(myenv) $ conda install -y matplotlib tqdm scikit-learn\n",
- "```\n",
- "\n",
- "For this notebook we need the Azure ML SDK and access to our workspace. The following cell imports the SDK, checks the version, and accesses our already configured AzureML workspace."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "install"
- ]
- },
- "outputs": [],
- "source": [
- "import azureml.core\n",
- "from azureml.core import Experiment, Run, Workspace\n",
- "\n",
- "# Check core SDK version number\n",
- "print(\"This notebook was created using version 1.0.2 of the Azure ML SDK\")\n",
- "print(\"You are currently using version\", azureml.core.VERSION, \"of the Azure ML SDK\")\n",
- "print(\"\")\n",
- "\n",
- "\n",
- "ws = Workspace.from_config()\n",
- "print('Workspace name: ' + ws.name, \n",
- " 'Azure region: ' + ws.location, \n",
- " 'Subscription id: ' + ws.subscription_id, \n",
- " 'Resource group: ' + ws.resource_group, sep='\\n')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "---\n",
- "\n",
- "## Data\n",
- "We will use the diabetes dataset for this experiement, a well-known small dataset that comes with scikit-learn. This cell loads the dataset and splits it into random training and testing sets.\n"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from sklearn.datasets import load_diabetes\n",
- "from sklearn.linear_model import Ridge\n",
- "from sklearn.metrics import mean_squared_error\n",
- "from sklearn.model_selection import train_test_split\n",
- "from sklearn.externals import joblib\n",
- "\n",
- "X, y = load_diabetes(return_X_y = True)\n",
- "columns = ['age', 'gender', 'bmi', 'bp', 's1', 's2', 's3', 's4', 's5', 's6']\n",
- "X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=0)\n",
- "data = {\n",
- " \"train\":{\"X\": X_train, \"y\": y_train}, \n",
- " \"test\":{\"X\": X_test, \"y\": y_test}\n",
- "}\n",
- "\n",
- "print (\"Data contains\", len(data['train']['X']), \"training samples and\",len(data['test']['X']), \"test samples\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "---\n",
- "## Train\n",
- "\n",
- "Let's use scikit-learn to train a simple Ridge regression model. We use AML to record interesting information about the model in an Experiment. An Experiment contains a series of trials called Runs. During this trial we use AML in the following way:\n",
- "* We access an experiment from our AML workspace by name, which will be created if it doesn't exist\n",
- "* We use `start_logging` to create a new run in this experiment\n",
- "* We use `run.log()` to record a parameter, alpha, and an accuracy measure - the Mean Squared Error (MSE) to the run. We will be able to review and compare these measures in the Azure Portal at a later time.\n",
- "* We store the resulting model in the **outputs** directory, which is automatically captured by AML when the run is complete.\n",
- "* We use `run.take_snapshot()` to capture *this* notebook so we can reproduce this experiment at a later time.\n",
- "* We use `run.complete()` to indicate that the run is over and results can be captured and finalized"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "local run",
- "outputs upload"
- ]
- },
- "outputs": [],
- "source": [
- "# Get an experiment object from Azure Machine Learning\n",
- "experiment = Experiment(workspace=ws, name=\"train-within-notebook\")\n",
- "\n",
- "# Create a run object in the experiment\n",
- "run = experiment.start_logging()# Log the algorithm parameter alpha to the run\n",
- "run.log('alpha', 0.03)\n",
- "\n",
- "# Create, fit, and test the scikit-learn Ridge regression model\n",
- "regression_model = Ridge(alpha=0.03)\n",
- "regression_model.fit(data['train']['X'], data['train']['y'])\n",
- "preds = regression_model.predict(data['test']['X'])\n",
- "\n",
- "# Output the Mean Squared Error to the notebook and to the run\n",
- "print('Mean Squared Error is', mean_squared_error(data['test']['y'], preds))\n",
- "run.log('mse', mean_squared_error(data['test']['y'], preds))\n",
- "\n",
- "# Save the model to the outputs directory for capture\n",
- "joblib.dump(value=regression_model, filename='outputs/model.pkl')\n",
- "\n",
- "# Take a snapshot of the directory containing this notebook\n",
- "run.take_snapshot('./')\n",
- "\n",
- "# Complete the run\n",
- "run.complete()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Viewing run results\n",
- "Azure Machine Learning stores all the details about the run in the Azure cloud. Let's access those details by retrieving a link to the run using the default run output. Clicking on the resulting link will take you to an interactive page presenting all run information."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "run"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Simple parameter sweep\n",
- "Now let's take the same concept from above and modify the **alpha** parameter. For each value of alpha we will create a run that will store metrics and the resulting model. In the end we can use the captured run history to determine which model was the best for us to deploy. \n",
- "\n",
- "Note that by using `with experiment.start_logging() as run` AML will automatically call `run.complete()` at the end of each loop.\n",
- "\n",
- "This example also uses the **tqdm** library to provide a thermometer feedback"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import numpy as np\n",
- "import os\n",
- "from tqdm import tqdm\n",
- "\n",
- "model_name = \"model.pkl\"\n",
- "\n",
- "# list of numbers from 0 to 1.0 with a 0.05 interval\n",
- "alphas = np.arange(0.0, 1.0, 0.05)\n",
- "\n",
- "# try a bunch of alpha values in a Linear Regression (Ridge) model\n",
- "for alpha in tqdm(alphas):\n",
- " # create a bunch of runs, each train a model with a different alpha value\n",
- " with experiment.start_logging() as run:\n",
- " # Use Ridge algorithm to build a regression model\n",
- " regression_model = Ridge(alpha=alpha)\n",
- " regression_model.fit(X=data[\"train\"][\"X\"], y=data[\"train\"][\"y\"])\n",
- " preds = regression_model.predict(X=data[\"test\"][\"X\"])\n",
- " mse = mean_squared_error(y_true=data[\"test\"][\"y\"], y_pred=preds)\n",
- "\n",
- " # log alpha, mean_squared_error and feature names in run history\n",
- " run.log(name=\"alpha\", value=alpha)\n",
- " run.log(name=\"mse\", value=mse)\n",
- "\n",
- " # Save the model to the outputs directory for capture\n",
- " joblib.dump(value=regression_model, filename='outputs/model.pkl')\n",
- " \n",
- " # Capture this notebook with the run\n",
- " run.take_snapshot('./')\n"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Viewing experiment results\n",
- "Similar to viewing the run, we can also view the entire experiment. The experiment report view in the Azure portal lets us view all the runs in a table, and also allows us to customize charts. This way, we can see how the alpha parameter impacts the quality of the model"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# now let's take a look at the experiment in Azure portal.\n",
- "experiment"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Select the best model \n",
- "Now that we've created many runs with different parameters, we need to determine which model is the best for deployment. For this, we will iterate over the set of runs. From each run we will take the *run id* using the `id` property, and examine the metrics by calling `run.get_metrics()`. \n",
- "\n",
- "Since each run may be different, we do need to check if the run has the metric that we are looking for, in this case, **mse**. To find the best run, we create a dictionary mapping the run id's to the metrics.\n",
- "\n",
- "Finally, we use the `tag` method to mark the best run to make it easier to find later. "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "runs = {}\n",
- "run_metrics = {}\n",
- "\n",
- "# Create dictionaries containing the runs and the metrics for all runs containing the 'mse' metric\n",
- "for r in tqdm(experiment.get_runs()):\n",
- " metrics = r.get_metrics()\n",
- " if 'mse' in metrics.keys():\n",
- " runs[r.id] = r\n",
- " run_metrics[r.id] = metrics\n",
- "\n",
- "# Find the run with the best (lowest) mean squared error and display the id and metrics\n",
- "best_run_id = min(run_metrics, key = lambda k: run_metrics[k]['mse'])\n",
- "best_run = runs[best_run_id]\n",
- "print('Best run is:', best_run_id)\n",
- "print('Metrics:', run_metrics[best_run_id])\n",
- "\n",
- "# Tag the best run for identification later\n",
- "best_run.tag(\"Best Run\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "---\n",
- "## Deploy\n",
- "Now that we have trained a set of models and identified the run containing the best model, we want to deploy the model for real time inferencing. The process of deploying a model involves\n",
- "* registering a model in your workspace\n",
- "* creating a scoring file containing init and run methods\n",
- "* creating an environment dependency file describing packages necessary for your scoring file\n",
- "* creating a docker image containing a properly described environment, your model, and your scoring file\n",
- "* deploying that docker image as a web service"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Register a model\n",
- "We have already identified which run contains the \"best model\" by our evaluation criteria. Each run has a file structure associated with it that contains various files collected during the run. Since a run can have many outputs we need to tell AML which file from those outputs represents the model that we want to use for our deployment. We can use the `run.get_file_names()` method to list the files associated with the run, and then use the `run.register_model()` method to place the model in the workspace's model registry.\n",
- "\n",
- "When using `run.register_model()` we supply a `model_name` that is meaningful for our scenario and the `model_path` of the model relative to the run. In this case, the model path is what is returned from `run.get_file_names()`"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "query history"
- ]
- },
- "outputs": [],
- "source": [
- "# View the files in the run\n",
- "for f in best_run.get_file_names():\n",
- " print(f)\n",
- " \n",
- "# Register the model with the workspace\n",
- "model = best_run.register_model(model_name='best_model', model_path='outputs/model.pkl')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Once a model is registered, it is accessible from the list of models on the AML workspace. If you register models with the same name multiple times, AML keeps a version history of those models for you. The `Model.list()` lists all models in a workspace, and can be filtered by name, tags, or model properties. "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "register model from history"
- ]
- },
- "outputs": [],
- "source": [
- "# Find all models called \"best_model\" and display their version numbers\n",
- "from azureml.core.model import Model\n",
- "models = Model.list(ws, name='best_model')\n",
- "for m in models:\n",
- " print(m.name, m.version)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create a scoring file\n",
- "\n",
- "Since your model file can essentially be anything you want it to be, you need to supply a scoring script that can load your model and then apply the model to new data. This script is your 'scoring file'. This scoring file is a python program containing, at a minimum, two methods `init()` and `run()`. The `init()` method is called once when your deployment is started so you can load your model and any other required objects. This method uses the `get_model_path` function to locate the registered model inside the docker container. The `run()` method is called interactively when the web service is called with one or more data samples to predict.\n",
- "\n",
- "The scoring file used for this exercise is [here](score.py). \n"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Describe your environment\n",
- "\n",
- "Each modelling process may require a unique set of packages. Therefore we need to create a dependency file providing instructions to AML on how to contstruct a docker image that can support the models and any other objects required for inferencing. In the following cell, we create a environment dependency file, *myenv.yml* that specifies which libraries are needed by the scoring script. You can create this file manually, or use the `CondaDependencies` class to create it for you.\n",
- "\n",
- "Next we use this environment file to describe the docker container that we need to create in order to deploy our model. This container is created using our environment description and includes our scoring script."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from azureml.core.conda_dependencies import CondaDependencies \n",
- "from azureml.core.image import ContainerImage\n",
- "\n",
- "# Create an empty conda environment and add the scikit-learn package\n",
- "env = CondaDependencies()\n",
- "env.add_conda_package(\"scikit-learn\")\n",
- "\n",
- "# Display the environment\n",
- "print(env.serialize_to_string())\n",
- "\n",
- "# Write the environment to disk\n",
- "with open(\"myenv.yml\",\"w\") as f:\n",
- " f.write(env.serialize_to_string())\n",
- "\n",
- "# Create a configuration object indicating how our deployment container needs to be created\n",
- "image_config = ContainerImage.image_configuration(execution_script=\"score.py\", \n",
- " runtime=\"python\", \n",
- " conda_file=\"myenv.yml\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Describe your target compute\n",
- "In addition to the container, we also need to describe the type of compute we want to allocate for our webservice. In in this example we are using an [Azure Container Instance](https://azure.microsoft.com/en-us/services/container-instances/) which is a good choice for quick and cost-effective dev/test deployment scenarios. ACI instances require the number of cores you want to run and memory you need. Tags and descriptions are available for you to identify the instances in AML when viewing the Compute tab in the AML Portal.\n",
- "\n",
- "For production workloads, it is better to use [Azure Kubernentes Service (AKS)](https://azure.microsoft.com/en-us/services/kubernetes-service/) instead. Try [this notebook](11.production-deploy-to-aks.ipynb) to see how that can be done from Azure ML.\n"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "deploy service",
- "aci"
- ]
- },
- "outputs": [],
- "source": [
- "from azureml.core.webservice import AciWebservice\n",
- "\n",
- "aciconfig = AciWebservice.deploy_configuration(cpu_cores=1, \n",
- " memory_gb=1, \n",
- " tags={'sample name': 'AML 101'}, \n",
- " description='This is a great example.')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Deploy your webservice\n",
- "The final step to deploying your webservice is to call `WebService.deploy_from_model()`. This function uses the deployment and image configurations created above to perform the following:\n",
- "* Build a docker image\n",
- "* Deploy to the docker image to an Azure Container Instance\n",
- "* Copy your model files to the Azure Container Instance\n",
- "* Call the `init()` function in your scoring file\n",
- "* Provide an HTTP endpoint for scoring calls\n",
- "\n",
- "The `deploy_from_model` method requires the following parameters\n",
- "* `workspace` - the workspace containing the service\n",
- "* `name` - a unique named used to identify the service in the workspace\n",
- "* `models` - an array of models to be deployed into the container\n",
- "* `image_config` - a configuration object describing the image environment\n",
- "* `deployment_config` - a configuration object describing the compute type\n",
- " \n",
- "**Note:** The web service creation can take several minutes. "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "deploy service",
- "aci"
- ]
- },
- "outputs": [],
- "source": [
- "%%time\n",
- "from azureml.core.webservice import Webservice\n",
- "\n",
- "# Create the webservice using all of the precreated configurations and our best model\n",
- "service = Webservice.deploy_from_model(name='my-aci-svc',\n",
- " deployment_config=aciconfig,\n",
- " models=[model],\n",
- " image_config=image_config,\n",
- " workspace=ws)\n",
- "\n",
- "# Wait for the service deployment to complete while displaying log output\n",
- "service.wait_for_deployment(show_output=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "\n",
- "### Test your webservice"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Now that your web service is runing you can send JSON data directly to the service using the `run` method. This cell pulls the first test sample from the original dataset into JSON and then sends it to the service."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "deploy service",
- "aci"
- ]
- },
- "outputs": [],
- "source": [
- "import json\n",
- "# scrape the first row from the test set.\n",
- "test_samples = json.dumps({\"data\": X_test[0:1, :].tolist()})\n",
- "\n",
- "#score on our service\n",
- "service.run(input_data = test_samples)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "This cell shows how you can send multiple rows to the webservice at once. It then calculates the residuals - that is, the errors - by subtracting out the actual values from the results. These residuals are used later to show a plotted result."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "deploy service",
- "aci"
- ]
- },
- "outputs": [],
- "source": [
- "# score the entire test set.\n",
- "test_samples = json.dumps({'data': X_test.tolist()})\n",
- "\n",
- "result = service.run(input_data = test_samples)\n",
- "residual = result - y_test"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "This cell shows how you can use the `service.scoring_uri` property to access the HTTP endpoint of the service and call it using standard POST operations."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "deploy service",
- "aci"
- ]
- },
- "outputs": [],
- "source": [
- "import requests\n",
- "import json\n",
- "\n",
- "# use the first row from the test set again\n",
- "test_samples = json.dumps({\"data\": X_test[0:1, :].tolist()})\n",
- "\n",
- "# create the required header\n",
- "headers = {'Content-Type':'application/json'}\n",
- "\n",
- "# post the request to the service and display the result\n",
- "resp = requests.post(service.scoring_uri, test_samples, headers = headers)\n",
- "print(resp.text)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Residual graph\n",
- "One way to understand the behavior of your model is to see how the data performs against data with known results. This cell uses matplotlib to create a histogram of the residual values, or errors, created from scoring the test samples.\n",
- "\n",
- "A good model should have residual values that cluster around 0 - that is, no error. Observing the resulting histogram can also show you if the model is skewed in any particular direction."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%matplotlib inline\n",
- "import matplotlib\n",
- "import matplotlib.pyplot as plt\n",
- "\n",
- "f, (a0, a1) = plt.subplots(1, 2, gridspec_kw={'width_ratios':[3, 1], 'wspace':0, 'hspace': 0})\n",
- "f.suptitle('Residual Values', fontsize = 18)\n",
- "\n",
- "f.set_figheight(6)\n",
- "f.set_figwidth(14)\n",
- "\n",
- "a0.plot(residual, 'bo', alpha=0.4);\n",
- "a0.plot([0,90], [0,0], 'r', lw=2)\n",
- "a0.set_ylabel('residue values', fontsize=14)\n",
- "a0.set_xlabel('test data set', fontsize=14)\n",
- "\n",
- "a1.hist(residual, orientation='horizontal', color='blue', bins=10, histtype='step');\n",
- "a1.hist(residual, orientation='horizontal', color='blue', alpha=0.2, bins=10);\n",
- "a1.set_yticklabels([])\n",
- "\n",
- "plt.show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Clean up"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Delete the ACI instance to stop the compute and any associated billing."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "deploy service",
- "aci"
- ]
- },
- "outputs": [],
- "source": [
- "%%time\n",
- "service.delete()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "---\n",
- "## Next Steps"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "In this example, you created a series of models inside the notebook using local data, stored them inside an AML experiment, found the best one and deployed it as a live service! From here you can continue to use Azure Machine Learning in this regard to run your own experiments and deploy your own models, or you can expand into further capabilities of AML!\n",
- "\n",
- "If you have a model that is difficult to process locally, either because the data is remote or the model is large, try the [train-on-remote-vm](../train-on-remote-vm) notebook to learn about submitting remote jobs.\n",
- "\n",
- "If you want to take advantage of multiple cloud machines to perform large parameter sweeps try the [train-hyperparameter-tune-deploy-with-pytorch](../../training-with-deep-learning/train-hyperparameter-tune-deploy-with-pytorch\n",
- ") sample.\n",
- "\n",
- "If you want to deploy models to a production cluster try the [production-deploy-to-aks](../../deployment/production-deploy-to-aks\n",
- ") notebook."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": []
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "roastala"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Train and deploy a model\n",
+ "_**Create and deploy a model directly from a notebook**_\n",
+ "\n",
+ "---\n",
+ "---\n",
+ "\n",
+ "## Contents\n",
+ "1. [Introduction](#Introduction)\n",
+ "1. [Setup](#Setup)\n",
+ "1. [Data](#Data)\n",
+ "1. [Train](#Train)\n",
+ " 1. Viewing run results\n",
+ " 1. Simple parameter sweep\n",
+ " 1. Viewing experiment results\n",
+ " 1. Select the best model\n",
+ "1. [Deploy](#Deploy)\n",
+ " 1. Register the model\n",
+ " 1. Create a scoring file\n",
+ " 1. Describe your environment\n",
+ " 1. Descrice your target compute\n",
+ " 1. Deploy your webservice\n",
+ " 1. Test your webservice\n",
+ " 1. Clean up\n",
+ "1. [Next Steps](#Next%20Steps)\n",
+ "\n",
+ "---\n",
+ "\n",
+ "## Introduction\n",
+ "Azure Machine Learning provides capabilities to control all aspects of model training and deployment directly from a notebook using the AML Python SDK. In this notebook we will\n",
+ "* connect to our AML Workspace\n",
+ "* create an experiment that contains multiple runs with tracked metrics\n",
+ "* choose the best model created across all runs\n",
+ "* deploy that model as a service\n",
+ "\n",
+ "In the end we will have a model deployed as a web service which we can call from an HTTP endpoint"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "---\n",
+ "\n",
+ "## Setup\n",
+ "Make sure you have completed the [Configuration](../../../configuration.ipnyb) notebook to set up your Azure Machine Learning workspace and ensure other common prerequisites are met. From the configuration, the important sections are the workspace configuration and ACI regristration.\n",
+ "\n",
+ "We will also need the following libraries install to our conda environment. If these are not installed, use the following command to do so and restart the notebook.\n",
+ "```shell\n",
+ "(myenv) $ conda install -y matplotlib tqdm scikit-learn\n",
+ "```\n",
+ "\n",
+ "For this notebook we need the Azure ML SDK and access to our workspace. The following cell imports the SDK, checks the version, and accesses our already configured AzureML workspace."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "install"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "import azureml.core\n",
+ "from azureml.core import Experiment, Workspace\n",
+ "\n",
+ "# Check core SDK version number\n",
+ "print(\"This notebook was created using version 1.0.2 of the Azure ML SDK\")\n",
+ "print(\"You are currently using version\", azureml.core.VERSION, \"of the Azure ML SDK\")\n",
+ "print(\"\")\n",
+ "\n",
+ "\n",
+ "ws = Workspace.from_config()\n",
+ "print('Workspace name: ' + ws.name, \n",
+ " 'Azure region: ' + ws.location, \n",
+ " 'Subscription id: ' + ws.subscription_id, \n",
+ " 'Resource group: ' + ws.resource_group, sep='\\n')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "---\n",
+ "\n",
+ "## Data\n",
+ "We will use the diabetes dataset for this experiement, a well-known small dataset that comes with scikit-learn. This cell loads the dataset and splits it into random training and testing sets.\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from sklearn.datasets import load_diabetes\n",
+ "from sklearn.linear_model import Ridge\n",
+ "from sklearn.metrics import mean_squared_error\n",
+ "from sklearn.model_selection import train_test_split\n",
+ "from sklearn.externals import joblib\n",
+ "\n",
+ "X, y = load_diabetes(return_X_y = True)\n",
+ "columns = ['age', 'gender', 'bmi', 'bp', 's1', 's2', 's3', 's4', 's5', 's6']\n",
+ "X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=0)\n",
+ "data = {\n",
+ " \"train\":{\"X\": X_train, \"y\": y_train}, \n",
+ " \"test\":{\"X\": X_test, \"y\": y_test}\n",
+ "}\n",
+ "\n",
+ "print (\"Data contains\", len(data['train']['X']), \"training samples and\",len(data['test']['X']), \"test samples\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "---\n",
+ "## Train\n",
+ "\n",
+ "Let's use scikit-learn to train a simple Ridge regression model. We use AML to record interesting information about the model in an Experiment. An Experiment contains a series of trials called Runs. During this trial we use AML in the following way:\n",
+ "* We access an experiment from our AML workspace by name, which will be created if it doesn't exist\n",
+ "* We use `start_logging` to create a new run in this experiment\n",
+ "* We use `run.log()` to record a parameter, alpha, and an accuracy measure - the Mean Squared Error (MSE) to the run. We will be able to review and compare these measures in the Azure Portal at a later time.\n",
+ "* We store the resulting model in the **outputs** directory, which is automatically captured by AML when the run is complete.\n",
+ "* We use `run.take_snapshot()` to capture *this* notebook so we can reproduce this experiment at a later time.\n",
+ "* We use `run.complete()` to indicate that the run is over and results can be captured and finalized"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "local run",
+ "outputs upload"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "# Get an experiment object from Azure Machine Learning\n",
+ "experiment = Experiment(workspace=ws, name=\"train-within-notebook\")\n",
+ "\n",
+ "# Create a run object in the experiment\n",
+ "run = experiment.start_logging()# Log the algorithm parameter alpha to the run\n",
+ "run.log('alpha', 0.03)\n",
+ "\n",
+ "# Create, fit, and test the scikit-learn Ridge regression model\n",
+ "regression_model = Ridge(alpha=0.03)\n",
+ "regression_model.fit(data['train']['X'], data['train']['y'])\n",
+ "preds = regression_model.predict(data['test']['X'])\n",
+ "\n",
+ "# Output the Mean Squared Error to the notebook and to the run\n",
+ "print('Mean Squared Error is', mean_squared_error(data['test']['y'], preds))\n",
+ "run.log('mse', mean_squared_error(data['test']['y'], preds))\n",
+ "\n",
+ "# Save the model to the outputs directory for capture\n",
+ "joblib.dump(value=regression_model, filename='outputs/model.pkl')\n",
+ "\n",
+ "# Take a snapshot of the directory containing this notebook\n",
+ "run.take_snapshot('./')\n",
+ "\n",
+ "# Complete the run\n",
+ "run.complete()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Viewing run results\n",
+ "Azure Machine Learning stores all the details about the run in the Azure cloud. Let's access those details by retrieving a link to the run using the default run output. Clicking on the resulting link will take you to an interactive page presenting all run information."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "run"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Simple parameter sweep\n",
+ "Now let's take the same concept from above and modify the **alpha** parameter. For each value of alpha we will create a run that will store metrics and the resulting model. In the end we can use the captured run history to determine which model was the best for us to deploy. \n",
+ "\n",
+ "Note that by using `with experiment.start_logging() as run` AML will automatically call `run.complete()` at the end of each loop.\n",
+ "\n",
+ "This example also uses the **tqdm** library to provide a thermometer feedback"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import numpy as np\n",
+ "import os\n",
+ "from tqdm import tqdm\n",
+ "\n",
+ "model_name = \"model.pkl\"\n",
+ "\n",
+ "# list of numbers from 0 to 1.0 with a 0.05 interval\n",
+ "alphas = np.arange(0.0, 1.0, 0.05)\n",
+ "\n",
+ "# try a bunch of alpha values in a Linear Regression (Ridge) model\n",
+ "for alpha in tqdm(alphas):\n",
+ " # create a bunch of runs, each train a model with a different alpha value\n",
+ " with experiment.start_logging() as run:\n",
+ " # Use Ridge algorithm to build a regression model\n",
+ " regression_model = Ridge(alpha=alpha)\n",
+ " regression_model.fit(X=data[\"train\"][\"X\"], y=data[\"train\"][\"y\"])\n",
+ " preds = regression_model.predict(X=data[\"test\"][\"X\"])\n",
+ " mse = mean_squared_error(y_true=data[\"test\"][\"y\"], y_pred=preds)\n",
+ "\n",
+ " # log alpha, mean_squared_error and feature names in run history\n",
+ " run.log(name=\"alpha\", value=alpha)\n",
+ " run.log(name=\"mse\", value=mse)\n",
+ "\n",
+ " # Save the model to the outputs directory for capture\n",
+ " joblib.dump(value=regression_model, filename='outputs/model.pkl')\n",
+ " \n",
+ " # Capture this notebook with the run\n",
+ " run.take_snapshot('./')\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Viewing experiment results\n",
+ "Similar to viewing the run, we can also view the entire experiment. The experiment report view in the Azure portal lets us view all the runs in a table, and also allows us to customize charts. This way, we can see how the alpha parameter impacts the quality of the model"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# now let's take a look at the experiment in Azure portal.\n",
+ "experiment"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Select the best model \n",
+ "Now that we've created many runs with different parameters, we need to determine which model is the best for deployment. For this, we will iterate over the set of runs. From each run we will take the *run id* using the `id` property, and examine the metrics by calling `run.get_metrics()`. \n",
+ "\n",
+ "Since each run may be different, we do need to check if the run has the metric that we are looking for, in this case, **mse**. To find the best run, we create a dictionary mapping the run id's to the metrics.\n",
+ "\n",
+ "Finally, we use the `tag` method to mark the best run to make it easier to find later. "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "runs = {}\n",
+ "run_metrics = {}\n",
+ "\n",
+ "# Create dictionaries containing the runs and the metrics for all runs containing the 'mse' metric\n",
+ "for r in tqdm(experiment.get_runs()):\n",
+ " metrics = r.get_metrics()\n",
+ " if 'mse' in metrics.keys():\n",
+ " runs[r.id] = r\n",
+ " run_metrics[r.id] = metrics\n",
+ "\n",
+ "# Find the run with the best (lowest) mean squared error and display the id and metrics\n",
+ "best_run_id = min(run_metrics, key = lambda k: run_metrics[k]['mse'])\n",
+ "best_run = runs[best_run_id]\n",
+ "print('Best run is:', best_run_id)\n",
+ "print('Metrics:', run_metrics[best_run_id])\n",
+ "\n",
+ "# Tag the best run for identification later\n",
+ "best_run.tag(\"Best Run\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "---\n",
+ "## Deploy\n",
+ "Now that we have trained a set of models and identified the run containing the best model, we want to deploy the model for real time inferencing. The process of deploying a model involves\n",
+ "* registering a model in your workspace\n",
+ "* creating a scoring file containing init and run methods\n",
+ "* creating an environment dependency file describing packages necessary for your scoring file\n",
+ "* creating a docker image containing a properly described environment, your model, and your scoring file\n",
+ "* deploying that docker image as a web service"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Register a model\n",
+ "We have already identified which run contains the \"best model\" by our evaluation criteria. Each run has a file structure associated with it that contains various files collected during the run. Since a run can have many outputs we need to tell AML which file from those outputs represents the model that we want to use for our deployment. We can use the `run.get_file_names()` method to list the files associated with the run, and then use the `run.register_model()` method to place the model in the workspace's model registry.\n",
+ "\n",
+ "When using `run.register_model()` we supply a `model_name` that is meaningful for our scenario and the `model_path` of the model relative to the run. In this case, the model path is what is returned from `run.get_file_names()`"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "query history"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "# View the files in the run\n",
+ "for f in best_run.get_file_names():\n",
+ " print(f)\n",
+ " \n",
+ "# Register the model with the workspace\n",
+ "model = best_run.register_model(model_name='best_model', model_path='outputs/model.pkl')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Once a model is registered, it is accessible from the list of models on the AML workspace. If you register models with the same name multiple times, AML keeps a version history of those models for you. The `Model.list()` lists all models in a workspace, and can be filtered by name, tags, or model properties. "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "register model from history"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "# Find all models called \"best_model\" and display their version numbers\n",
+ "from azureml.core.model import Model\n",
+ "models = Model.list(ws, name='best_model')\n",
+ "for m in models:\n",
+ " print(m.name, m.version)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create a scoring file\n",
+ "\n",
+ "Since your model file can essentially be anything you want it to be, you need to supply a scoring script that can load your model and then apply the model to new data. This script is your 'scoring file'. This scoring file is a python program containing, at a minimum, two methods `init()` and `run()`. The `init()` method is called once when your deployment is started so you can load your model and any other required objects. This method uses the `get_model_path` function to locate the registered model inside the docker container. The `run()` method is called interactively when the web service is called with one or more data samples to predict.\n",
+ "\n",
+ "The scoring file used for this exercise is [here](score.py). \n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Describe your environment\n",
+ "\n",
+ "Each modelling process may require a unique set of packages. Therefore we need to create a dependency file providing instructions to AML on how to contstruct a docker image that can support the models and any other objects required for inferencing. In the following cell, we create a environment dependency file, *myenv.yml* that specifies which libraries are needed by the scoring script. You can create this file manually, or use the `CondaDependencies` class to create it for you.\n",
+ "\n",
+ "Next we use this environment file to describe the docker container that we need to create in order to deploy our model. This container is created using our environment description and includes our scoring script."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from azureml.core.conda_dependencies import CondaDependencies \n",
+ "from azureml.core.image import ContainerImage\n",
+ "\n",
+ "# Create an empty conda environment and add the scikit-learn package\n",
+ "env = CondaDependencies()\n",
+ "env.add_conda_package(\"scikit-learn\")\n",
+ "\n",
+ "# Display the environment\n",
+ "print(env.serialize_to_string())\n",
+ "\n",
+ "# Write the environment to disk\n",
+ "with open(\"myenv.yml\",\"w\") as f:\n",
+ " f.write(env.serialize_to_string())\n",
+ "\n",
+ "# Create a configuration object indicating how our deployment container needs to be created\n",
+ "image_config = ContainerImage.image_configuration(execution_script=\"score.py\", \n",
+ " runtime=\"python\", \n",
+ " conda_file=\"myenv.yml\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Describe your target compute\n",
+ "In addition to the container, we also need to describe the type of compute we want to allocate for our webservice. In in this example we are using an [Azure Container Instance](https://azure.microsoft.com/en-us/services/container-instances/) which is a good choice for quick and cost-effective dev/test deployment scenarios. ACI instances require the number of cores you want to run and memory you need. Tags and descriptions are available for you to identify the instances in AML when viewing the Compute tab in the AML Portal.\n",
+ "\n",
+ "For production workloads, it is better to use [Azure Kubernentes Service (AKS)](https://azure.microsoft.com/en-us/services/kubernetes-service/) instead. Try [this notebook](11.production-deploy-to-aks.ipynb) to see how that can be done from Azure ML.\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "deploy service",
+ "aci"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "from azureml.core.webservice import AciWebservice\n",
+ "\n",
+ "aciconfig = AciWebservice.deploy_configuration(cpu_cores=1, \n",
+ " memory_gb=1, \n",
+ " tags={'sample name': 'AML 101'}, \n",
+ " description='This is a great example.')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Deploy your webservice\n",
+ "The final step to deploying your webservice is to call `WebService.deploy_from_model()`. This function uses the deployment and image configurations created above to perform the following:\n",
+ "* Build a docker image\n",
+ "* Deploy to the docker image to an Azure Container Instance\n",
+ "* Copy your model files to the Azure Container Instance\n",
+ "* Call the `init()` function in your scoring file\n",
+ "* Provide an HTTP endpoint for scoring calls\n",
+ "\n",
+ "The `deploy_from_model` method requires the following parameters\n",
+ "* `workspace` - the workspace containing the service\n",
+ "* `name` - a unique named used to identify the service in the workspace\n",
+ "* `models` - an array of models to be deployed into the container\n",
+ "* `image_config` - a configuration object describing the image environment\n",
+ "* `deployment_config` - a configuration object describing the compute type\n",
+ " \n",
+ "**Note:** The web service creation can take several minutes. "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "deploy service",
+ "aci"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "%%time\n",
+ "from azureml.core.webservice import Webservice\n",
+ "\n",
+ "# Create the webservice using all of the precreated configurations and our best model\n",
+ "service = Webservice.deploy_from_model(name='my-aci-svc',\n",
+ " deployment_config=aciconfig,\n",
+ " models=[model],\n",
+ " image_config=image_config,\n",
+ " workspace=ws)\n",
+ "\n",
+ "# Wait for the service deployment to complete while displaying log output\n",
+ "service.wait_for_deployment(show_output=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "\n",
+ "### Test your webservice"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Now that your web service is runing you can send JSON data directly to the service using the `run` method. This cell pulls the first test sample from the original dataset into JSON and then sends it to the service."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "deploy service",
+ "aci"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "import json\n",
+ "# scrape the first row from the test set.\n",
+ "test_samples = json.dumps({\"data\": X_test[0:1, :].tolist()})\n",
+ "\n",
+ "#score on our service\n",
+ "service.run(input_data = test_samples)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "This cell shows how you can send multiple rows to the webservice at once. It then calculates the residuals - that is, the errors - by subtracting out the actual values from the results. These residuals are used later to show a plotted result."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "deploy service",
+ "aci"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "# score the entire test set.\n",
+ "test_samples = json.dumps({'data': X_test.tolist()})\n",
+ "\n",
+ "result = service.run(input_data = test_samples)\n",
+ "residual = result - y_test"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "This cell shows how you can use the `service.scoring_uri` property to access the HTTP endpoint of the service and call it using standard POST operations."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "deploy service",
+ "aci"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "import requests\n",
+ "\n",
+ "# use the first row from the test set again\n",
+ "test_samples = json.dumps({\"data\": X_test[0:1, :].tolist()})\n",
+ "\n",
+ "# create the required header\n",
+ "headers = {'Content-Type':'application/json'}\n",
+ "\n",
+ "# post the request to the service and display the result\n",
+ "resp = requests.post(service.scoring_uri, test_samples, headers = headers)\n",
+ "print(resp.text)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Residual graph\n",
+ "One way to understand the behavior of your model is to see how the data performs against data with known results. This cell uses matplotlib to create a histogram of the residual values, or errors, created from scoring the test samples.\n",
+ "\n",
+ "A good model should have residual values that cluster around 0 - that is, no error. Observing the resulting histogram can also show you if the model is skewed in any particular direction."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%matplotlib inline\n",
+ "import matplotlib.pyplot as plt\n",
+ "\n",
+ "f, (a0, a1) = plt.subplots(1, 2, gridspec_kw={'width_ratios':[3, 1], 'wspace':0, 'hspace': 0})\n",
+ "f.suptitle('Residual Values', fontsize = 18)\n",
+ "\n",
+ "f.set_figheight(6)\n",
+ "f.set_figwidth(14)\n",
+ "\n",
+ "a0.plot(residual, 'bo', alpha=0.4)\n",
+ "a0.plot([0,90], [0,0], 'r', lw=2)\n",
+ "a0.set_ylabel('residue values', fontsize=14)\n",
+ "a0.set_xlabel('test data set', fontsize=14)\n",
+ "\n",
+ "a1.hist(residual, orientation='horizontal', color='blue', bins=10, histtype='step')\n",
+ "a1.hist(residual, orientation='horizontal', color='blue', alpha=0.2, bins=10)\n",
+ "a1.set_yticklabels([])\n",
+ "\n",
+ "plt.show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Clean up"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Delete the ACI instance to stop the compute and any associated billing."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "deploy service",
+ "aci"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "%%time\n",
+ "service.delete()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "---\n",
+ "## Next Steps"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "In this example, you created a series of models inside the notebook using local data, stored them inside an AML experiment, found the best one and deployed it as a live service! From here you can continue to use Azure Machine Learning in this regard to run your own experiments and deploy your own models, or you can expand into further capabilities of AML!\n",
+ "\n",
+ "If you have a model that is difficult to process locally, either because the data is remote or the model is large, try the [train-on-remote-vm](../train-on-remote-vm) notebook to learn about submitting remote jobs.\n",
+ "\n",
+ "If you want to take advantage of multiple cloud machines to perform large parameter sweeps try the [train-hyperparameter-tune-deploy-with-pytorch](../../training-with-deep-learning/train-hyperparameter-tune-deploy-with-pytorch\n",
+ ") sample.\n",
+ "\n",
+ "If you want to deploy models to a production cluster try the [production-deploy-to-aks](../../deployment/production-deploy-to-aks\n",
+ ") notebook."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": []
+ }
],
- "kernelspec": {
- "display_name": "Python [Python 3.6]",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "roastala"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.6"
+ }
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- }
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/tutorials/img-classification-part1-training.ipynb b/tutorials/img-classification-part1-training.ipynb
index 29f32d88..1a4d2d9c 100644
--- a/tutorials/img-classification-part1-training.ipynb
+++ b/tutorials/img-classification-part1-training.ipynb
@@ -1,717 +1,717 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Tutorial #1: Train an image classification model with Azure Machine Learning\n",
- "\n",
- "In this tutorial, you train a machine learning model both locally and on remote compute resources. You'll use the training and deployment workflow for Azure Machine Learning service (preview) in a Python Jupyter notebook. You can then use the notebook as a template to train your own machine learning model with your own data. This tutorial is **part one of a two-part tutorial series**. \n",
- "\n",
- "This tutorial trains a simple logistic regression using the [MNIST](http://yann.lecun.com/exdb/mnist/) dataset and [scikit-learn](http://scikit-learn.org) with Azure Machine Learning. MNIST is a popular dataset consisting of 70,000 grayscale images. Each image is a handwritten digit of 28x28 pixels, representing a number from 0 to 9. The goal is to create a multi-class classifier to identify the digit a given image represents. \n",
- "\n",
- "Learn how to:\n",
- "\n",
- "> * Set up your development environment\n",
- "> * Access and examine the data\n",
- "> * Train a simple logistic regression model locally using the popular scikit-learn machine learning library \n",
- "> * Train multiple models on a remote cluster\n",
- "> * Review training results, find and register the best model\n",
- "\n",
- "You'll learn how to select a model and deploy it in [part two of this tutorial](deploy-models.ipynb) later. \n",
- "\n",
- "## Prerequisites\n",
- "\n",
- "Use [these instructions](https://aka.ms/aml-how-to-configure-environment) to: \n",
- "* Create a workspace and its configuration file (**config.json**) \n",
- "* Save your **config.json** to the same folder as this notebook"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Set up your development environment\n",
- "\n",
- "All the setup for your development work can be accomplished in a Python notebook. Setup includes:\n",
- "\n",
- "* Importing Python packages\n",
- "* Connecting to a workspace to enable communication between your local computer and remote resources\n",
- "* Creating an experiment to track all your runs\n",
- "* Creating a remote compute target to use for training\n",
- "\n",
- "### Import packages\n",
- "\n",
- "Import Python packages you need in this session. Also display the Azure Machine Learning SDK version."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "check version"
- ]
- },
- "outputs": [],
- "source": [
- "%matplotlib inline\n",
- "import numpy as np\n",
- "import matplotlib\n",
- "import matplotlib.pyplot as plt\n",
- "\n",
- "import azureml\n",
- "from azureml.core import Workspace, Run\n",
- "\n",
- "# check core SDK version number\n",
- "print(\"Azure ML SDK Version: \", azureml.core.VERSION)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Connect to workspace\n",
- "\n",
- "Create a workspace object from the existing workspace. `Workspace.from_config()` reads the file **config.json** and loads the details into an object named `ws`."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "load workspace"
- ]
- },
- "outputs": [],
- "source": [
- "# load workspace configuration from the config.json file in the current folder.\n",
- "ws = Workspace.from_config()\n",
- "print(ws.name, ws.location, ws.resource_group, ws.location, sep = '\\t')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create experiment\n",
- "\n",
- "Create an experiment to track the runs in your workspace. A workspace can have muliple experiments. "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "create experiment"
- ]
- },
- "outputs": [],
- "source": [
- "experiment_name = 'sklearn-mnist'\n",
- "\n",
- "from azureml.core import Experiment\n",
- "exp = Experiment(workspace=ws, name=experiment_name)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create or Attach existing AmlCompute\n",
- "You will need to create a [compute target](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#compute-target) for training your model. In this tutorial, you create `AmlCompute` as your training compute resource.\n",
- "\n",
- "**Creation of AmlCompute takes approximately 5 minutes.** If the AmlCompute with that name is already in your workspace this code will skip the creation process."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "create mlc",
- "amlcompute"
- ]
- },
- "outputs": [],
- "source": [
- "from azureml.core.compute import AmlCompute\n",
- "from azureml.core.compute import ComputeTarget\n",
- "import os\n",
- "\n",
- "# choose a name for your cluster\n",
- "compute_name = os.environ.get(\"AML_COMPUTE_CLUSTER_NAME\", \"cpucluster\")\n",
- "compute_min_nodes = os.environ.get(\"AML_COMPUTE_CLUSTER_MIN_NODES\", 0)\n",
- "compute_max_nodes = os.environ.get(\"AML_COMPUTE_CLUSTER_MAX_NODES\", 4)\n",
- "\n",
- "# This example uses CPU VM. For using GPU VM, set SKU to STANDARD_NC6\n",
- "vm_size = os.environ.get(\"AML_COMPUTE_CLUSTER_SKU\", \"STANDARD_D2_V2\")\n",
- "\n",
- "\n",
- "if compute_name in ws.compute_targets:\n",
- " compute_target = ws.compute_targets[compute_name]\n",
- " if compute_target and type(compute_target) is AmlCompute:\n",
- " print('found compute target. just use it. ' + compute_name)\n",
- "else:\n",
- " print('creating a new compute target...')\n",
- " provisioning_config = AmlCompute.provisioning_configuration(vm_size = vm_size,\n",
- " min_nodes = compute_min_nodes, \n",
- " max_nodes = compute_max_nodes)\n",
- "\n",
- " # create the cluster\n",
- " compute_target = ComputeTarget.create(ws, compute_name, provisioning_config)\n",
- " \n",
- " # can poll for a minimum number of nodes and for a specific timeout. \n",
- " # if no min node count is provided it will use the scale settings for the cluster\n",
- " compute_target.wait_for_completion(show_output=True, min_node_count=None, timeout_in_minutes=20)\n",
- " \n",
- " # For a more detailed view of current AmlCompute status, use the 'status' property \n",
- " print(compute_target.status.serialize())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "You now have the necessary packages and compute resources to train a model in the cloud. \n",
- "\n",
- "## Explore data\n",
- "\n",
- "Before you train a model, you need to understand the data that you are using to train it. You also need to copy the data into the cloud so it can be accessed by your cloud training environment. In this section you learn how to:\n",
- "\n",
- "* Download the MNIST dataset\n",
- "* Display some sample images\n",
- "* Upload data to the cloud\n",
- "\n",
- "### Download the MNIST dataset\n",
- "\n",
- "Download the MNIST dataset and save the files into a `data` directory locally. Images and labels for both training and testing are downloaded."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import os\n",
- "import urllib.request\n",
- "\n",
- "os.makedirs('./data', exist_ok = True)\n",
- "\n",
- "urllib.request.urlretrieve('http://yann.lecun.com/exdb/mnist/train-images-idx3-ubyte.gz', filename='./data/train-images.gz')\n",
- "urllib.request.urlretrieve('http://yann.lecun.com/exdb/mnist/train-labels-idx1-ubyte.gz', filename='./data/train-labels.gz')\n",
- "urllib.request.urlretrieve('http://yann.lecun.com/exdb/mnist/t10k-images-idx3-ubyte.gz', filename='./data/test-images.gz')\n",
- "urllib.request.urlretrieve('http://yann.lecun.com/exdb/mnist/t10k-labels-idx1-ubyte.gz', filename='./data/test-labels.gz')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Display some sample images\n",
- "\n",
- "Load the compressed files into `numpy` arrays. Then use `matplotlib` to plot 30 random images from the dataset with their labels above them. Note this step requires a `load_data` function that's included in an `util.py` file. This file is included in the sample folder. Please make sure it is placed in the same folder as this notebook. The `load_data` function simply parses the compresse files into numpy arrays."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# make sure utils.py is in the same directory as this code\n",
- "from utils import load_data\n",
- "\n",
- "# note we also shrink the intensity values (X) from 0-255 to 0-1. This helps the model converge faster.\n",
- "X_train = load_data('./data/train-images.gz', False) / 255.0\n",
- "y_train = load_data('./data/train-labels.gz', True).reshape(-1)\n",
- "\n",
- "X_test = load_data('./data/test-images.gz', False) / 255.0\n",
- "y_test = load_data('./data/test-labels.gz', True).reshape(-1)\n",
- "\n",
- "# now let's show some randomly chosen images from the traininng set.\n",
- "count = 0\n",
- "sample_size = 30\n",
- "plt.figure(figsize = (16, 6))\n",
- "for i in np.random.permutation(X_train.shape[0])[:sample_size]:\n",
- " count = count + 1\n",
- " plt.subplot(1, sample_size, count)\n",
- " plt.axhline('')\n",
- " plt.axvline('')\n",
- " plt.text(x=10, y=-10, s=y_train[i], fontsize=18)\n",
- " plt.imshow(X_train[i].reshape(28, 28), cmap=plt.cm.Greys)\n",
- "plt.show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Now you have an idea of what these images look like and the expected prediction outcome.\n",
- "\n",
- "### Upload data to the cloud\n",
- "\n",
- "Now make the data accessible remotely by uploading that data from your local machine into Azure so it can be accessed for remote training. The datastore is a convenient construct associated with your workspace for you to upload/download data, and interact with it from your remote compute targets. It is backed by Azure blob storage account.\n",
- "\n",
- "The MNIST files are uploaded into a directory named `mnist` at the root of the datastore."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "use datastore"
- ]
- },
- "outputs": [],
- "source": [
- "ds = ws.get_default_datastore()\n",
- "print(ds.datastore_type, ds.account_name, ds.container_name)\n",
- "\n",
- "ds.upload(src_dir='./data', target_path='mnist', overwrite=True, show_progress=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "You now have everything you need to start training a model. \n",
- "\n",
- "## Train a local model\n",
- "\n",
- "Train a simple logistic regression model using scikit-learn locally.\n",
- "\n",
- "**Training locally can take a minute or two** depending on your computer configuration."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%time\n",
- "from sklearn.linear_model import LogisticRegression\n",
- "\n",
- "clf = LogisticRegression()\n",
- "clf.fit(X_train, y_train)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Next, make predictions using the test set and calculate the accuracy."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "y_hat = clf.predict(X_test)\n",
- "print(np.average(y_hat == y_test))"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "With just a few lines of code, you have a 92% accuracy.\n",
- "\n",
- "## Train on a remote cluster\n",
- "\n",
- "Now you can expand on this simple model by building a model with a different regularization rate. This time you'll train the model on a remote resource. \n",
- "\n",
- "For this task, submit the job to the remote training cluster you set up earlier. To submit a job you:\n",
- "* Create a directory\n",
- "* Create a training script\n",
- "* Create an estimator object\n",
- "* Submit the job \n",
- "\n",
- "### Create a directory\n",
- "\n",
- "Create a directory to deliver the necessary code from your computer to the remote resource."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import os\n",
- "script_folder = './sklearn-mnist'\n",
- "os.makedirs(script_folder, exist_ok=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create a training script\n",
- "\n",
- "To submit the job to the cluster, first create a training script. Run the following code to create the training script called `train.py` in the directory you just created. This training adds a regularization rate to the training algorithm, so produces a slightly different model than the local version."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%writefile $script_folder/train.py\n",
- "\n",
- "import argparse\n",
- "import os\n",
- "import numpy as np\n",
- "\n",
- "from sklearn.linear_model import LogisticRegression\n",
- "from sklearn.externals import joblib\n",
- "\n",
- "from azureml.core import Run\n",
- "from utils import load_data\n",
- "\n",
- "# let user feed in 2 parameters, the location of the data files (from datastore), and the regularization rate of the logistic regression model\n",
- "parser = argparse.ArgumentParser()\n",
- "parser.add_argument('--data-folder', type=str, dest='data_folder', help='data folder mounting point')\n",
- "parser.add_argument('--regularization', type=float, dest='reg', default=0.01, help='regularization rate')\n",
- "args = parser.parse_args()\n",
- "\n",
- "data_folder = os.path.join(args.data_folder, 'mnist')\n",
- "print('Data folder:', data_folder)\n",
- "\n",
- "# load train and test set into numpy arrays\n",
- "# note we scale the pixel intensity values to 0-1 (by dividing it with 255.0) so the model can converge faster.\n",
- "X_train = load_data(os.path.join(data_folder, 'train-images.gz'), False) / 255.0\n",
- "X_test = load_data(os.path.join(data_folder, 'test-images.gz'), False) / 255.0\n",
- "y_train = load_data(os.path.join(data_folder, 'train-labels.gz'), True).reshape(-1)\n",
- "y_test = load_data(os.path.join(data_folder, 'test-labels.gz'), True).reshape(-1)\n",
- "print(X_train.shape, y_train.shape, X_test.shape, y_test.shape, sep = '\\n')\n",
- "\n",
- "# get hold of the current run\n",
- "run = Run.get_context()\n",
- "\n",
- "print('Train a logistic regression model with regularizaion rate of', args.reg)\n",
- "clf = LogisticRegression(C=1.0/args.reg, random_state=42)\n",
- "clf.fit(X_train, y_train)\n",
- "\n",
- "print('Predict the test set')\n",
- "y_hat = clf.predict(X_test)\n",
- "\n",
- "# calculate accuracy on the prediction\n",
- "acc = np.average(y_hat == y_test)\n",
- "print('Accuracy is', acc)\n",
- "\n",
- "run.log('regularization rate', np.float(args.reg))\n",
- "run.log('accuracy', np.float(acc))\n",
- "\n",
- "os.makedirs('outputs', exist_ok=True)\n",
- "# note file saved in the outputs folder is automatically uploaded into experiment record\n",
- "joblib.dump(value=clf, filename='outputs/sklearn_mnist_model.pkl')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Notice how the script gets data and saves models:\n",
- "\n",
- "+ The training script reads an argument to find the directory containing the data. When you submit the job later, you point to the datastore for this argument:\n",
- "`parser.add_argument('--data-folder', type=str, dest='data_folder', help='data directory mounting point')`"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "\n",
- "+ The training script saves your model into a directory named outputs.
\n",
- "`joblib.dump(value=clf, filename='outputs/sklearn_mnist_model.pkl')`
\n",
- "Anything written in this directory is automatically uploaded into your workspace. You'll access your model from this directory later in the tutorial."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "The file `utils.py` is referenced from the training script to load the dataset correctly. Copy this script into the script folder so that it can be accessed along with the training script on the remote resource."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import shutil\n",
- "shutil.copy('utils.py', script_folder)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create an estimator\n",
- "\n",
- "An estimator object is used to submit the run. Create your estimator by running the following code to define:\n",
- "\n",
- "* The name of the estimator object, `est`\n",
- "* The directory that contains your scripts. All the files in this directory are uploaded into the cluster nodes for execution. \n",
- "* The compute target. In this case you will use the AmlCompute you created\n",
- "* The training script name, train.py\n",
- "* Parameters required from the training script \n",
- "* Python packages needed for training\n",
- "\n",
- "In this tutorial, this target is AmlCompute. All files in the script folder are uploaded into the cluster nodes for execution. The data_folder is set to use the datastore (`ds.as_mount()`)."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "configure estimator"
- ]
- },
- "outputs": [],
- "source": [
- "from azureml.train.estimator import Estimator\n",
- "\n",
- "script_params = {\n",
- " '--data-folder': ds.as_mount(),\n",
- " '--regularization': 0.8\n",
- "}\n",
- "\n",
- "est = Estimator(source_directory=script_folder,\n",
- " script_params=script_params,\n",
- " compute_target=compute_target,\n",
- " entry_script='train.py',\n",
- " conda_packages=['scikit-learn'])"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Submit the job to the cluster\n",
- "\n",
- "Run the experiment by submitting the estimator object."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "remote run",
- "amlcompute",
- "scikit-learn"
- ]
- },
- "outputs": [],
- "source": [
- "run = exp.submit(config=est)\n",
- "run"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Since the call is asynchronous, it returns a **Preparing** or **Running** state as soon as the job is started.\n",
- "\n",
- "## Monitor a remote run\n",
- "\n",
- "In total, the first run takes **approximately 10 minutes**. But for subsequent runs, as long as the script dependencies don't change, the same image is reused and hence the container start up time is much faster.\n",
- "\n",
- "Here is what's happening while you wait:\n",
- "\n",
- "- **Image creation**: A Docker image is created matching the Python environment specified by the estimator. The image is uploaded to the workspace. Image creation and uploading takes **about 5 minutes**. \n",
- "\n",
- " This stage happens once for each Python environment since the container is cached for subsequent runs. During image creation, logs are streamed to the run history. You can monitor the image creation progress using these logs.\n",
- "\n",
- "- **Scaling**: If the remote cluster requires more nodes to execute the run than currently available, additional nodes are added automatically. Scaling typically takes **about 5 minutes.**\n",
- "\n",
- "- **Running**: In this stage, the necessary scripts and files are sent to the compute target, then data stores are mounted/copied, then the entry_script is run. While the job is running, stdout and the ./logs directory are streamed to the run history. You can monitor the run's progress using these logs.\n",
- "\n",
- "- **Post-Processing**: The ./outputs directory of the run is copied over to the run history in your workspace so you can access these results.\n",
- "\n",
- "\n",
- "You can check the progress of a running job in multiple ways. This tutorial uses a Jupyter widget as well as a `wait_for_completion` method. \n",
- "\n",
- "### Jupyter widget\n",
- "\n",
- "Watch the progress of the run with a Jupyter widget. Like the run submission, the widget is asynchronous and provides live updates every 10-15 seconds until the job completes."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "use notebook widget"
- ]
- },
- "outputs": [],
- "source": [
- "from azureml.widgets import RunDetails\n",
- "RunDetails(run).show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Get log results upon completion\n",
- "\n",
- "Model training and monitoring happen in the background. Wait until the model has completed training before running more code. Use `wait_for_completion` to show when the model training is complete."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "remote run",
- "amlcompute",
- "scikit-learn"
- ]
- },
- "outputs": [],
- "source": [
- "run.wait_for_completion(show_output=False) # specify True for a verbose log"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Display run results\n",
- "\n",
- "You now have a model trained on a remote cluster. Retrieve the accuracy of the model:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "get metrics"
- ]
- },
- "outputs": [],
- "source": [
- "print(run.get_metrics())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "In the next tutorial you will explore this model in more detail.\n",
- "\n",
- "## Register model\n",
- "\n",
- "The last step in the training script wrote the file `outputs/sklearn_mnist_model.pkl` in a directory named `outputs` in the VM of the cluster where the job is executed. `outputs` is a special directory in that all content in this directory is automatically uploaded to your workspace. This content appears in the run record in the experiment under your workspace. Hence, the model file is now also available in your workspace.\n",
- "\n",
- "You can see files associated with that run."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "query history"
- ]
- },
- "outputs": [],
- "source": [
- "print(run.get_file_names())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Register the model in the workspace so that you (or other collaborators) can later query, examine, and deploy this model."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "register model from history"
- ]
- },
- "outputs": [],
- "source": [
- "# register model \n",
- "model = run.register_model(model_name='sklearn_mnist', model_path='outputs/sklearn_mnist_model.pkl')\n",
- "print(model.name, model.id, model.version, sep = '\\t')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Next steps\n",
- "\n",
- "In this Azure Machine Learning tutorial, you used Python to:\n",
- "\n",
- "> * Set up your development environment\n",
- "> * Access and examine the data\n",
- "> * Train a simple logistic regression locally using the popular scikit-learn machine learning library\n",
- "> * Train multiple models on a remote cluster\n",
- "> * Review training details and register the best model\n",
- "\n",
- "You are ready to deploy this registered model using the instructions in the next part of the tutorial series:\n",
- "\n",
- "> [Tutorial 2 - Deploy models](img-classification-part2-deploy.ipynb)"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "roastala"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Tutorial #1: Train an image classification model with Azure Machine Learning\n",
+ "\n",
+ "In this tutorial, you train a machine learning model both locally and on remote compute resources. You'll use the training and deployment workflow for Azure Machine Learning service (preview) in a Python Jupyter notebook. You can then use the notebook as a template to train your own machine learning model with your own data. This tutorial is **part one of a two-part tutorial series**. \n",
+ "\n",
+ "This tutorial trains a simple logistic regression using the [MNIST](http://yann.lecun.com/exdb/mnist/) dataset and [scikit-learn](http://scikit-learn.org) with Azure Machine Learning. MNIST is a popular dataset consisting of 70,000 grayscale images. Each image is a handwritten digit of 28x28 pixels, representing a number from 0 to 9. The goal is to create a multi-class classifier to identify the digit a given image represents. \n",
+ "\n",
+ "Learn how to:\n",
+ "\n",
+ "> * Set up your development environment\n",
+ "> * Access and examine the data\n",
+ "> * Train a simple logistic regression model locally using the popular scikit-learn machine learning library \n",
+ "> * Train multiple models on a remote cluster\n",
+ "> * Review training results, find and register the best model\n",
+ "\n",
+ "You'll learn how to select a model and deploy it in [part two of this tutorial](deploy-models.ipynb) later. \n",
+ "\n",
+ "## Prerequisites\n",
+ "\n",
+ "Use [these instructions](https://aka.ms/aml-how-to-configure-environment) to: \n",
+ "* Create a workspace and its configuration file (**config.json**) \n",
+ "* Save your **config.json** to the same folder as this notebook"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Set up your development environment\n",
+ "\n",
+ "All the setup for your development work can be accomplished in a Python notebook. Setup includes:\n",
+ "\n",
+ "* Importing Python packages\n",
+ "* Connecting to a workspace to enable communication between your local computer and remote resources\n",
+ "* Creating an experiment to track all your runs\n",
+ "* Creating a remote compute target to use for training\n",
+ "\n",
+ "### Import packages\n",
+ "\n",
+ "Import Python packages you need in this session. Also display the Azure Machine Learning SDK version."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "check version"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "%matplotlib inline\n",
+ "import numpy as np\n",
+ "import matplotlib\n",
+ "import matplotlib.pyplot as plt\n",
+ "\n",
+ "import azureml\n",
+ "from azureml.core import Workspace, Run\n",
+ "\n",
+ "# check core SDK version number\n",
+ "print(\"Azure ML SDK Version: \", azureml.core.VERSION)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Connect to workspace\n",
+ "\n",
+ "Create a workspace object from the existing workspace. `Workspace.from_config()` reads the file **config.json** and loads the details into an object named `ws`."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "load workspace"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "# load workspace configuration from the config.json file in the current folder.\n",
+ "ws = Workspace.from_config()\n",
+ "print(ws.name, ws.location, ws.resource_group, ws.location, sep = '\\t')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create experiment\n",
+ "\n",
+ "Create an experiment to track the runs in your workspace. A workspace can have muliple experiments. "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "create experiment"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "experiment_name = 'sklearn-mnist'\n",
+ "\n",
+ "from azureml.core import Experiment\n",
+ "exp = Experiment(workspace=ws, name=experiment_name)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create or Attach existing AmlCompute\n",
+ "You will need to create a [compute target](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#compute-target) for training your model. In this tutorial, you create `AmlCompute` as your training compute resource.\n",
+ "\n",
+ "**Creation of AmlCompute takes approximately 5 minutes.** If the AmlCompute with that name is already in your workspace this code will skip the creation process."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "create mlc",
+ "amlcompute"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "from azureml.core.compute import AmlCompute\n",
+ "from azureml.core.compute import ComputeTarget\n",
+ "import os\n",
+ "\n",
+ "# choose a name for your cluster\n",
+ "compute_name = os.environ.get(\"AML_COMPUTE_CLUSTER_NAME\", \"cpucluster\")\n",
+ "compute_min_nodes = os.environ.get(\"AML_COMPUTE_CLUSTER_MIN_NODES\", 0)\n",
+ "compute_max_nodes = os.environ.get(\"AML_COMPUTE_CLUSTER_MAX_NODES\", 4)\n",
+ "\n",
+ "# This example uses CPU VM. For using GPU VM, set SKU to STANDARD_NC6\n",
+ "vm_size = os.environ.get(\"AML_COMPUTE_CLUSTER_SKU\", \"STANDARD_D2_V2\")\n",
+ "\n",
+ "\n",
+ "if compute_name in ws.compute_targets:\n",
+ " compute_target = ws.compute_targets[compute_name]\n",
+ " if compute_target and type(compute_target) is AmlCompute:\n",
+ " print('found compute target. just use it. ' + compute_name)\n",
+ "else:\n",
+ " print('creating a new compute target...')\n",
+ " provisioning_config = AmlCompute.provisioning_configuration(vm_size = vm_size,\n",
+ " min_nodes = compute_min_nodes, \n",
+ " max_nodes = compute_max_nodes)\n",
+ "\n",
+ " # create the cluster\n",
+ " compute_target = ComputeTarget.create(ws, compute_name, provisioning_config)\n",
+ " \n",
+ " # can poll for a minimum number of nodes and for a specific timeout. \n",
+ " # if no min node count is provided it will use the scale settings for the cluster\n",
+ " compute_target.wait_for_completion(show_output=True, min_node_count=None, timeout_in_minutes=20)\n",
+ " \n",
+ " # For a more detailed view of current AmlCompute status, use get_status()\n",
+ " print(compute_target.get_status().serialize())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "You now have the necessary packages and compute resources to train a model in the cloud. \n",
+ "\n",
+ "## Explore data\n",
+ "\n",
+ "Before you train a model, you need to understand the data that you are using to train it. You also need to copy the data into the cloud so it can be accessed by your cloud training environment. In this section you learn how to:\n",
+ "\n",
+ "* Download the MNIST dataset\n",
+ "* Display some sample images\n",
+ "* Upload data to the cloud\n",
+ "\n",
+ "### Download the MNIST dataset\n",
+ "\n",
+ "Download the MNIST dataset and save the files into a `data` directory locally. Images and labels for both training and testing are downloaded."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import os\n",
+ "import urllib.request\n",
+ "\n",
+ "os.makedirs('./data', exist_ok = True)\n",
+ "\n",
+ "urllib.request.urlretrieve('http://yann.lecun.com/exdb/mnist/train-images-idx3-ubyte.gz', filename='./data/train-images.gz')\n",
+ "urllib.request.urlretrieve('http://yann.lecun.com/exdb/mnist/train-labels-idx1-ubyte.gz', filename='./data/train-labels.gz')\n",
+ "urllib.request.urlretrieve('http://yann.lecun.com/exdb/mnist/t10k-images-idx3-ubyte.gz', filename='./data/test-images.gz')\n",
+ "urllib.request.urlretrieve('http://yann.lecun.com/exdb/mnist/t10k-labels-idx1-ubyte.gz', filename='./data/test-labels.gz')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Display some sample images\n",
+ "\n",
+ "Load the compressed files into `numpy` arrays. Then use `matplotlib` to plot 30 random images from the dataset with their labels above them. Note this step requires a `load_data` function that's included in an `util.py` file. This file is included in the sample folder. Please make sure it is placed in the same folder as this notebook. The `load_data` function simply parses the compresse files into numpy arrays."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# make sure utils.py is in the same directory as this code\n",
+ "from utils import load_data\n",
+ "\n",
+ "# note we also shrink the intensity values (X) from 0-255 to 0-1. This helps the model converge faster.\n",
+ "X_train = load_data('./data/train-images.gz', False) / 255.0\n",
+ "y_train = load_data('./data/train-labels.gz', True).reshape(-1)\n",
+ "\n",
+ "X_test = load_data('./data/test-images.gz', False) / 255.0\n",
+ "y_test = load_data('./data/test-labels.gz', True).reshape(-1)\n",
+ "\n",
+ "# now let's show some randomly chosen images from the traininng set.\n",
+ "count = 0\n",
+ "sample_size = 30\n",
+ "plt.figure(figsize = (16, 6))\n",
+ "for i in np.random.permutation(X_train.shape[0])[:sample_size]:\n",
+ " count = count + 1\n",
+ " plt.subplot(1, sample_size, count)\n",
+ " plt.axhline('')\n",
+ " plt.axvline('')\n",
+ " plt.text(x=10, y=-10, s=y_train[i], fontsize=18)\n",
+ " plt.imshow(X_train[i].reshape(28, 28), cmap=plt.cm.Greys)\n",
+ "plt.show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Now you have an idea of what these images look like and the expected prediction outcome.\n",
+ "\n",
+ "### Upload data to the cloud\n",
+ "\n",
+ "Now make the data accessible remotely by uploading that data from your local machine into Azure so it can be accessed for remote training. The datastore is a convenient construct associated with your workspace for you to upload/download data, and interact with it from your remote compute targets. It is backed by Azure blob storage account.\n",
+ "\n",
+ "The MNIST files are uploaded into a directory named `mnist` at the root of the datastore."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "use datastore"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "ds = ws.get_default_datastore()\n",
+ "print(ds.datastore_type, ds.account_name, ds.container_name)\n",
+ "\n",
+ "ds.upload(src_dir='./data', target_path='mnist', overwrite=True, show_progress=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "You now have everything you need to start training a model. \n",
+ "\n",
+ "## Train a local model\n",
+ "\n",
+ "Train a simple logistic regression model using scikit-learn locally.\n",
+ "\n",
+ "**Training locally can take a minute or two** depending on your computer configuration."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%time\n",
+ "from sklearn.linear_model import LogisticRegression\n",
+ "\n",
+ "clf = LogisticRegression()\n",
+ "clf.fit(X_train, y_train)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Next, make predictions using the test set and calculate the accuracy."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "y_hat = clf.predict(X_test)\n",
+ "print(np.average(y_hat == y_test))"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "With just a few lines of code, you have a 92% accuracy.\n",
+ "\n",
+ "## Train on a remote cluster\n",
+ "\n",
+ "Now you can expand on this simple model by building a model with a different regularization rate. This time you'll train the model on a remote resource. \n",
+ "\n",
+ "For this task, submit the job to the remote training cluster you set up earlier. To submit a job you:\n",
+ "* Create a directory\n",
+ "* Create a training script\n",
+ "* Create an estimator object\n",
+ "* Submit the job \n",
+ "\n",
+ "### Create a directory\n",
+ "\n",
+ "Create a directory to deliver the necessary code from your computer to the remote resource."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import os\n",
+ "script_folder = './sklearn-mnist'\n",
+ "os.makedirs(script_folder, exist_ok=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create a training script\n",
+ "\n",
+ "To submit the job to the cluster, first create a training script. Run the following code to create the training script called `train.py` in the directory you just created. This training adds a regularization rate to the training algorithm, so produces a slightly different model than the local version."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%writefile $script_folder/train.py\n",
+ "\n",
+ "import argparse\n",
+ "import os\n",
+ "import numpy as np\n",
+ "\n",
+ "from sklearn.linear_model import LogisticRegression\n",
+ "from sklearn.externals import joblib\n",
+ "\n",
+ "from azureml.core import Run\n",
+ "from utils import load_data\n",
+ "\n",
+ "# let user feed in 2 parameters, the location of the data files (from datastore), and the regularization rate of the logistic regression model\n",
+ "parser = argparse.ArgumentParser()\n",
+ "parser.add_argument('--data-folder', type=str, dest='data_folder', help='data folder mounting point')\n",
+ "parser.add_argument('--regularization', type=float, dest='reg', default=0.01, help='regularization rate')\n",
+ "args = parser.parse_args()\n",
+ "\n",
+ "data_folder = os.path.join(args.data_folder, 'mnist')\n",
+ "print('Data folder:', data_folder)\n",
+ "\n",
+ "# load train and test set into numpy arrays\n",
+ "# note we scale the pixel intensity values to 0-1 (by dividing it with 255.0) so the model can converge faster.\n",
+ "X_train = load_data(os.path.join(data_folder, 'train-images.gz'), False) / 255.0\n",
+ "X_test = load_data(os.path.join(data_folder, 'test-images.gz'), False) / 255.0\n",
+ "y_train = load_data(os.path.join(data_folder, 'train-labels.gz'), True).reshape(-1)\n",
+ "y_test = load_data(os.path.join(data_folder, 'test-labels.gz'), True).reshape(-1)\n",
+ "print(X_train.shape, y_train.shape, X_test.shape, y_test.shape, sep = '\\n')\n",
+ "\n",
+ "# get hold of the current run\n",
+ "run = Run.get_context()\n",
+ "\n",
+ "print('Train a logistic regression model with regularizaion rate of', args.reg)\n",
+ "clf = LogisticRegression(C=1.0/args.reg, random_state=42)\n",
+ "clf.fit(X_train, y_train)\n",
+ "\n",
+ "print('Predict the test set')\n",
+ "y_hat = clf.predict(X_test)\n",
+ "\n",
+ "# calculate accuracy on the prediction\n",
+ "acc = np.average(y_hat == y_test)\n",
+ "print('Accuracy is', acc)\n",
+ "\n",
+ "run.log('regularization rate', np.float(args.reg))\n",
+ "run.log('accuracy', np.float(acc))\n",
+ "\n",
+ "os.makedirs('outputs', exist_ok=True)\n",
+ "# note file saved in the outputs folder is automatically uploaded into experiment record\n",
+ "joblib.dump(value=clf, filename='outputs/sklearn_mnist_model.pkl')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Notice how the script gets data and saves models:\n",
+ "\n",
+ "+ The training script reads an argument to find the directory containing the data. When you submit the job later, you point to the datastore for this argument:\n",
+ "`parser.add_argument('--data-folder', type=str, dest='data_folder', help='data directory mounting point')`"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "\n",
+ "+ The training script saves your model into a directory named outputs.
\n",
+ "`joblib.dump(value=clf, filename='outputs/sklearn_mnist_model.pkl')`
\n",
+ "Anything written in this directory is automatically uploaded into your workspace. You'll access your model from this directory later in the tutorial."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "The file `utils.py` is referenced from the training script to load the dataset correctly. Copy this script into the script folder so that it can be accessed along with the training script on the remote resource."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import shutil\n",
+ "shutil.copy('utils.py', script_folder)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create an estimator\n",
+ "\n",
+ "An estimator object is used to submit the run. Create your estimator by running the following code to define:\n",
+ "\n",
+ "* The name of the estimator object, `est`\n",
+ "* The directory that contains your scripts. All the files in this directory are uploaded into the cluster nodes for execution. \n",
+ "* The compute target. In this case you will use the AmlCompute you created\n",
+ "* The training script name, train.py\n",
+ "* Parameters required from the training script \n",
+ "* Python packages needed for training\n",
+ "\n",
+ "In this tutorial, this target is AmlCompute. All files in the script folder are uploaded into the cluster nodes for execution. The data_folder is set to use the datastore (`ds.as_mount()`)."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "configure estimator"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "from azureml.train.estimator import Estimator\n",
+ "\n",
+ "script_params = {\n",
+ " '--data-folder': ds.as_mount(),\n",
+ " '--regularization': 0.8\n",
+ "}\n",
+ "\n",
+ "est = Estimator(source_directory=script_folder,\n",
+ " script_params=script_params,\n",
+ " compute_target=compute_target,\n",
+ " entry_script='train.py',\n",
+ " conda_packages=['scikit-learn'])"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Submit the job to the cluster\n",
+ "\n",
+ "Run the experiment by submitting the estimator object."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "remote run",
+ "amlcompute",
+ "scikit-learn"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "run = exp.submit(config=est)\n",
+ "run"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Since the call is asynchronous, it returns a **Preparing** or **Running** state as soon as the job is started.\n",
+ "\n",
+ "## Monitor a remote run\n",
+ "\n",
+ "In total, the first run takes **approximately 10 minutes**. But for subsequent runs, as long as the script dependencies don't change, the same image is reused and hence the container start up time is much faster.\n",
+ "\n",
+ "Here is what's happening while you wait:\n",
+ "\n",
+ "- **Image creation**: A Docker image is created matching the Python environment specified by the estimator. The image is uploaded to the workspace. Image creation and uploading takes **about 5 minutes**. \n",
+ "\n",
+ " This stage happens once for each Python environment since the container is cached for subsequent runs. During image creation, logs are streamed to the run history. You can monitor the image creation progress using these logs.\n",
+ "\n",
+ "- **Scaling**: If the remote cluster requires more nodes to execute the run than currently available, additional nodes are added automatically. Scaling typically takes **about 5 minutes.**\n",
+ "\n",
+ "- **Running**: In this stage, the necessary scripts and files are sent to the compute target, then data stores are mounted/copied, then the entry_script is run. While the job is running, stdout and the ./logs directory are streamed to the run history. You can monitor the run's progress using these logs.\n",
+ "\n",
+ "- **Post-Processing**: The ./outputs directory of the run is copied over to the run history in your workspace so you can access these results.\n",
+ "\n",
+ "\n",
+ "You can check the progress of a running job in multiple ways. This tutorial uses a Jupyter widget as well as a `wait_for_completion` method. \n",
+ "\n",
+ "### Jupyter widget\n",
+ "\n",
+ "Watch the progress of the run with a Jupyter widget. Like the run submission, the widget is asynchronous and provides live updates every 10-15 seconds until the job completes."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "use notebook widget"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "from azureml.widgets import RunDetails\n",
+ "RunDetails(run).show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Get log results upon completion\n",
+ "\n",
+ "Model training and monitoring happen in the background. Wait until the model has completed training before running more code. Use `wait_for_completion` to show when the model training is complete."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "remote run",
+ "amlcompute",
+ "scikit-learn"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "run.wait_for_completion(show_output=False) # specify True for a verbose log"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Display run results\n",
+ "\n",
+ "You now have a model trained on a remote cluster. Retrieve the accuracy of the model:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "get metrics"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "print(run.get_metrics())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "In the next tutorial you will explore this model in more detail.\n",
+ "\n",
+ "## Register model\n",
+ "\n",
+ "The last step in the training script wrote the file `outputs/sklearn_mnist_model.pkl` in a directory named `outputs` in the VM of the cluster where the job is executed. `outputs` is a special directory in that all content in this directory is automatically uploaded to your workspace. This content appears in the run record in the experiment under your workspace. Hence, the model file is now also available in your workspace.\n",
+ "\n",
+ "You can see files associated with that run."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "query history"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "print(run.get_file_names())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Register the model in the workspace so that you (or other collaborators) can later query, examine, and deploy this model."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "register model from history"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "# register model \n",
+ "model = run.register_model(model_name='sklearn_mnist', model_path='outputs/sklearn_mnist_model.pkl')\n",
+ "print(model.name, model.id, model.version, sep = '\\t')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Next steps\n",
+ "\n",
+ "In this Azure Machine Learning tutorial, you used Python to:\n",
+ "\n",
+ "> * Set up your development environment\n",
+ "> * Access and examine the data\n",
+ "> * Train a simple logistic regression locally using the popular scikit-learn machine learning library\n",
+ "> * Train multiple models on a remote cluster\n",
+ "> * Review training details and register the best model\n",
+ "\n",
+ "You are ready to deploy this registered model using the instructions in the next part of the tutorial series:\n",
+ "\n",
+ "> [Tutorial 2 - Deploy models](img-classification-part2-deploy.ipynb)"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "roastala"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.2"
+ },
+ "msauthor": "sgilley"
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.2"
- },
- "msauthor": "sgilley"
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/tutorials/img-classification-part2-deploy.ipynb b/tutorials/img-classification-part2-deploy.ipynb
index 65537187..3ec090c4 100644
--- a/tutorials/img-classification-part2-deploy.ipynb
+++ b/tutorials/img-classification-part2-deploy.ipynb
@@ -1,615 +1,615 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Tutorial #2: Deploy an image classification model in Azure Container Instance (ACI)\n",
- "\n",
- "This tutorial is **part two of a two-part tutorial series**. In the [previous tutorial](img-classification-part1-training.ipynb), you trained machine learning models and then registered a model in your workspace on the cloud. \n",
- "\n",
- "Now, you're ready to deploy the model as a web service in [Azure Container Instances](https://docs.microsoft.com/azure/container-instances/) (ACI). A web service is an image, in this case a Docker image, that encapsulates the scoring logic and the model itself. \n",
- "\n",
- "In this part of the tutorial, you use Azure Machine Learning service (Preview) to:\n",
- "\n",
- "> * Set up your testing environment\n",
- "> * Retrieve the model from your workspace\n",
- "> * Test the model locally\n",
- "> * Deploy the model to ACI\n",
- "> * Test the deployed model\n",
- "\n",
- "ACI is not ideal for production deployments, but it is great for testing and understanding the workflow. For scalable production deployments, consider using AKS.\n",
- "\n",
- "\n",
- "## Prerequisites\n",
- "\n",
- "Complete the model training in the [Tutorial #1: Train an image classification model with Azure Machine Learning](train-models.ipynb) notebook. \n",
- "\n"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "register model from file"
- ]
- },
- "outputs": [],
- "source": [
- "# If you did NOT complete the tutorial, you can instead run this cell \n",
- "# This will register a model and download the data needed for this tutorial\n",
- "# These prerequisites are created in the training tutorial\n",
- "# Feel free to skip this cell if you completed the training tutorial \n",
- "\n",
- "# register a model\n",
- "from azureml.core import Workspace\n",
- "ws = Workspace.from_config()\n",
- "\n",
- "from azureml.core.model import Model\n",
- "\n",
- "model_name = \"sklearn_mnist\"\n",
- "model = Model.register(model_path=\"sklearn_mnist_model.pkl\",\n",
- " model_name=model_name,\n",
- " tags={\"data\": \"mnist\", \"model\": \"classification\"},\n",
- " description=\"Mnist handwriting recognition\",\n",
- " workspace=ws)\n",
- "\n",
- "# download test data\n",
- "import os\n",
- "import urllib.request\n",
- "\n",
- "os.makedirs('./data', exist_ok=True)\n",
- "\n",
- "urllib.request.urlretrieve('http://yann.lecun.com/exdb/mnist/t10k-images-idx3-ubyte.gz', filename='./data/test-images.gz')\n",
- "urllib.request.urlretrieve('http://yann.lecun.com/exdb/mnist/t10k-labels-idx1-ubyte.gz', filename='./data/test-labels.gz')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Set up the environment\n",
- "\n",
- "Start by setting up a testing environment.\n",
- "\n",
- "### Import packages\n",
- "\n",
- "Import the Python packages needed for this tutorial."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "check version"
- ]
- },
- "outputs": [],
- "source": [
- "%matplotlib inline\n",
- "import numpy as np\n",
- "import matplotlib\n",
- "import matplotlib.pyplot as plt\n",
- " \n",
- "import azureml\n",
- "from azureml.core import Workspace, Run\n",
- "\n",
- "# display the core SDK version number\n",
- "print(\"Azure ML SDK Version: \", azureml.core.VERSION)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Retrieve the model\n",
- "\n",
- "You registered a model in your workspace in the previous tutorial. Now, load this workspace and download the model to your local directory."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "load workspace",
- "download model"
- ]
- },
- "outputs": [],
- "source": [
- "from azureml.core import Workspace\n",
- "from azureml.core.model import Model\n",
- "\n",
- "ws = Workspace.from_config()\n",
- "model=Model(ws, 'sklearn_mnist')\n",
- "model.download(target_dir='.', exist_ok=True)\n",
- "import os \n",
- "# verify the downloaded model file\n",
- "os.stat('./sklearn_mnist_model.pkl')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Test model locally\n",
- "\n",
- "Before deploying, make sure your model is working locally by:\n",
- "* Loading test data\n",
- "* Predicting test data\n",
- "* Examining the confusion matrix\n",
- "\n",
- "### Load test data\n",
- "\n",
- "Load the test data from the **./data/** directory created during the training tutorial."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from utils import load_data\n",
- "\n",
- "# note we also shrink the intensity values (X) from 0-255 to 0-1. This helps the neural network converge faster\n",
- "X_test = load_data('./data/test-images.gz', False) / 255.0\n",
- "y_test = load_data('./data/test-labels.gz', True).reshape(-1)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Predict test data\n",
- "\n",
- "Feed the test dataset to the model to get predictions."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import pickle\n",
- "from sklearn.externals import joblib\n",
- "\n",
- "clf = joblib.load('./sklearn_mnist_model.pkl')\n",
- "y_hat = clf.predict(X_test)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Examine the confusion matrix\n",
- "\n",
- "Generate a confusion matrix to see how many samples from the test set are classified correctly. Notice the mis-classified value for the incorrect predictions."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from sklearn.metrics import confusion_matrix\n",
- "\n",
- "conf_mx = confusion_matrix(y_test, y_hat)\n",
- "print(conf_mx)\n",
- "print('Overall accuracy:', np.average(y_hat == y_test))"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Use `matplotlib` to display the confusion matrix as a graph. In this graph, the X axis represents the actual values, and the Y axis represents the predicted values. The color in each grid represents the error rate. The lighter the color, the higher the error rate is. For example, many 5's are mis-classified as 3's. Hence you see a bright grid at (5,3)."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "# normalize the diagnal cells so that they don't overpower the rest of the cells when visualized\n",
- "row_sums = conf_mx.sum(axis=1, keepdims=True)\n",
- "norm_conf_mx = conf_mx / row_sums\n",
- "np.fill_diagonal(norm_conf_mx, 0)\n",
- "\n",
- "fig = plt.figure(figsize=(8,5))\n",
- "ax = fig.add_subplot(111)\n",
- "cax = ax.matshow(norm_conf_mx, cmap=plt.cm.bone)\n",
- "ticks = np.arange(0, 10, 1)\n",
- "ax.set_xticks(ticks)\n",
- "ax.set_yticks(ticks)\n",
- "ax.set_xticklabels(ticks)\n",
- "ax.set_yticklabels(ticks)\n",
- "fig.colorbar(cax)\n",
- "plt.ylabel('true labels', fontsize=14)\n",
- "plt.xlabel('predicted values', fontsize=14)\n",
- "plt.savefig('conf.png')\n",
- "plt.show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Deploy as web service\n",
- "\n",
- "Once you've tested the model and are satisfied with the results, deploy the model as a web service hosted in ACI. \n",
- "\n",
- "To build the correct environment for ACI, provide the following:\n",
- "* A scoring script to show how to use the model\n",
- "* An environment file to show what packages need to be installed\n",
- "* A configuration file to build the ACI\n",
- "* The model you trained before\n",
- "\n",
- "### Create scoring script\n",
- "\n",
- "Create the scoring script, called score.py, used by the web service call to show how to use the model.\n",
- "\n",
- "You must include two required functions into the scoring script:\n",
- "* The `init()` function, which typically loads the model into a global object. This function is run only once when the Docker container is started. \n",
- "\n",
- "* The `run(input_data)` function uses the model to predict a value based on the input data. Inputs and outputs to the run typically use JSON for serialization and de-serialization, but other formats are supported.\n"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "%%writefile score.py\n",
- "import json\n",
- "import numpy as np\n",
- "import os\n",
- "import pickle\n",
- "from sklearn.externals import joblib\n",
- "from sklearn.linear_model import LogisticRegression\n",
- "\n",
- "from azureml.core.model import Model\n",
- "\n",
- "def init():\n",
- " global model\n",
- " # retreive the path to the model file using the model name\n",
- " model_path = Model.get_model_path('sklearn_mnist')\n",
- " model = joblib.load(model_path)\n",
- "\n",
- "def run(raw_data):\n",
- " data = np.array(json.loads(raw_data)['data'])\n",
- " # make prediction\n",
- " y_hat = model.predict(data)\n",
- " # you can return any data type as long as it is JSON-serializable\n",
- " return y_hat.tolist()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create environment file\n",
- "\n",
- "Next, create an environment file, called myenv.yml, that specifies all of the script's package dependencies. This file is used to ensure that all of those dependencies are installed in the Docker image. This model needs `scikit-learn` and `azureml-sdk`."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "set conda dependencies"
- ]
- },
- "outputs": [],
- "source": [
- "from azureml.core.conda_dependencies import CondaDependencies \n",
- "\n",
- "myenv = CondaDependencies()\n",
- "myenv.add_conda_package(\"scikit-learn\")\n",
- "\n",
- "with open(\"myenv.yml\",\"w\") as f:\n",
- " f.write(myenv.serialize_to_string())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Review the content of the `myenv.yml` file."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "with open(\"myenv.yml\",\"r\") as f:\n",
- " print(f.read())"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Create configuration file\n",
- "\n",
- "Create a deployment configuration file and specify the number of CPUs and gigabyte of RAM needed for your ACI container. While it depends on your model, the default of 1 core and 1 gigabyte of RAM is usually sufficient for many models. If you feel you need more later, you would have to recreate the image and redeploy the service."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "configure web service",
- "aci"
- ]
- },
- "outputs": [],
- "source": [
- "from azureml.core.webservice import AciWebservice\n",
- "\n",
- "aciconfig = AciWebservice.deploy_configuration(cpu_cores=1, \n",
- " memory_gb=1, \n",
- " tags={\"data\": \"MNIST\", \"method\" : \"sklearn\"}, \n",
- " description='Predict MNIST with sklearn')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Deploy in ACI\n",
- "Estimated time to complete: **about 7-8 minutes**\n",
- "\n",
- "Configure the image and deploy. The following code goes through these steps:\n",
- "\n",
- "1. Build an image using:\n",
- " * The scoring file (`score.py`)\n",
- " * The environment file (`myenv.yml`)\n",
- " * The model file\n",
- "1. Register that image under the workspace. \n",
- "1. Send the image to the ACI container.\n",
- "1. Start up a container in ACI using the image.\n",
- "1. Get the web service HTTP endpoint."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "configure image",
- "create image",
- "deploy web service",
- "aci"
- ]
- },
- "outputs": [],
- "source": [
- "%%time\n",
- "from azureml.core.webservice import Webservice\n",
- "from azureml.core.image import ContainerImage\n",
- "\n",
- "# configure the image\n",
- "image_config = ContainerImage.image_configuration(execution_script=\"score.py\", \n",
- " runtime=\"python\", \n",
- " conda_file=\"myenv.yml\")\n",
- "\n",
- "service = Webservice.deploy_from_model(workspace=ws,\n",
- " name='sklearn-mnist-svc',\n",
- " deployment_config=aciconfig,\n",
- " models=[model],\n",
- " image_config=image_config)\n",
- "\n",
- "service.wait_for_deployment(show_output=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Get the scoring web service's HTTP endpoint, which accepts REST client calls. This endpoint can be shared with anyone who wants to test the web service or integrate it into an application."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "get scoring uri"
- ]
- },
- "outputs": [],
- "source": [
- "print(service.scoring_uri)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Test deployed service\n",
- "\n",
- "Earlier you scored all the test data with the local version of the model. Now, you can test the deployed model with a random sample of 30 images from the test data. \n",
- "\n",
- "The following code goes through these steps:\n",
- "1. Send the data as a JSON array to the web service hosted in ACI. \n",
- "\n",
- "1. Use the SDK's `run` API to invoke the service. You can also make raw calls using any HTTP tool such as curl.\n",
- "\n",
- "1. Print the returned predictions and plot them along with the input images. Red font and inverse image (white on black) is used to highlight the misclassified samples. \n",
- "\n",
- " Since the model accuracy is high, you might have to run the following code a few times before you can see a misclassified sample."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "score web service"
- ]
- },
- "outputs": [],
- "source": [
- "import json\n",
- "\n",
- "# find 30 random samples from test set\n",
- "n = 30\n",
- "sample_indices = np.random.permutation(X_test.shape[0])[0:n]\n",
- "\n",
- "test_samples = json.dumps({\"data\": X_test[sample_indices].tolist()})\n",
- "test_samples = bytes(test_samples, encoding='utf8')\n",
- "\n",
- "# predict using the deployed model\n",
- "result = service.run(input_data=test_samples)\n",
- "\n",
- "# compare actual value vs. the predicted values:\n",
- "i = 0\n",
- "plt.figure(figsize = (20, 1))\n",
- "\n",
- "for s in sample_indices:\n",
- " plt.subplot(1, n, i + 1)\n",
- " plt.axhline('')\n",
- " plt.axvline('')\n",
- " \n",
- " # use different color for misclassified sample\n",
- " font_color = 'red' if y_test[s] != result[i] else 'black'\n",
- " clr_map = plt.cm.gray if y_test[s] != result[i] else plt.cm.Greys\n",
- " \n",
- " plt.text(x=10, y =-10, s=result[i], fontsize=18, color=font_color)\n",
- " plt.imshow(X_test[s].reshape(28, 28), cmap=clr_map)\n",
- " \n",
- " i = i + 1\n",
- "plt.show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "You can also send raw HTTP request to test the web service."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "score web service"
- ]
- },
- "outputs": [],
- "source": [
- "import requests\n",
- "import json\n",
- "\n",
- "# send a random row from the test set to score\n",
- "random_index = np.random.randint(0, len(X_test)-1)\n",
- "input_data = \"{\\\"data\\\": [\" + str(list(X_test[random_index])) + \"]}\"\n",
- "\n",
- "headers = {'Content-Type':'application/json'}\n",
- "\n",
- "# for AKS deployment you'd need to the service key in the header as well\n",
- "# api_key = service.get_key()\n",
- "# headers = {'Content-Type':'application/json', 'Authorization':('Bearer '+ api_key)} \n",
- "\n",
- "resp = requests.post(service.scoring_uri, input_data, headers=headers)\n",
- "\n",
- "print(\"POST to url\", service.scoring_uri)\n",
- "#print(\"input data:\", input_data)\n",
- "print(\"label:\", y_test[random_index])\n",
- "print(\"prediction:\", resp.text)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Clean up resources\n",
- "\n",
- "To keep the resource group and workspace for other tutorials and exploration, you can delete only the ACI deployment using this API call:"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "delete web service"
- ]
- },
- "outputs": [],
- "source": [
- "service.delete()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "\n",
- "If you're not going to use what you've created here, delete the resources you just created with this quickstart so you don't incur any charges. In the Azure portal, select and delete your resource group. You can also keep the resource group, but delete a single workspace by displaying the workspace properties and selecting the Delete button.\n",
- "\n",
- "\n",
- "## Next steps\n",
- "\n",
- "In this Azure Machine Learning tutorial, you used Python to:\n",
- "\n",
- "> * Set up your testing environment\n",
- "> * Retrieve the model from your workspace\n",
- "> * Test the model locally\n",
- "> * Deploy the model to ACI\n",
- "> * Test the deployed model\n",
- " \n",
- "You can also try out the [regression tutorial](regression-part1-data-prep.ipynb)."
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "roastala"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Tutorial #2: Deploy an image classification model in Azure Container Instance (ACI)\n",
+ "\n",
+ "This tutorial is **part two of a two-part tutorial series**. In the [previous tutorial](img-classification-part1-training.ipynb), you trained machine learning models and then registered a model in your workspace on the cloud. \n",
+ "\n",
+ "Now, you're ready to deploy the model as a web service in [Azure Container Instances](https://docs.microsoft.com/azure/container-instances/) (ACI). A web service is an image, in this case a Docker image, that encapsulates the scoring logic and the model itself. \n",
+ "\n",
+ "In this part of the tutorial, you use Azure Machine Learning service (Preview) to:\n",
+ "\n",
+ "> * Set up your testing environment\n",
+ "> * Retrieve the model from your workspace\n",
+ "> * Test the model locally\n",
+ "> * Deploy the model to ACI\n",
+ "> * Test the deployed model\n",
+ "\n",
+ "ACI is not ideal for production deployments, but it is great for testing and understanding the workflow. For scalable production deployments, consider using AKS.\n",
+ "\n",
+ "\n",
+ "## Prerequisites\n",
+ "\n",
+ "Complete the model training in the [Tutorial #1: Train an image classification model with Azure Machine Learning](train-models.ipynb) notebook. \n",
+ "\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "register model from file"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "# If you did NOT complete the tutorial, you can instead run this cell \n",
+ "# This will register a model and download the data needed for this tutorial\n",
+ "# These prerequisites are created in the training tutorial\n",
+ "# Feel free to skip this cell if you completed the training tutorial \n",
+ "\n",
+ "# register a model\n",
+ "from azureml.core import Workspace\n",
+ "ws = Workspace.from_config()\n",
+ "\n",
+ "from azureml.core.model import Model\n",
+ "\n",
+ "model_name = \"sklearn_mnist\"\n",
+ "model = Model.register(model_path=\"sklearn_mnist_model.pkl\",\n",
+ " model_name=model_name,\n",
+ " tags={\"data\": \"mnist\", \"model\": \"classification\"},\n",
+ " description=\"Mnist handwriting recognition\",\n",
+ " workspace=ws)\n",
+ "\n",
+ "# download test data\n",
+ "import os\n",
+ "import urllib.request\n",
+ "\n",
+ "os.makedirs('./data', exist_ok=True)\n",
+ "\n",
+ "urllib.request.urlretrieve('http://yann.lecun.com/exdb/mnist/t10k-images-idx3-ubyte.gz', filename='./data/test-images.gz')\n",
+ "urllib.request.urlretrieve('http://yann.lecun.com/exdb/mnist/t10k-labels-idx1-ubyte.gz', filename='./data/test-labels.gz')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Set up the environment\n",
+ "\n",
+ "Start by setting up a testing environment.\n",
+ "\n",
+ "### Import packages\n",
+ "\n",
+ "Import the Python packages needed for this tutorial."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "check version"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "%matplotlib inline\n",
+ "import numpy as np\n",
+ "import matplotlib\n",
+ "import matplotlib.pyplot as plt\n",
+ " \n",
+ "import azureml\n",
+ "from azureml.core import Workspace, Run\n",
+ "\n",
+ "# display the core SDK version number\n",
+ "print(\"Azure ML SDK Version: \", azureml.core.VERSION)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Retrieve the model\n",
+ "\n",
+ "You registered a model in your workspace in the previous tutorial. Now, load this workspace and download the model to your local directory."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "load workspace",
+ "download model"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "from azureml.core import Workspace\n",
+ "from azureml.core.model import Model\n",
+ "\n",
+ "ws = Workspace.from_config()\n",
+ "model=Model(ws, 'sklearn_mnist')\n",
+ "model.download(target_dir='.', exist_ok=True)\n",
+ "import os \n",
+ "# verify the downloaded model file\n",
+ "os.stat('./sklearn_mnist_model.pkl')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Test model locally\n",
+ "\n",
+ "Before deploying, make sure your model is working locally by:\n",
+ "* Loading test data\n",
+ "* Predicting test data\n",
+ "* Examining the confusion matrix\n",
+ "\n",
+ "### Load test data\n",
+ "\n",
+ "Load the test data from the **./data/** directory created during the training tutorial."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from utils import load_data\n",
+ "\n",
+ "# note we also shrink the intensity values (X) from 0-255 to 0-1. This helps the neural network converge faster\n",
+ "X_test = load_data('./data/test-images.gz', False) / 255.0\n",
+ "y_test = load_data('./data/test-labels.gz', True).reshape(-1)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Predict test data\n",
+ "\n",
+ "Feed the test dataset to the model to get predictions."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import pickle\n",
+ "from sklearn.externals import joblib\n",
+ "\n",
+ "clf = joblib.load('./sklearn_mnist_model.pkl')\n",
+ "y_hat = clf.predict(X_test)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Examine the confusion matrix\n",
+ "\n",
+ "Generate a confusion matrix to see how many samples from the test set are classified correctly. Notice the mis-classified value for the incorrect predictions."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from sklearn.metrics import confusion_matrix\n",
+ "\n",
+ "conf_mx = confusion_matrix(y_test, y_hat)\n",
+ "print(conf_mx)\n",
+ "print('Overall accuracy:', np.average(y_hat == y_test))"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Use `matplotlib` to display the confusion matrix as a graph. In this graph, the X axis represents the actual values, and the Y axis represents the predicted values. The color in each grid represents the error rate. The lighter the color, the higher the error rate is. For example, many 5's are mis-classified as 3's. Hence you see a bright grid at (5,3)."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "# normalize the diagnal cells so that they don't overpower the rest of the cells when visualized\n",
+ "row_sums = conf_mx.sum(axis=1, keepdims=True)\n",
+ "norm_conf_mx = conf_mx / row_sums\n",
+ "np.fill_diagonal(norm_conf_mx, 0)\n",
+ "\n",
+ "fig = plt.figure(figsize=(8,5))\n",
+ "ax = fig.add_subplot(111)\n",
+ "cax = ax.matshow(norm_conf_mx, cmap=plt.cm.bone)\n",
+ "ticks = np.arange(0, 10, 1)\n",
+ "ax.set_xticks(ticks)\n",
+ "ax.set_yticks(ticks)\n",
+ "ax.set_xticklabels(ticks)\n",
+ "ax.set_yticklabels(ticks)\n",
+ "fig.colorbar(cax)\n",
+ "plt.ylabel('true labels', fontsize=14)\n",
+ "plt.xlabel('predicted values', fontsize=14)\n",
+ "plt.savefig('conf.png')\n",
+ "plt.show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Deploy as web service\n",
+ "\n",
+ "Once you've tested the model and are satisfied with the results, deploy the model as a web service hosted in ACI. \n",
+ "\n",
+ "To build the correct environment for ACI, provide the following:\n",
+ "* A scoring script to show how to use the model\n",
+ "* An environment file to show what packages need to be installed\n",
+ "* A configuration file to build the ACI\n",
+ "* The model you trained before\n",
+ "\n",
+ "### Create scoring script\n",
+ "\n",
+ "Create the scoring script, called score.py, used by the web service call to show how to use the model.\n",
+ "\n",
+ "You must include two required functions into the scoring script:\n",
+ "* The `init()` function, which typically loads the model into a global object. This function is run only once when the Docker container is started. \n",
+ "\n",
+ "* The `run(input_data)` function uses the model to predict a value based on the input data. Inputs and outputs to the run typically use JSON for serialization and de-serialization, but other formats are supported.\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "%%writefile score.py\n",
+ "import json\n",
+ "import numpy as np\n",
+ "import os\n",
+ "import pickle\n",
+ "from sklearn.externals import joblib\n",
+ "from sklearn.linear_model import LogisticRegression\n",
+ "\n",
+ "from azureml.core.model import Model\n",
+ "\n",
+ "def init():\n",
+ " global model\n",
+ " # retreive the path to the model file using the model name\n",
+ " model_path = Model.get_model_path('sklearn_mnist')\n",
+ " model = joblib.load(model_path)\n",
+ "\n",
+ "def run(raw_data):\n",
+ " data = np.array(json.loads(raw_data)['data'])\n",
+ " # make prediction\n",
+ " y_hat = model.predict(data)\n",
+ " # you can return any data type as long as it is JSON-serializable\n",
+ " return y_hat.tolist()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create environment file\n",
+ "\n",
+ "Next, create an environment file, called myenv.yml, that specifies all of the script's package dependencies. This file is used to ensure that all of those dependencies are installed in the Docker image. This model needs `scikit-learn` and `azureml-sdk`."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "set conda dependencies"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "from azureml.core.conda_dependencies import CondaDependencies \n",
+ "\n",
+ "myenv = CondaDependencies()\n",
+ "myenv.add_conda_package(\"scikit-learn\")\n",
+ "\n",
+ "with open(\"myenv.yml\",\"w\") as f:\n",
+ " f.write(myenv.serialize_to_string())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Review the content of the `myenv.yml` file."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "with open(\"myenv.yml\",\"r\") as f:\n",
+ " print(f.read())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Create configuration file\n",
+ "\n",
+ "Create a deployment configuration file and specify the number of CPUs and gigabyte of RAM needed for your ACI container. While it depends on your model, the default of 1 core and 1 gigabyte of RAM is usually sufficient for many models. If you feel you need more later, you would have to recreate the image and redeploy the service."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "configure web service",
+ "aci"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "from azureml.core.webservice import AciWebservice\n",
+ "\n",
+ "aciconfig = AciWebservice.deploy_configuration(cpu_cores=1, \n",
+ " memory_gb=1, \n",
+ " tags={\"data\": \"MNIST\", \"method\" : \"sklearn\"}, \n",
+ " description='Predict MNIST with sklearn')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Deploy in ACI\n",
+ "Estimated time to complete: **about 7-8 minutes**\n",
+ "\n",
+ "Configure the image and deploy. The following code goes through these steps:\n",
+ "\n",
+ "1. Build an image using:\n",
+ " * The scoring file (`score.py`)\n",
+ " * The environment file (`myenv.yml`)\n",
+ " * The model file\n",
+ "1. Register that image under the workspace. \n",
+ "1. Send the image to the ACI container.\n",
+ "1. Start up a container in ACI using the image.\n",
+ "1. Get the web service HTTP endpoint."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "configure image",
+ "create image",
+ "deploy web service",
+ "aci"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "%%time\n",
+ "from azureml.core.webservice import Webservice\n",
+ "from azureml.core.image import ContainerImage\n",
+ "\n",
+ "# configure the image\n",
+ "image_config = ContainerImage.image_configuration(execution_script=\"score.py\", \n",
+ " runtime=\"python\", \n",
+ " conda_file=\"myenv.yml\")\n",
+ "\n",
+ "service = Webservice.deploy_from_model(workspace=ws,\n",
+ " name='sklearn-mnist-svc',\n",
+ " deployment_config=aciconfig,\n",
+ " models=[model],\n",
+ " image_config=image_config)\n",
+ "\n",
+ "service.wait_for_deployment(show_output=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Get the scoring web service's HTTP endpoint, which accepts REST client calls. This endpoint can be shared with anyone who wants to test the web service or integrate it into an application."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "get scoring uri"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "print(service.scoring_uri)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Test deployed service\n",
+ "\n",
+ "Earlier you scored all the test data with the local version of the model. Now, you can test the deployed model with a random sample of 30 images from the test data. \n",
+ "\n",
+ "The following code goes through these steps:\n",
+ "1. Send the data as a JSON array to the web service hosted in ACI. \n",
+ "\n",
+ "1. Use the SDK's `run` API to invoke the service. You can also make raw calls using any HTTP tool such as curl.\n",
+ "\n",
+ "1. Print the returned predictions and plot them along with the input images. Red font and inverse image (white on black) is used to highlight the misclassified samples. \n",
+ "\n",
+ " Since the model accuracy is high, you might have to run the following code a few times before you can see a misclassified sample."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "score web service"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "import json\n",
+ "\n",
+ "# find 30 random samples from test set\n",
+ "n = 30\n",
+ "sample_indices = np.random.permutation(X_test.shape[0])[0:n]\n",
+ "\n",
+ "test_samples = json.dumps({\"data\": X_test[sample_indices].tolist()})\n",
+ "test_samples = bytes(test_samples, encoding='utf8')\n",
+ "\n",
+ "# predict using the deployed model\n",
+ "result = service.run(input_data=test_samples)\n",
+ "\n",
+ "# compare actual value vs. the predicted values:\n",
+ "i = 0\n",
+ "plt.figure(figsize = (20, 1))\n",
+ "\n",
+ "for s in sample_indices:\n",
+ " plt.subplot(1, n, i + 1)\n",
+ " plt.axhline('')\n",
+ " plt.axvline('')\n",
+ " \n",
+ " # use different color for misclassified sample\n",
+ " font_color = 'red' if y_test[s] != result[i] else 'black'\n",
+ " clr_map = plt.cm.gray if y_test[s] != result[i] else plt.cm.Greys\n",
+ " \n",
+ " plt.text(x=10, y =-10, s=result[i], fontsize=18, color=font_color)\n",
+ " plt.imshow(X_test[s].reshape(28, 28), cmap=clr_map)\n",
+ " \n",
+ " i = i + 1\n",
+ "plt.show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "You can also send raw HTTP request to test the web service."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "score web service"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "import requests\n",
+ "import json\n",
+ "\n",
+ "# send a random row from the test set to score\n",
+ "random_index = np.random.randint(0, len(X_test)-1)\n",
+ "input_data = \"{\\\"data\\\": [\" + str(list(X_test[random_index])) + \"]}\"\n",
+ "\n",
+ "headers = {'Content-Type':'application/json'}\n",
+ "\n",
+ "# for AKS deployment you'd need to the service key in the header as well\n",
+ "# api_key = service.get_key()\n",
+ "# headers = {'Content-Type':'application/json', 'Authorization':('Bearer '+ api_key)} \n",
+ "\n",
+ "resp = requests.post(service.scoring_uri, input_data, headers=headers)\n",
+ "\n",
+ "print(\"POST to url\", service.scoring_uri)\n",
+ "#print(\"input data:\", input_data)\n",
+ "print(\"label:\", y_test[random_index])\n",
+ "print(\"prediction:\", resp.text)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Clean up resources\n",
+ "\n",
+ "To keep the resource group and workspace for other tutorials and exploration, you can delete only the ACI deployment using this API call:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "delete web service"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "service.delete()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "\n",
+ "If you're not going to use what you've created here, delete the resources you just created with this quickstart so you don't incur any charges. In the Azure portal, select and delete your resource group. You can also keep the resource group, but delete a single workspace by displaying the workspace properties and selecting the Delete button.\n",
+ "\n",
+ "\n",
+ "## Next steps\n",
+ "\n",
+ "In this Azure Machine Learning tutorial, you used Python to:\n",
+ "\n",
+ "> * Set up your testing environment\n",
+ "> * Retrieve the model from your workspace\n",
+ "> * Test the model locally\n",
+ "> * Deploy the model to ACI\n",
+ "> * Test the deployed model\n",
+ " \n",
+ "You can also try out the [Automatic algorithm selection tutorial](03.auto-train-models.ipynb) to see how Azure Machine Learning can auto-select and tune the best algorithm for your model and build that model for you."
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "roastala"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.6"
+ },
+ "msauthor": "sgilley"
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.6"
- },
- "msauthor": "sgilley"
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/tutorials/regression-part1-data-prep.ipynb b/tutorials/regression-part1-data-prep.ipynb
index 402bfdc7..b54f70d5 100644
--- a/tutorials/regression-part1-data-prep.ipynb
+++ b/tutorials/regression-part1-data-prep.ipynb
@@ -1,631 +1,631 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved.\n",
- "\n",
- "Licensed under the MIT License."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Tutorial (part 1): Prepare data for regression modeling"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "In this tutorial, you learn how to prep data for regression modeling using the Azure Machine Learning Data Prep SDK. Perform various transformations to filter and combine two different NYC Taxi data sets. The end goal of this tutorial set is to predict the cost of a taxi trip by training a model on data features including pickup hour, day of week, number of passengers, and coordinates. This tutorial is part one of a two-part tutorial series.\n",
- "\n",
- "In this tutorial, you:\n",
- "\n",
- "\n",
- "> * Setup a Python environment and import packages\n",
- "> * Load two datasets with different field names\n",
- "> * Cleanse data to remove anomalies\n",
- "> * Transform data using intelligent transforms to create new features\n",
- "> * Save your dataflow object to use in a regression model\n",
- "\n",
- "You can prepare your data in Python using the [Azure Machine Learning Data Prep SDK](https://aka.ms/data-prep-sdk)."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Import packages\n",
- "Begin by importing the SDK."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import azureml.dataprep as dprep"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Load data\n",
- "Download two different NYC Taxi data sets into dataflow objects. These datasets contain slightly different fields. The method `auto_read_file()` automatically recognizes the input file type."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "dataset_root = \"https://dprepdata.blob.core.windows.net/demo\"\n",
- "\n",
- "green_path = \"/\".join([dataset_root, \"green-small/*\"])\n",
- "yellow_path = \"/\".join([dataset_root, \"yellow-small/*\"])\n",
- "\n",
- "green_df = dprep.read_csv(path=green_path, header=dprep.PromoteHeadersMode.GROUPED)\n",
- "# auto_read_file will automatically identify and parse the file type, and is useful if you don't know the file type\n",
- "yellow_df = dprep.auto_read_file(path=yellow_path)\n",
- "\n",
- "green_df.head(5)\n",
- "yellow_df.head(5)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Cleanse data"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Now you populate some variables with shortcut transforms that will apply to all dataflows. The variable `drop_if_all_null` will be used to delete records where all fields are null. The variable `useful_columns` holds an array of column descriptions that are retained in each dataflow."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "all_columns = dprep.ColumnSelector(term=\".*\", use_regex=True)\n",
- "drop_if_all_null = [all_columns, dprep.ColumnRelationship(dprep.ColumnRelationship.ALL)]\n",
- "useful_columns = [\n",
- " \"cost\", \"distance\", \"dropoff_datetime\", \"dropoff_latitude\", \"dropoff_longitude\",\n",
- " \"passengers\", \"pickup_datetime\", \"pickup_latitude\", \"pickup_longitude\", \"store_forward\", \"vendor\"\n",
- "]"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "You first work with the green taxi data and get it into a valid shape that can be combined with the yellow taxi data. Create a temporary dataflow `tmp_df`, and call the `replace_na()`, `drop_nulls()`, and `keep_columns()` functions using the shortcut transform variables you created. Additionally, rename all the columns in the dataframe to match the names in `useful_columns`."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "tmp_df = (green_df\n",
- " .replace_na(columns=all_columns)\n",
- " .drop_nulls(*drop_if_all_null)\n",
- " .rename_columns(column_pairs={\n",
- " \"VendorID\": \"vendor\",\n",
- " \"lpep_pickup_datetime\": \"pickup_datetime\",\n",
- " \"Lpep_dropoff_datetime\": \"dropoff_datetime\",\n",
- " \"lpep_dropoff_datetime\": \"dropoff_datetime\",\n",
- " \"Store_and_fwd_flag\": \"store_forward\",\n",
- " \"store_and_fwd_flag\": \"store_forward\",\n",
- " \"Pickup_longitude\": \"pickup_longitude\",\n",
- " \"Pickup_latitude\": \"pickup_latitude\",\n",
- " \"Dropoff_longitude\": \"dropoff_longitude\",\n",
- " \"Dropoff_latitude\": \"dropoff_latitude\",\n",
- " \"Passenger_count\": \"passengers\",\n",
- " \"Fare_amount\": \"cost\",\n",
- " \"Trip_distance\": \"distance\"\n",
- " })\n",
- " .keep_columns(columns=useful_columns))\n",
- "tmp_df.head(5)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Overwrite the `green_df` variable with the transforms performed on `tmp_df` in the previous step."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "green_df = tmp_df"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Perform the same transformation steps to the yellow taxi data."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "tmp_df = (yellow_df\n",
- " .replace_na(columns=all_columns)\n",
- " .drop_nulls(*drop_if_all_null)\n",
- " .rename_columns(column_pairs={\n",
- " \"vendor_name\": \"vendor\",\n",
- " \"VendorID\": \"vendor\",\n",
- " \"vendor_id\": \"vendor\",\n",
- " \"Trip_Pickup_DateTime\": \"pickup_datetime\",\n",
- " \"tpep_pickup_datetime\": \"pickup_datetime\",\n",
- " \"Trip_Dropoff_DateTime\": \"dropoff_datetime\",\n",
- " \"tpep_dropoff_datetime\": \"dropoff_datetime\",\n",
- " \"store_and_forward\": \"store_forward\",\n",
- " \"store_and_fwd_flag\": \"store_forward\",\n",
- " \"Start_Lon\": \"pickup_longitude\",\n",
- " \"Start_Lat\": \"pickup_latitude\",\n",
- " \"End_Lon\": \"dropoff_longitude\",\n",
- " \"End_Lat\": \"dropoff_latitude\",\n",
- " \"Passenger_Count\": \"passengers\",\n",
- " \"passenger_count\": \"passengers\",\n",
- " \"Fare_Amt\": \"cost\",\n",
- " \"fare_amount\": \"cost\",\n",
- " \"Trip_Distance\": \"distance\",\n",
- " \"trip_distance\": \"distance\"\n",
- " })\n",
- " .keep_columns(columns=useful_columns))\n",
- "tmp_df.head(5)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Again, overwrite `yellow_df` with `tmp_df`, and then call the `append_rows()` function on the green taxi data to append the yellow taxi data, creating a new combined dataframe."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "yellow_df = tmp_df\n",
- "combined_df = green_df.append_rows([yellow_df])"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Convert types and filter "
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Examine the pickup and drop-off coordinates summary statistics to see how the data is distributed. First define a `TypeConverter` object to change the lat/long fields to decimal type. Next, call the `keep_columns()` function to restrict output to only the lat/long fields, and then call `get_profile()`."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "decimal_type = dprep.TypeConverter(data_type=dprep.FieldType.DECIMAL)\n",
- "combined_df = combined_df.set_column_types(type_conversions={\n",
- " \"pickup_longitude\": decimal_type,\n",
- " \"pickup_latitude\": decimal_type,\n",
- " \"dropoff_longitude\": decimal_type,\n",
- " \"dropoff_latitude\": decimal_type\n",
- "})\n",
- "combined_df.keep_columns(columns=[\n",
- " \"pickup_longitude\", \"pickup_latitude\", \n",
- " \"dropoff_longitude\", \"dropoff_latitude\"\n",
- "]).get_profile()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "From the summary statistics output, you see that there are coordinates that are missing, and coordinates that are not in New York City. Filter out coordinates not in the city border by chaining column filter commands within the `filter()` function, and defining minimum and maximum bounds for each field. Then call `get_profile()` again to verify the transformation."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "tmp_df = (combined_df\n",
- " .drop_nulls(\n",
- " columns=[\"pickup_longitude\", \"pickup_latitude\", \"dropoff_longitude\", \"dropoff_latitude\"],\n",
- " column_relationship=dprep.ColumnRelationship(dprep.ColumnRelationship.ANY)\n",
- " ) \n",
- " .filter(dprep.f_and(\n",
- " dprep.col(\"pickup_longitude\") <= -73.72,\n",
- " dprep.col(\"pickup_longitude\") >= -74.09,\n",
- " dprep.col(\"pickup_latitude\") <= 40.88,\n",
- " dprep.col(\"pickup_latitude\") >= 40.53,\n",
- " dprep.col(\"dropoff_longitude\") <= -73.72,\n",
- " dprep.col(\"dropoff_longitude\") >= -74.09,\n",
- " dprep.col(\"dropoff_latitude\") <= 40.88,\n",
- " dprep.col(\"dropoff_latitude\") >= 40.53\n",
- " )))\n",
- "tmp_df.keep_columns(columns=[\n",
- " \"pickup_longitude\", \"pickup_latitude\", \n",
- " \"dropoff_longitude\", \"dropoff_latitude\"\n",
- "]).get_profile()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Overwrite `combined_df` with the transformations you made to `tmp_df`."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "combined_df = tmp_df"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Split and rename columns"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Look at the data profile for the `store_forward` column."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "combined_df.keep_columns(columns='store_forward').get_profile()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "From the data profile output of `store_forward`, you see that the data is inconsistent and there are missing/null values. Replace these values using the `replace()` and `fill_nulls()` functions, and in both cases change to the string \"N\"."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "combined_df = combined_df.replace(columns=\"store_forward\", find=\"0\", replace_with=\"N\").fill_nulls(\"store_forward\", \"N\")"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Execute another `replace` function, this time on the `distance` field. This reformats distance values that are incorrectly labeled as `.00`, and fills any nulls with zeros. Convert the `distance` field to numerical format."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "combined_df = combined_df.replace(columns=\"distance\", find=\".00\", replace_with=0).fill_nulls(\"distance\", 0)\n",
- "combined_df = combined_df.to_number([\"distance\"])"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Split the pick up and drop off datetimes into respective date and time columns. Use `split_column_by_example()` to perform the split. In this case, the optional `example` parameter of `split_column_by_example()` is omitted. Therefore the function will automatically determine where to split based on the data."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "tmp_df = (combined_df\n",
- " .split_column_by_example(source_column=\"pickup_datetime\")\n",
- " .split_column_by_example(source_column=\"dropoff_datetime\"))\n",
- "tmp_df.head(5)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Rename the columns generated by `split_column_by_example()` into meaningful names."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "tmp_df_renamed = (tmp_df\n",
- " .rename_columns(column_pairs={\n",
- " \"pickup_datetime_1\": \"pickup_date\",\n",
- " \"pickup_datetime_2\": \"pickup_time\",\n",
- " \"dropoff_datetime_1\": \"dropoff_date\",\n",
- " \"dropoff_datetime_2\": \"dropoff_time\"\n",
- " }))\n",
- "tmp_df_renamed.head(5)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Overwrite `combined_df` with the executed transformations, and then call `get_profile()` to see full summary statistics after all transformations."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "combined_df = tmp_df_renamed\n",
- "combined_df.get_profile()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Transform data"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Split the pickup and drop-off date further into day of week, day of month, and month. To get day of week, use the `derive_column_by_example()` function. This function takes as a parameter an array of example objects that define the input data, and the desired output. The function then automatically determines your desired transformation. For pickup and drop-off time columns, split into hour, minute, and second using the `split_column_by_example()` function with no example parameter.\n",
- "\n",
- "Once you have generated these new features, delete the original fields in favor of the newly generated features using `drop_columns()`. Rename all remaining fields to accurate descriptions."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "tmp_df = (combined_df\n",
- " .derive_column_by_example(\n",
- " source_columns=\"pickup_date\", \n",
- " new_column_name=\"pickup_weekday\", \n",
- " example_data=[(\"2009-01-04\", \"Sunday\"), (\"2013-08-22\", \"Thursday\")]\n",
- " )\n",
- " .derive_column_by_example(\n",
- " source_columns=\"dropoff_date\",\n",
- " new_column_name=\"dropoff_weekday\",\n",
- " example_data=[(\"2013-08-22\", \"Thursday\"), (\"2013-11-03\", \"Sunday\")]\n",
- " )\n",
- " \n",
- " .split_column_by_example(source_column=\"pickup_time\")\n",
- " .split_column_by_example(source_column=\"dropoff_time\")\n",
- " # the following two split_column_by_example calls reference the generated column names from the above two calls\n",
- " .split_column_by_example(source_column=\"pickup_time_1\")\n",
- " .split_column_by_example(source_column=\"dropoff_time_1\")\n",
- " .drop_columns(columns=[\n",
- " \"pickup_date\", \"pickup_time\", \"dropoff_date\", \"dropoff_time\", \n",
- " \"pickup_date_1\", \"dropoff_date_1\", \"pickup_time_1\", \"dropoff_time_1\"\n",
- " ])\n",
- " \n",
- " .rename_columns(column_pairs={\n",
- " \"pickup_date_2\": \"pickup_month\",\n",
- " \"pickup_date_3\": \"pickup_monthday\",\n",
- " \"pickup_time_1_1\": \"pickup_hour\",\n",
- " \"pickup_time_1_2\": \"pickup_minute\",\n",
- " \"pickup_time_2\": \"pickup_second\",\n",
- " \"dropoff_date_2\": \"dropoff_month\",\n",
- " \"dropoff_date_3\": \"dropoff_monthday\",\n",
- " \"dropoff_time_1_1\": \"dropoff_hour\",\n",
- " \"dropoff_time_1_2\": \"dropoff_minute\",\n",
- " \"dropoff_time_2\": \"dropoff_second\"\n",
- " }))\n",
- "\n",
- "tmp_df.head(5)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "From the data above, you see that the pickup and drop-off date and time components produced from the derived transformations are correct. Drop the `pickup_datetime` and `dropoff_datetime` columns as they are no longer needed."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "tmp_df = tmp_df.drop_columns(columns=[\"pickup_datetime\", \"dropoff_datetime\"])"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Use the type inference functionality to automatically check the data type of each field, and display the inference results."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "type_infer = tmp_df.builders.set_column_types()\n",
- "type_infer.learn()\n",
- "type_infer"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "The inference results look correct based on the data, now apply the type conversions to the dataflow."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "tmp_df = type_infer.to_dataflow()\n",
- "tmp_df.get_profile()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Before packaging the dataflow, perform two final filters on the data set. To eliminate incorrect data points, filter the dataflow on records where both the `cost` and `distance` are greater than zero."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "tmp_df = tmp_df.filter(dprep.col(\"distance\") > 0)\n",
- "tmp_df = tmp_df.filter(dprep.col(\"cost\") > 0)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "At this point, you have a fully transformed and prepared dataflow object to use in a machine learning model. The DataPrep SDK includes object serialization functionality, which is used as follows."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import os\n",
- "file_path = os.path.join(os.getcwd(), \"dflows.dprep\")\n",
- "\n",
- "dflow_prepared = tmp_df\n",
- "package = dprep.Package([dflow_prepared])\n",
- "package.save(file_path)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Clean up resources"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Delete the file `dflows.dprep` (whether you are running locally or in Azure Notebooks) in your current directory if you do not wish to continue with part two of the tutorial. If you continue on to part two, you will need the `dflows.dprep` file in the current directory."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Next steps"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "In this Azure Machine Learning Data Prep SDK tutorial, you:\n",
- "\n",
- "> * Set up your development environment\n",
- "> * Loaded and cleansed data sets\n",
- "> * Used smart transforms to predict your logic based on an example\n",
- "> * Merged and packaged datasets for machine learning training\n",
- "\n",
- "You are ready to use this training data in the next part of the tutorial series:\n",
- "\n",
- "\n",
- "> [Tutorial #2: Train regression model](regression-part2-automated-ml.ipynb)"
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "cforbe"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved.\n",
+ "\n",
+ "Licensed under the MIT License."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Tutorial (part 1): Prepare data for regression modeling"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "In this tutorial, you learn how to prep data for regression modeling using the Azure Machine Learning Data Prep SDK. Perform various transformations to filter and combine two different NYC Taxi data sets. The end goal of this tutorial set is to predict the cost of a taxi trip by training a model on data features including pickup hour, day of week, number of passengers, and coordinates. This tutorial is part one of a two-part tutorial series.\n",
+ "\n",
+ "In this tutorial, you:\n",
+ "\n",
+ "\n",
+ "> * Setup a Python environment and import packages\n",
+ "> * Load two datasets with different field names\n",
+ "> * Cleanse data to remove anomalies\n",
+ "> * Transform data using intelligent transforms to create new features\n",
+ "> * Save your dataflow object to use in a regression model\n",
+ "\n",
+ "You can prepare your data in Python using the [Azure Machine Learning Data Prep SDK](https://aka.ms/data-prep-sdk)."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Import packages\n",
+ "Begin by importing the SDK."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import azureml.dataprep as dprep"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Load data\n",
+ "Download two different NYC Taxi data sets into dataflow objects. These datasets contain slightly different fields. The method `auto_read_file()` automatically recognizes the input file type."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "dataset_root = \"https://dprepdata.blob.core.windows.net/demo\"\n",
+ "\n",
+ "green_path = \"/\".join([dataset_root, \"green-small/*\"])\n",
+ "yellow_path = \"/\".join([dataset_root, \"yellow-small/*\"])\n",
+ "\n",
+ "green_df = dprep.read_csv(path=green_path, header=dprep.PromoteHeadersMode.GROUPED)\n",
+ "# auto_read_file will automatically identify and parse the file type, and is useful if you don't know the file type\n",
+ "yellow_df = dprep.auto_read_file(path=yellow_path)\n",
+ "\n",
+ "green_df.head(5)\n",
+ "yellow_df.head(5)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Cleanse data"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Now you populate some variables with shortcut transforms that will apply to all dataflows. The variable `drop_if_all_null` will be used to delete records where all fields are null. The variable `useful_columns` holds an array of column descriptions that are retained in each dataflow."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "all_columns = dprep.ColumnSelector(term=\".*\", use_regex=True)\n",
+ "drop_if_all_null = [all_columns, dprep.ColumnRelationship(dprep.ColumnRelationship.ALL)]\n",
+ "useful_columns = [\n",
+ " \"cost\", \"distance\", \"dropoff_datetime\", \"dropoff_latitude\", \"dropoff_longitude\",\n",
+ " \"passengers\", \"pickup_datetime\", \"pickup_latitude\", \"pickup_longitude\", \"store_forward\", \"vendor\"\n",
+ "]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "You first work with the green taxi data and get it into a valid shape that can be combined with the yellow taxi data. Create a temporary dataflow `tmp_df`, and call the `replace_na()`, `drop_nulls()`, and `keep_columns()` functions using the shortcut transform variables you created. Additionally, rename all the columns in the dataframe to match the names in `useful_columns`."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "tmp_df = (green_df\n",
+ " .replace_na(columns=all_columns)\n",
+ " .drop_nulls(*drop_if_all_null)\n",
+ " .rename_columns(column_pairs={\n",
+ " \"VendorID\": \"vendor\",\n",
+ " \"lpep_pickup_datetime\": \"pickup_datetime\",\n",
+ " \"Lpep_dropoff_datetime\": \"dropoff_datetime\",\n",
+ " \"lpep_dropoff_datetime\": \"dropoff_datetime\",\n",
+ " \"Store_and_fwd_flag\": \"store_forward\",\n",
+ " \"store_and_fwd_flag\": \"store_forward\",\n",
+ " \"Pickup_longitude\": \"pickup_longitude\",\n",
+ " \"Pickup_latitude\": \"pickup_latitude\",\n",
+ " \"Dropoff_longitude\": \"dropoff_longitude\",\n",
+ " \"Dropoff_latitude\": \"dropoff_latitude\",\n",
+ " \"Passenger_count\": \"passengers\",\n",
+ " \"Fare_amount\": \"cost\",\n",
+ " \"Trip_distance\": \"distance\"\n",
+ " })\n",
+ " .keep_columns(columns=useful_columns))\n",
+ "tmp_df.head(5)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Overwrite the `green_df` variable with the transforms performed on `tmp_df` in the previous step."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "green_df = tmp_df"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Perform the same transformation steps to the yellow taxi data."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "tmp_df = (yellow_df\n",
+ " .replace_na(columns=all_columns)\n",
+ " .drop_nulls(*drop_if_all_null)\n",
+ " .rename_columns(column_pairs={\n",
+ " \"vendor_name\": \"vendor\",\n",
+ " \"VendorID\": \"vendor\",\n",
+ " \"vendor_id\": \"vendor\",\n",
+ " \"Trip_Pickup_DateTime\": \"pickup_datetime\",\n",
+ " \"tpep_pickup_datetime\": \"pickup_datetime\",\n",
+ " \"Trip_Dropoff_DateTime\": \"dropoff_datetime\",\n",
+ " \"tpep_dropoff_datetime\": \"dropoff_datetime\",\n",
+ " \"store_and_forward\": \"store_forward\",\n",
+ " \"store_and_fwd_flag\": \"store_forward\",\n",
+ " \"Start_Lon\": \"pickup_longitude\",\n",
+ " \"Start_Lat\": \"pickup_latitude\",\n",
+ " \"End_Lon\": \"dropoff_longitude\",\n",
+ " \"End_Lat\": \"dropoff_latitude\",\n",
+ " \"Passenger_Count\": \"passengers\",\n",
+ " \"passenger_count\": \"passengers\",\n",
+ " \"Fare_Amt\": \"cost\",\n",
+ " \"fare_amount\": \"cost\",\n",
+ " \"Trip_Distance\": \"distance\",\n",
+ " \"trip_distance\": \"distance\"\n",
+ " })\n",
+ " .keep_columns(columns=useful_columns))\n",
+ "tmp_df.head(5)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Again, overwrite `yellow_df` with `tmp_df`, and then call the `append_rows()` function on the green taxi data to append the yellow taxi data, creating a new combined dataframe."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "yellow_df = tmp_df\n",
+ "combined_df = green_df.append_rows([yellow_df])"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Convert types and filter "
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Examine the pickup and drop-off coordinates summary statistics to see how the data is distributed. First define a `TypeConverter` object to change the lat/long fields to decimal type. Next, call the `keep_columns()` function to restrict output to only the lat/long fields, and then call `get_profile()`."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "decimal_type = dprep.TypeConverter(data_type=dprep.FieldType.DECIMAL)\n",
+ "combined_df = combined_df.set_column_types(type_conversions={\n",
+ " \"pickup_longitude\": decimal_type,\n",
+ " \"pickup_latitude\": decimal_type,\n",
+ " \"dropoff_longitude\": decimal_type,\n",
+ " \"dropoff_latitude\": decimal_type\n",
+ "})\n",
+ "combined_df.keep_columns(columns=[\n",
+ " \"pickup_longitude\", \"pickup_latitude\", \n",
+ " \"dropoff_longitude\", \"dropoff_latitude\"\n",
+ "]).get_profile()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "From the summary statistics output, you see that there are coordinates that are missing, and coordinates that are not in New York City. Filter out coordinates not in the city border by chaining column filter commands within the `filter()` function, and defining minimum and maximum bounds for each field. Then call `get_profile()` again to verify the transformation."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "tmp_df = (combined_df\n",
+ " .drop_nulls(\n",
+ " columns=[\"pickup_longitude\", \"pickup_latitude\", \"dropoff_longitude\", \"dropoff_latitude\"],\n",
+ " column_relationship=dprep.ColumnRelationship(dprep.ColumnRelationship.ANY)\n",
+ " ) \n",
+ " .filter(dprep.f_and(\n",
+ " dprep.col(\"pickup_longitude\") <= -73.72,\n",
+ " dprep.col(\"pickup_longitude\") >= -74.09,\n",
+ " dprep.col(\"pickup_latitude\") <= 40.88,\n",
+ " dprep.col(\"pickup_latitude\") >= 40.53,\n",
+ " dprep.col(\"dropoff_longitude\") <= -73.72,\n",
+ " dprep.col(\"dropoff_longitude\") >= -74.09,\n",
+ " dprep.col(\"dropoff_latitude\") <= 40.88,\n",
+ " dprep.col(\"dropoff_latitude\") >= 40.53\n",
+ " )))\n",
+ "tmp_df.keep_columns(columns=[\n",
+ " \"pickup_longitude\", \"pickup_latitude\", \n",
+ " \"dropoff_longitude\", \"dropoff_latitude\"\n",
+ "]).get_profile()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Overwrite `combined_df` with the transformations you made to `tmp_df`."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "combined_df = tmp_df"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Split and rename columns"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Look at the data profile for the `store_forward` column."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "combined_df.keep_columns(columns='store_forward').get_profile()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "From the data profile output of `store_forward`, you see that the data is inconsistent and there are missing/null values. Replace these values using the `replace()` and `fill_nulls()` functions, and in both cases change to the string \"N\"."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "combined_df = combined_df.replace(columns=\"store_forward\", find=\"0\", replace_with=\"N\").fill_nulls(\"store_forward\", \"N\")"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Execute another `replace` function, this time on the `distance` field. This reformats distance values that are incorrectly labeled as `.00`, and fills any nulls with zeros. Convert the `distance` field to numerical format."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "combined_df = combined_df.replace(columns=\"distance\", find=\".00\", replace_with=0).fill_nulls(\"distance\", 0)\n",
+ "combined_df = combined_df.to_number([\"distance\"])"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Split the pick up and drop off datetimes into respective date and time columns. Use `split_column_by_example()` to perform the split. In this case, the optional `example` parameter of `split_column_by_example()` is omitted. Therefore the function will automatically determine where to split based on the data."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "tmp_df = (combined_df\n",
+ " .split_column_by_example(source_column=\"pickup_datetime\")\n",
+ " .split_column_by_example(source_column=\"dropoff_datetime\"))\n",
+ "tmp_df.head(5)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Rename the columns generated by `split_column_by_example()` into meaningful names."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "tmp_df_renamed = (tmp_df\n",
+ " .rename_columns(column_pairs={\n",
+ " \"pickup_datetime_1\": \"pickup_date\",\n",
+ " \"pickup_datetime_2\": \"pickup_time\",\n",
+ " \"dropoff_datetime_1\": \"dropoff_date\",\n",
+ " \"dropoff_datetime_2\": \"dropoff_time\"\n",
+ " }))\n",
+ "tmp_df_renamed.head(5)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Overwrite `combined_df` with the executed transformations, and then call `get_profile()` to see full summary statistics after all transformations."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "combined_df = tmp_df_renamed\n",
+ "combined_df.get_profile()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Transform data"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Split the pickup and drop-off date further into day of week, day of month, and month. To get day of week, use the `derive_column_by_example()` function. This function takes as a parameter an array of example objects that define the input data, and the desired output. The function then automatically determines your desired transformation. For pickup and drop-off time columns, split into hour, minute, and second using the `split_column_by_example()` function with no example parameter.\n",
+ "\n",
+ "Once you have generated these new features, delete the original fields in favor of the newly generated features using `drop_columns()`. Rename all remaining fields to accurate descriptions."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "tmp_df = (combined_df\n",
+ " .derive_column_by_example(\n",
+ " source_columns=\"pickup_date\", \n",
+ " new_column_name=\"pickup_weekday\", \n",
+ " example_data=[(\"2009-01-04\", \"Sunday\"), (\"2013-08-22\", \"Thursday\")]\n",
+ " )\n",
+ " .derive_column_by_example(\n",
+ " source_columns=\"dropoff_date\",\n",
+ " new_column_name=\"dropoff_weekday\",\n",
+ " example_data=[(\"2013-08-22\", \"Thursday\"), (\"2013-11-03\", \"Sunday\")]\n",
+ " )\n",
+ " \n",
+ " .split_column_by_example(source_column=\"pickup_time\")\n",
+ " .split_column_by_example(source_column=\"dropoff_time\")\n",
+ " # the following two split_column_by_example calls reference the generated column names from the above two calls\n",
+ " .split_column_by_example(source_column=\"pickup_time_1\")\n",
+ " .split_column_by_example(source_column=\"dropoff_time_1\")\n",
+ " .drop_columns(columns=[\n",
+ " \"pickup_date\", \"pickup_time\", \"dropoff_date\", \"dropoff_time\", \n",
+ " \"pickup_date_1\", \"dropoff_date_1\", \"pickup_time_1\", \"dropoff_time_1\"\n",
+ " ])\n",
+ " \n",
+ " .rename_columns(column_pairs={\n",
+ " \"pickup_date_2\": \"pickup_month\",\n",
+ " \"pickup_date_3\": \"pickup_monthday\",\n",
+ " \"pickup_time_1_1\": \"pickup_hour\",\n",
+ " \"pickup_time_1_2\": \"pickup_minute\",\n",
+ " \"pickup_time_2\": \"pickup_second\",\n",
+ " \"dropoff_date_2\": \"dropoff_month\",\n",
+ " \"dropoff_date_3\": \"dropoff_monthday\",\n",
+ " \"dropoff_time_1_1\": \"dropoff_hour\",\n",
+ " \"dropoff_time_1_2\": \"dropoff_minute\",\n",
+ " \"dropoff_time_2\": \"dropoff_second\"\n",
+ " }))\n",
+ "\n",
+ "tmp_df.head(5)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "From the data above, you see that the pickup and drop-off date and time components produced from the derived transformations are correct. Drop the `pickup_datetime` and `dropoff_datetime` columns as they are no longer needed."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "tmp_df = tmp_df.drop_columns(columns=[\"pickup_datetime\", \"dropoff_datetime\"])"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Use the type inference functionality to automatically check the data type of each field, and display the inference results."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "type_infer = tmp_df.builders.set_column_types()\n",
+ "type_infer.learn()\n",
+ "type_infer"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "The inference results look correct based on the data, now apply the type conversions to the dataflow."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "tmp_df = type_infer.to_dataflow()\n",
+ "tmp_df.get_profile()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Before packaging the dataflow, perform two final filters on the data set. To eliminate incorrect data points, filter the dataflow on records where both the `cost` and `distance` are greater than zero."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "tmp_df = tmp_df.filter(dprep.col(\"distance\") > 0)\n",
+ "tmp_df = tmp_df.filter(dprep.col(\"cost\") > 0)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "At this point, you have a fully transformed and prepared dataflow object to use in a machine learning model. The DataPrep SDK includes object serialization functionality, which is used as follows."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import os\n",
+ "file_path = os.path.join(os.getcwd(), \"dflows.dprep\")\n",
+ "\n",
+ "dflow_prepared = tmp_df\n",
+ "package = dprep.Package([dflow_prepared])\n",
+ "package.save(file_path)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Clean up resources"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Delete the file `dflows.dprep` (whether you are running locally or in Azure Notebooks) in your current directory if you do not wish to continue with part two of the tutorial. If you continue on to part two, you will need the `dflows.dprep` file in the current directory."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Next steps"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "In this Azure Machine Learning Data Prep SDK tutorial, you:\n",
+ "\n",
+ "> * Set up your development environment\n",
+ "> * Loaded and cleansed data sets\n",
+ "> * Used smart transforms to predict your logic based on an example\n",
+ "> * Merged and packaged datasets for machine learning training\n",
+ "\n",
+ "You are ready to use this training data in the next part of the tutorial series:\n",
+ "\n",
+ "\n",
+ "> [Tutorial #2: Train regression model](regression-part2-automated-ml.ipynb)"
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "cforbe"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.7"
+ },
+ "msauthor": "trbye"
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.7"
- },
- "msauthor": "trbye"
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file
diff --git a/tutorials/regression-part2-automated-ml.ipynb b/tutorials/regression-part2-automated-ml.ipynb
index b9504783..dca81bb6 100644
--- a/tutorials/regression-part2-automated-ml.ipynb
+++ b/tutorials/regression-part2-automated-ml.ipynb
@@ -1,502 +1,502 @@
{
- "cells": [
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Copyright (c) Microsoft Corporation. All rights reserved."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "# Tutorial (part 2): Use automated machine learning to build your regression model \n",
- "\n",
- "This tutorial is **part two of a two-part tutorial series**. In the previous tutorial, you [prepared the NYC taxi data for regression modeling](regression-part1-data-prep.ipynb).\n",
- "\n",
- "Now, you're ready to start building your model with Azure Machine Learning service. In this part of the tutorial, you will use the prepared data and automatically generate a regression model to predict taxi fare prices. Using the automated ML capabilities of the service, you define your machine learning goals and constraints, launch the automated machine learning process and then allow the algorithm selection and hyperparameter-tuning to happen for you. The automated ML technique iterates over many combinations of algorithms and hyperparameters until it finds the best model based on your criterion.\n",
- "\n",
- "In this tutorial, you learn how to:\n",
- "\n",
- "> * Setup a Python environment and import the SDK packages\n",
- "> * Configure an Azure Machine Learning service workspace\n",
- "> * Auto-train a regression model \n",
- "> * Run the model locally with custom parameters\n",
- "> * Explore the results\n",
- "> * Register the best model\n",
- "\n",
- "If you don’t have an Azure subscription, create a [free account](https://aka.ms/AMLfree) before you begin. \n",
- "\n",
- "> Code in this article was tested with Azure Machine Learning SDK version 1.0.0\n",
- "\n",
- "\n",
- "## Prerequisites\n",
- "\n",
- "> * [Run the data preparation tutorial](regression-part1-data-prep.ipynb)\n",
- "\n",
- "> * Automated machine learning configured environment e.g. Azure notebooks, Local Python environment or Data Science Virtual Machine. [Setup](https://docs.microsoft.com/azure/machine-learning/service/samples-notebooks) automated machine learning."
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Import packages\n",
- "Import Python packages you need in this tutorial."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import azureml.core\n",
- "import pandas as pd\n",
- "from azureml.core.workspace import Workspace\n",
- "import logging"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Configure workspace\n",
- "\n",
- "Create a workspace object from the existing workspace. A `Workspace` is a class that accepts your Azure subscription and resource information, and creates a cloud resource to monitor and track your model runs. `Workspace.from_config()` reads the file **aml_config/config.json** and loads the details into an object named `ws`. `ws` is used throughout the rest of the code in this tutorial.\n",
- "\n",
- "Once you have a workspace object, specify a name for the experiment and create and register a local directory with the workspace. The history of all runs is recorded under the specified experiment."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "ws = Workspace.from_config()\n",
- "# choose a name for the run history container in the workspace\n",
- "experiment_name = 'automated-ml-regression'\n",
- "# project folder\n",
- "project_folder = './automated-ml-regression'\n",
- "\n",
- "import os\n",
- "\n",
- "output = {}\n",
- "output['SDK version'] = azureml.core.VERSION\n",
- "output['Subscription ID'] = ws.subscription_id\n",
- "output['Workspace'] = ws.name\n",
- "output['Resource Group'] = ws.resource_group\n",
- "output['Location'] = ws.location\n",
- "output['Project Directory'] = project_folder\n",
- "pd.set_option('display.max_colwidth', -1)\n",
- "outputDf = pd.DataFrame(data = output, index = [''])\n",
- "outputDf.T"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Explore data\n",
- "\n",
- "Utilize the data flow object created in the previous tutorial. Open and execute the data flow and review the results."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import azureml.dataprep as dprep\n",
- "\n",
- "file_path = os.path.join(os.getcwd(), \"dflows.dprep\")\n",
- "\n",
- "package_saved = dprep.Package.open(file_path)\n",
- "dflow_prepared = package_saved.dataflows[0]\n",
- "dflow_prepared.get_profile()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "You prepare the data for the experiment by adding columns to `dflow_X` to be features for our model creation. You define `dflow_y` to be our prediction value; cost.\n"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "dflow_X = dflow_prepared.keep_columns(['pickup_weekday','pickup_hour', 'distance','passengers', 'vendor'])\n",
- "dflow_y = dflow_prepared.keep_columns('cost')"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Split data into train and test sets\n",
- "\n",
- "Now you split the data into training and test sets using the `train_test_split` function in the `sklearn` library. This function segregates the data into the x (features) data set for model training and the y (values to predict) data set for testing. The `test_size` parameter determines the percentage of data to allocate to testing. The `random_state` parameter sets a seed to the random generator, so that your train-test splits are always deterministic."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from sklearn.model_selection import train_test_split\n",
- "\n",
- "\n",
- "x_df = dflow_X.to_pandas_dataframe()\n",
- "y_df = dflow_y.to_pandas_dataframe()\n",
- "\n",
- "x_train, x_test, y_train, y_test = train_test_split(x_df, y_df, test_size=0.2, random_state=223)\n",
- "# flatten y_train to 1d array\n",
- "y_train.values.flatten()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "You now have the necessary packages and data ready for auto training for your model. \n",
- "\n",
- "## Automatically train a model\n",
- "\n",
- "To automatically train a model:\n",
- "1. Define settings for the experiment run\n",
- "1. Submit the experiment for model tuning\n",
- "\n",
- "\n",
- "### Define settings for autogeneration and tuning\n",
- "\n",
- "Define the experiment parameters and models settings for autogeneration and tuning. View the full list of [settings](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-configure-auto-train).\n",
- "\n",
- "\n",
- "|Property| Value in this tutorial |Description|\n",
- "|----|----|---|\n",
- "|**iteration_timeout_minutes**|10|Time limit in minutes for each iteration|\n",
- "|**iterations**|30|Number of iterations. In each iteration, the model trains with the data with a specific pipeline|\n",
- "|**primary_metric**|spearman_correlation | Metric that you want to optimize.|\n",
- "|**preprocess**| True | True enables experiment to perform preprocessing on the input.|\n",
- "|**verbosity**| logging.INFO | Controls the level of logging.|\n",
- "|**n_cross_validationss**|5|Number of cross validation splits\n"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "automl_settings = {\n",
- " \"iteration_timeout_minutes\" : 10,\n",
- " \"iterations\" : 30,\n",
- " \"primary_metric\" : 'spearman_correlation',\n",
- " \"preprocess\" : True,\n",
- " \"verbosity\" : logging.INFO,\n",
- " \"n_cross_validations\": 5\n",
- "}"
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "configure automl"
- ]
- },
- "outputs": [],
- "source": [
- "from azureml.train.automl import AutoMLConfig\n",
- "\n",
- "# local compute \n",
- "automated_ml_config = AutoMLConfig(task = 'regression',\n",
- " debug_log = 'automated_ml_errors.log',\n",
- " path = project_folder,\n",
- " X = x_train.values,\n",
- " y = y_train.values.flatten(),\n",
- " **automl_settings)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Train the automatic regression model\n",
- "\n",
- "Start the experiment to run locally. Pass the defined `automated_ml_config` object to the experiment, and set the output to `true` to view progress during the experiment."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "local submitted run",
- "automl"
- ]
- },
- "outputs": [],
- "source": [
- "from azureml.core.experiment import Experiment\n",
- "experiment=Experiment(ws, experiment_name)\n",
- "local_run = experiment.submit(automated_ml_config, show_output=True)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Explore the results\n",
- "\n",
- "Explore the results of automatic training with a Jupyter widget or by examining the experiment history.\n",
- "\n",
- "### Option 1: Add a Jupyter widget to see results\n",
- "\n",
- "Use the Jupyter notebook widget to see a graph and a table of all results."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "use notebook widget"
- ]
- },
- "outputs": [],
- "source": [
- "from azureml.widgets import RunDetails\n",
- "RunDetails(local_run).show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "### Option 2: Get and examine all run iterations in Python\n",
- "\n",
- "Alternatively, you can retrieve the history of each experiment and explore the individual metrics for each iteration run."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {
- "tags": [
- "get metrics",
- "query history"
- ]
- },
- "outputs": [],
- "source": [
- "children = list(local_run.get_children())\n",
- "metricslist = {}\n",
- "for run in children:\n",
- " properties = run.get_properties()\n",
- " metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)}\n",
- " metricslist[int(properties['iteration'])] = metrics\n",
- "\n",
- "rundata = pd.DataFrame(metricslist).sort_index(1)\n",
- "rundata"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Retrieve the best model\n",
- "\n",
- "Select the best pipeline from our iterations. The `get_output` method on `automl_classifier` returns the best run and the fitted model for the last fit invocation. There are overloads on `get_output` that allow you to retrieve the best run and fitted model for any logged metric or a particular iteration."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "best_run, fitted_model = local_run.get_output()\n",
- "print(best_run)\n",
- "print(fitted_model)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Register the model\n",
- "\n",
- "Register the model in your Azure Machine Learning Workspace."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "description = 'Automated Machine Learning Model'\n",
- "tags = None\n",
- "local_run.register_model(description=description, tags=tags)\n",
- "print(local_run.model_id) # Use this id to deploy the model as a web service in Azure"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Test the best model accuracy\n",
- "\n",
- "Use the best model to run predictions on the test data set. The function `predict` uses the best model, and predicts the values of y (trip cost) from the `x_test` data set. Print the first 10 predicted cost values from `y_predict`."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "y_predict = fitted_model.predict(x_test.values) \n",
- "print(y_predict[:10])"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Create a scatter plot to visualize the predicted cost values compared to the actual cost values. The following code uses the `distance` feature as the x-axis, and trip `cost` as the y-axis. The first 100 predicted and actual cost values are created as separate series, in order to compare the variance of predicted cost at each trip distance value. Examining the plot shows that the distance/cost relationship is nearly linear, and the predicted cost values are in most cases very close to the actual cost values for the same trip distance."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "import matplotlib.pyplot as plt\n",
- "\n",
- "fig = plt.figure(figsize=(14, 10))\n",
- "ax1 = fig.add_subplot(111)\n",
- "\n",
- "distance_vals = [x[4] for x in x_test.values]\n",
- "y_actual = y_test.values.flatten().tolist()\n",
- "\n",
- "ax1.scatter(distance_vals[:100], y_predict[:100], s=18, c='b', marker=\"s\", label='Predicted')\n",
- "ax1.scatter(distance_vals[:100], y_actual[:100], s=18, c='r', marker=\"o\", label='Actual')\n",
- "\n",
- "ax1.set_xlabel('distance (mi)')\n",
- "ax1.set_title('Predicted and Actual Cost/Distance')\n",
- "ax1.set_ylabel('Cost ($)')\n",
- "\n",
- "plt.legend(loc='upper left', prop={'size': 12})\n",
- "plt.rcParams.update({'font.size': 14})\n",
- "plt.show()"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Calculate the `root mean squared error` of the results. Use the `y_test` dataframe, and convert it to a list to compare to the predicted values. The function `mean_squared_error` takes two arrays of values, and calculates the average squared error between them. Taking the square root of the result gives an error in the same units as the y variable (cost), and indicates roughly how far your predictions are from the actual value. "
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "from sklearn.metrics import mean_squared_error\n",
- "from math import sqrt\n",
- "\n",
- "rmse = sqrt(mean_squared_error(y_actual, y_predict))\n",
- "rmse"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "Run the following code to calculate MAPE (mean absolute percent error) using the full `y_actual` and `y_predict` data sets. This metric calculates an absolute difference between each predicted and actual value, sums all the differences, and then expresses that sum as a percent of the total of the actual values."
- ]
- },
- {
- "cell_type": "code",
- "execution_count": null,
- "metadata": {},
- "outputs": [],
- "source": [
- "sum_actuals = sum_errors = 0\n",
- "\n",
- "for actual_val, predict_val in zip(y_actual, y_predict):\n",
- " abs_error = actual_val - predict_val\n",
- " if abs_error < 0:\n",
- " abs_error = abs_error * -1\n",
- " \n",
- " sum_errors = sum_errors + abs_error\n",
- " sum_actuals = sum_actuals + actual_val\n",
- " \n",
- "mean_abs_percent_error = sum_errors / sum_actuals\n",
- "print(\"Model MAPE:\")\n",
- "print(mean_abs_percent_error)\n",
- "print()\n",
- "print(\"Model Accuracy:\")\n",
- "print(1 - mean_abs_percent_error)"
- ]
- },
- {
- "cell_type": "markdown",
- "metadata": {},
- "source": [
- "## Next steps\n",
- "\n",
- "In this automated machine learning tutorial, you:\n",
- "\n",
- "\n",
- "> * Configured a workspace and prepared data for an experiment\n",
- "> * Trained using an automated regression model locally with custom parameters\n",
- "> * Explored and reviewed training results\n",
- "> * Registered the best model\n",
- "\n",
- "You can also try out the [image classification tutorial](img-classification-part1-training.ipynb)."
- ]
- }
- ],
- "metadata": {
- "authors": [
- {
- "name": "jeffshep"
- }
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Copyright (c) Microsoft Corporation. All rights reserved."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Tutorial (part 2): Use automated machine learning to build your regression model \n",
+ "\n",
+ "This tutorial is **part two of a two-part tutorial series**. In the previous tutorial, you [prepared the NYC taxi data for regression modeling](regression-part1-data-prep.ipynb).\n",
+ "\n",
+ "Now, you're ready to start building your model with Azure Machine Learning service. In this part of the tutorial, you will use the prepared data and automatically generate a regression model to predict taxi fare prices. Using the automated ML capabilities of the service, you define your machine learning goals and constraints, launch the automated machine learning process and then allow the algorithm selection and hyperparameter-tuning to happen for you. The automated ML technique iterates over many combinations of algorithms and hyperparameters until it finds the best model based on your criterion.\n",
+ "\n",
+ "In this tutorial, you learn how to:\n",
+ "\n",
+ "> * Setup a Python environment and import the SDK packages\n",
+ "> * Configure an Azure Machine Learning service workspace\n",
+ "> * Auto-train a regression model \n",
+ "> * Run the model locally with custom parameters\n",
+ "> * Explore the results\n",
+ "> * Register the best model\n",
+ "\n",
+ "If you don\u00e2\u20ac\u2122t have an Azure subscription, create a [free account](https://aka.ms/AMLfree) before you begin. \n",
+ "\n",
+ "> Code in this article was tested with Azure Machine Learning SDK version 1.0.0\n",
+ "\n",
+ "\n",
+ "## Prerequisites\n",
+ "\n",
+ "> * [Run the data preparation tutorial](regression-part1-data-prep.ipynb)\n",
+ "\n",
+ "> * Automated machine learning configured environment e.g. Azure notebooks, Local Python environment or Data Science Virtual Machine. [Setup](https://docs.microsoft.com/azure/machine-learning/service/samples-notebooks) automated machine learning."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Import packages\n",
+ "Import Python packages you need in this tutorial."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import azureml.core\n",
+ "import pandas as pd\n",
+ "from azureml.core.workspace import Workspace\n",
+ "import logging"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Configure workspace\n",
+ "\n",
+ "Create a workspace object from the existing workspace. A `Workspace` is a class that accepts your Azure subscription and resource information, and creates a cloud resource to monitor and track your model runs. `Workspace.from_config()` reads the file **aml_config/config.json** and loads the details into an object named `ws`. `ws` is used throughout the rest of the code in this tutorial.\n",
+ "\n",
+ "Once you have a workspace object, specify a name for the experiment and create and register a local directory with the workspace. The history of all runs is recorded under the specified experiment."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "ws = Workspace.from_config()\n",
+ "# choose a name for the run history container in the workspace\n",
+ "experiment_name = 'automated-ml-regression'\n",
+ "# project folder\n",
+ "project_folder = './automated-ml-regression'\n",
+ "\n",
+ "import os\n",
+ "\n",
+ "output = {}\n",
+ "output['SDK version'] = azureml.core.VERSION\n",
+ "output['Subscription ID'] = ws.subscription_id\n",
+ "output['Workspace'] = ws.name\n",
+ "output['Resource Group'] = ws.resource_group\n",
+ "output['Location'] = ws.location\n",
+ "output['Project Directory'] = project_folder\n",
+ "pd.set_option('display.max_colwidth', -1)\n",
+ "outputDf = pd.DataFrame(data = output, index = [''])\n",
+ "outputDf.T"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Explore data\n",
+ "\n",
+ "Utilize the data flow object created in the previous tutorial. Open and execute the data flow and review the results."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import azureml.dataprep as dprep\n",
+ "\n",
+ "file_path = os.path.join(os.getcwd(), \"dflows.dprep\")\n",
+ "\n",
+ "package_saved = dprep.Package.open(file_path)\n",
+ "dflow_prepared = package_saved.dataflows[0]\n",
+ "dflow_prepared.get_profile()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "You prepare the data for the experiment by adding columns to `dflow_X` to be features for our model creation. You define `dflow_y` to be our prediction value; cost.\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "dflow_X = dflow_prepared.keep_columns(['pickup_weekday','pickup_hour', 'distance','passengers', 'vendor'])\n",
+ "dflow_y = dflow_prepared.keep_columns('cost')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Split data into train and test sets\n",
+ "\n",
+ "Now you split the data into training and test sets using the `train_test_split` function in the `sklearn` library. This function segregates the data into the x (features) data set for model training and the y (values to predict) data set for testing. The `test_size` parameter determines the percentage of data to allocate to testing. The `random_state` parameter sets a seed to the random generator, so that your train-test splits are always deterministic."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from sklearn.model_selection import train_test_split\n",
+ "\n",
+ "\n",
+ "x_df = dflow_X.to_pandas_dataframe()\n",
+ "y_df = dflow_y.to_pandas_dataframe()\n",
+ "\n",
+ "x_train, x_test, y_train, y_test = train_test_split(x_df, y_df, test_size=0.2, random_state=223)\n",
+ "# flatten y_train to 1d array\n",
+ "y_train.values.flatten()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "You now have the necessary packages and data ready for auto training for your model. \n",
+ "\n",
+ "## Automatically train a model\n",
+ "\n",
+ "To automatically train a model:\n",
+ "1. Define settings for the experiment run\n",
+ "1. Submit the experiment for model tuning\n",
+ "\n",
+ "\n",
+ "### Define settings for autogeneration and tuning\n",
+ "\n",
+ "Define the experiment parameters and models settings for autogeneration and tuning. View the full list of [settings](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-configure-auto-train).\n",
+ "\n",
+ "\n",
+ "|Property| Value in this tutorial |Description|\n",
+ "|----|----|---|\n",
+ "|**iteration_timeout_minutes**|10|Time limit in minutes for each iteration|\n",
+ "|**iterations**|30|Number of iterations. In each iteration, the model trains with the data with a specific pipeline|\n",
+ "|**primary_metric**|spearman_correlation | Metric that you want to optimize.|\n",
+ "|**preprocess**| True | True enables experiment to perform preprocessing on the input.|\n",
+ "|**verbosity**| logging.INFO | Controls the level of logging.|\n",
+ "|**n_cross_validationss**|5|Number of cross validation splits\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "automl_settings = {\n",
+ " \"iteration_timeout_minutes\" : 10,\n",
+ " \"iterations\" : 30,\n",
+ " \"primary_metric\" : 'spearman_correlation',\n",
+ " \"preprocess\" : True,\n",
+ " \"verbosity\" : logging.INFO,\n",
+ " \"n_cross_validations\": 5\n",
+ "}"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "configure automl"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "from azureml.train.automl import AutoMLConfig\n",
+ "\n",
+ "# local compute \n",
+ "automated_ml_config = AutoMLConfig(task = 'regression',\n",
+ " debug_log = 'automated_ml_errors.log',\n",
+ " path = project_folder,\n",
+ " X = x_train.values,\n",
+ " y = y_train.values.flatten(),\n",
+ " **automl_settings)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Train the automatic regression model\n",
+ "\n",
+ "Start the experiment to run locally. Pass the defined `automated_ml_config` object to the experiment, and set the output to `true` to view progress during the experiment."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "local submitted run",
+ "automl"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "from azureml.core.experiment import Experiment\n",
+ "experiment=Experiment(ws, experiment_name)\n",
+ "local_run = experiment.submit(automated_ml_config, show_output=True)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Explore the results\n",
+ "\n",
+ "Explore the results of automatic training with a Jupyter widget or by examining the experiment history.\n",
+ "\n",
+ "### Option 1: Add a Jupyter widget to see results\n",
+ "\n",
+ "Use the Jupyter notebook widget to see a graph and a table of all results."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "use notebook widget"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "from azureml.widgets import RunDetails\n",
+ "RunDetails(local_run).show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Option 2: Get and examine all run iterations in Python\n",
+ "\n",
+ "Alternatively, you can retrieve the history of each experiment and explore the individual metrics for each iteration run."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "tags": [
+ "get metrics",
+ "query history"
+ ]
+ },
+ "outputs": [],
+ "source": [
+ "children = list(local_run.get_children())\n",
+ "metricslist = {}\n",
+ "for run in children:\n",
+ " properties = run.get_properties()\n",
+ " metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)}\n",
+ " metricslist[int(properties['iteration'])] = metrics\n",
+ "\n",
+ "rundata = pd.DataFrame(metricslist).sort_index(1)\n",
+ "rundata"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Retrieve the best model\n",
+ "\n",
+ "Select the best pipeline from our iterations. The `get_output` method on `automl_classifier` returns the best run and the fitted model for the last fit invocation. There are overloads on `get_output` that allow you to retrieve the best run and fitted model for any logged metric or a particular iteration."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "best_run, fitted_model = local_run.get_output()\n",
+ "print(best_run)\n",
+ "print(fitted_model)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Register the model\n",
+ "\n",
+ "Register the model in your Azure Machine Learning Workspace."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "description = 'Automated Machine Learning Model'\n",
+ "tags = None\n",
+ "local_run.register_model(description=description, tags=tags)\n",
+ "print(local_run.model_id) # Use this id to deploy the model as a web service in Azure"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Test the best model accuracy\n",
+ "\n",
+ "Use the best model to run predictions on the test data set. The function `predict` uses the best model, and predicts the values of y (trip cost) from the `x_test` data set. Print the first 10 predicted cost values from `y_predict`."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "y_predict = fitted_model.predict(x_test.values) \n",
+ "print(y_predict[:10])"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Create a scatter plot to visualize the predicted cost values compared to the actual cost values. The following code uses the `distance` feature as the x-axis, and trip `cost` as the y-axis. The first 100 predicted and actual cost values are created as separate series, in order to compare the variance of predicted cost at each trip distance value. Examining the plot shows that the distance/cost relationship is nearly linear, and the predicted cost values are in most cases very close to the actual cost values for the same trip distance."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "import matplotlib.pyplot as plt\n",
+ "\n",
+ "fig = plt.figure(figsize=(14, 10))\n",
+ "ax1 = fig.add_subplot(111)\n",
+ "\n",
+ "distance_vals = [x[4] for x in x_test.values]\n",
+ "y_actual = y_test.values.flatten().tolist()\n",
+ "\n",
+ "ax1.scatter(distance_vals[:100], y_predict[:100], s=18, c='b', marker=\"s\", label='Predicted')\n",
+ "ax1.scatter(distance_vals[:100], y_actual[:100], s=18, c='r', marker=\"o\", label='Actual')\n",
+ "\n",
+ "ax1.set_xlabel('distance (mi)')\n",
+ "ax1.set_title('Predicted and Actual Cost/Distance')\n",
+ "ax1.set_ylabel('Cost ($)')\n",
+ "\n",
+ "plt.legend(loc='upper left', prop={'size': 12})\n",
+ "plt.rcParams.update({'font.size': 14})\n",
+ "plt.show()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Calculate the `root mean squared error` of the results. Use the `y_test` dataframe, and convert it to a list to compare to the predicted values. The function `mean_squared_error` takes two arrays of values, and calculates the average squared error between them. Taking the square root of the result gives an error in the same units as the y variable (cost), and indicates roughly how far your predictions are from the actual value. "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "from sklearn.metrics import mean_squared_error\n",
+ "from math import sqrt\n",
+ "\n",
+ "rmse = sqrt(mean_squared_error(y_actual, y_predict))\n",
+ "rmse"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Run the following code to calculate MAPE (mean absolute percent error) using the full `y_actual` and `y_predict` data sets. This metric calculates an absolute difference between each predicted and actual value, sums all the differences, and then expresses that sum as a percent of the total of the actual values."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {},
+ "outputs": [],
+ "source": [
+ "sum_actuals = sum_errors = 0\n",
+ "\n",
+ "for actual_val, predict_val in zip(y_actual, y_predict):\n",
+ " abs_error = actual_val - predict_val\n",
+ " if abs_error < 0:\n",
+ " abs_error = abs_error * -1\n",
+ " \n",
+ " sum_errors = sum_errors + abs_error\n",
+ " sum_actuals = sum_actuals + actual_val\n",
+ " \n",
+ "mean_abs_percent_error = sum_errors / sum_actuals\n",
+ "print(\"Model MAPE:\")\n",
+ "print(mean_abs_percent_error)\n",
+ "print()\n",
+ "print(\"Model Accuracy:\")\n",
+ "print(1 - mean_abs_percent_error)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Next steps\n",
+ "\n",
+ "In this automated machine learning tutorial, you:\n",
+ "\n",
+ "\n",
+ "> * Configured a workspace and prepared data for an experiment\n",
+ "> * Trained using an automated regression model locally with custom parameters\n",
+ "> * Explored and reviewed training results\n",
+ "> * Registered the best model\n",
+ "\n",
+ "[Deploy your model](02.deploy-models.ipynb) with Azure Machine Learning."
+ ]
+ }
],
- "kernelspec": {
- "display_name": "Python 3.6",
- "language": "python",
- "name": "python36"
+ "metadata": {
+ "authors": [
+ {
+ "name": "jeffshep"
+ }
+ ],
+ "kernelspec": {
+ "display_name": "Python 3.6",
+ "language": "python",
+ "name": "python36"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.6.7"
+ },
+ "msauthor": "sgilley"
},
- "language_info": {
- "codemirror_mode": {
- "name": "ipython",
- "version": 3
- },
- "file_extension": ".py",
- "mimetype": "text/x-python",
- "name": "python",
- "nbconvert_exporter": "python",
- "pygments_lexer": "ipython3",
- "version": "3.6.7"
- },
- "msauthor": "sgilley"
- },
- "nbformat": 4,
- "nbformat_minor": 2
-}
+ "nbformat": 4,
+ "nbformat_minor": 2
+}
\ No newline at end of file